A genome-wide search for pleiotropy in more than 100,000 harmonized longitudinal cognitive domain scores
- PMID: 37349795
- PMCID: PMC10286470
- DOI: 10.1186/s13024-023-00633-4
A genome-wide search for pleiotropy in more than 100,000 harmonized longitudinal cognitive domain scores
Abstract
Background: More than 75 common variant loci account for only a portion of the heritability for Alzheimer's disease (AD). A more complete understanding of the genetic basis of AD can be deduced by exploring associations with AD-related endophenotypes.
Methods: We conducted genome-wide scans for cognitive domain performance using harmonized and co-calibrated scores derived by confirmatory factor analyses for executive function, language, and memory. We analyzed 103,796 longitudinal observations from 23,066 members of community-based (FHS, ACT, and ROSMAP) and clinic-based (ADRCs and ADNI) cohorts using generalized linear mixed models including terms for SNP, age, SNP × age interaction, sex, education, and five ancestry principal components. Significance was determined based on a joint test of the SNP's main effect and interaction with age. Results across datasets were combined using inverse-variance meta-analysis. Genome-wide tests of pleiotropy for each domain pair as the outcome were performed using PLACO software.
Results: Individual domain and pleiotropy analyses revealed genome-wide significant (GWS) associations with five established loci for AD and AD-related disorders (BIN1, CR1, GRN, MS4A6A, and APOE) and eight novel loci. ULK2 was associated with executive function in the community-based cohorts (rs157405, P = 2.19 × 10-9). GWS associations for language were identified with CDK14 in the clinic-based cohorts (rs705353, P = 1.73 × 10-8) and LINC02712 in the total sample (rs145012974, P = 3.66 × 10-8). GRN (rs5848, P = 4.21 × 10-8) and PURG (rs117523305, P = 1.73 × 10-8) were associated with memory in the total and community-based cohorts, respectively. GWS pleiotropy was observed for language and memory with LOC107984373 (rs73005629, P = 3.12 × 10-8) in the clinic-based cohorts, and with NCALD (rs56162098, P = 1.23 × 10-9) and PTPRD (rs145989094, P = 8.34 × 10-9) in the community-based cohorts. GWS pleiotropy was also found for executive function and memory with OSGIN1 (rs12447050, P = 4.09 × 10-8) and PTPRD (rs145989094, P = 3.85 × 10-8) in the community-based cohorts. Functional studies have previously linked AD to ULK2, NCALD, and PTPRD.
Conclusion: Our results provide some insight into biological pathways underlying processes leading to domain-specific cognitive impairment and AD, as well as a conduit toward a syndrome-specific precision medicine approach to AD. Increasing the number of participants with harmonized cognitive domain scores will enhance the discovery of additional genetic factors of cognitive decline leading to AD and related dementias.
Keywords: Alzheimer’s disease; Cognitive domains; Genome-wide association study; Longitudinal measures; Pathway analysis; Pleiotropy.
© 2023. The Author(s).
Conflict of interest statement
The authors declare they have no competing interests.
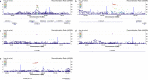
Figures

Similar articles
-
CpG-related SNPs in the MS4A region have a dose-dependent effect on risk of late-onset Alzheimer disease.Aging Cell. 2019 Aug;18(4):e12964. doi: 10.1111/acel.12964. Epub 2019 May 29. Aging Cell. 2019. PMID: 31144443 Free PMC article.
-
A novel Alzheimer disease locus located near the gene encoding tau protein.Mol Psychiatry. 2016 Jan;21(1):108-17. doi: 10.1038/mp.2015.23. Epub 2015 Mar 17. Mol Psychiatry. 2016. PMID: 25778476 Free PMC article.
-
Meta-analysis for genome-wide association study identifies multiple variants at the BIN1 locus associated with late-onset Alzheimer's disease.PLoS One. 2011 Feb 24;6(2):e16616. doi: 10.1371/journal.pone.0016616. PLoS One. 2011. PMID: 21390209 Free PMC article.
-
Gene-based aggregate SNP associations between candidate AD genes and cognitive decline.Age (Dordr). 2016 Apr;38(2):41. doi: 10.1007/s11357-016-9885-2. Epub 2016 Mar 22. Age (Dordr). 2016. PMID: 27005436 Free PMC article. Review.
-
Genetic studies of quantitative MCI and AD phenotypes in ADNI: Progress, opportunities, and plans.Alzheimers Dement. 2015 Jul;11(7):792-814. doi: 10.1016/j.jalz.2015.05.009. Alzheimers Dement. 2015. PMID: 26194313 Free PMC article. Review.
Cited by
-
Mechanisms and Functions of the Cerebral-Cognitive Reserve in Patients with Alzheimer's Disease: A Narrative Review.Consort Psychiatr. 2024 Aug 26;5(3):17-29. doi: 10.17816/CP15526. eCollection 2024. Consort Psychiatr. 2024. PMID: 39526013 Free PMC article. Review.
-
Genetics of Alzheimer's Disease in the African American Population.J Clin Med. 2023 Aug 9;12(16):5189. doi: 10.3390/jcm12165189. J Clin Med. 2023. PMID: 37629231 Free PMC article. Review.
-
The Human Accelerated Region HAR202 Controls NPAS3 Expression in the Developing Forebrain Displaying Differential Enhancer Activity Between Modern and Archaic Human Sequences.Mol Biol Evol. 2024 Oct 4;41(10):msae186. doi: 10.1093/molbev/msae186. Mol Biol Evol. 2024. PMID: 39241178 Free PMC article.
-
A cross-trait study of lung cancer and its related respiratory diseases based on large-scale exome sequencing population.Transl Lung Cancer Res. 2024 Mar 29;13(3):512-525. doi: 10.21037/tlcr-24-4. Epub 2024 Mar 14. Transl Lung Cancer Res. 2024. PMID: 38601445 Free PMC article.
-
A Transcriptional Signature of Induced Neurons Differentiates Virologically Suppressed People Living With HIV from People Without HIV.bioRxiv [Preprint]. 2024 Oct 22:2024.10.22.619617. doi: 10.1101/2024.10.22.619617. bioRxiv. 2024. PMID: 39484396 Free PMC article. Preprint.
References
-
- Lee SH, Harold D, Nyholt DR, Consortium AN. International Endogene C. Genetic. Environmental Risk for Alzheimer’s disease C. Goddard ME, Zondervan KT, Williams J, et al. Estimation and partitioning of polygenic variation captured by common SNPs for Alzheimer’s disease, multiple sclerosis and endometriosis. Hum Mol Genet. 2013;22:832–841. doi: 10.1093/hmg/dds491. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- U24 AG021886/AG/NIA NIH HHS/United States
- R01 AG073439/AG/NIA NIH HHS/United States
- U01 AG068057/AG/NIA NIH HHS/United States
- U01 AG016976/AG/NIA NIH HHS/United States
- U24 AG074855/AG/NIA NIH HHS/United States
- R01-AG048927/AG/NIA NIH HHS/United States
- R01 AG059716/AG/NIA NIH HHS/United States
- U01-AG068057/AG/NIA NIH HHS/United States
- U54-AG052427/AG/NIA NIH HHS/United States
- P30-AG072878/AG/NIA NIH HHS/United States
- R01-AG059716/AG/NIA NIH HHS/United States
- U01-AG062602/AG/NIA NIH HHS/United States
- U01-AG058654/AG/NIA NIH HHS/United States
- U24-AG074855/AG/NIA NIH HHS/United States
- U01-AG032984/AG/NIA NIH HHS/United States
- U19-AG068753/AG/NIA NIH HHS/United States
- RF1-AG057519/AG/NIA NIH HHS/United States
- U24 AG041689/AG/NIA NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

