Membrane-bound mRNA immunogens lower the threshold to activate HIV Env V2 apex-directed broadly neutralizing B cell precursors in humanized mice
- PMID: 36179690
- PMCID: PMC9671093
- DOI: 10.1016/j.immuni.2022.09.003
Membrane-bound mRNA immunogens lower the threshold to activate HIV Env V2 apex-directed broadly neutralizing B cell precursors in humanized mice
Abstract
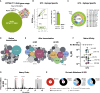
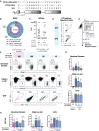
Eliciting broadly neutralizing antibodies (bnAbs) is the core of HIV vaccine design. bnAbs specific to the V2-apex region of the HIV envelope acquire breadth and potency with modest somatic hypermutation, making them attractive vaccination targets. To evaluate Apex germline-targeting (ApexGT) vaccine candidates, we engineered knockin (KI) mouse models expressing the germline B cell receptor (BCR) of the bnAb PCT64. We found that high affinity of the ApexGT immunogen for PCT64-germline BCRs was necessary to specifically activate KI B cells at human physiological frequencies, recruit them to germinal centers, and select for mature bnAb mutations. Relative to protein, mRNA-encoded membrane-bound ApexGT immunization significantly increased activation and recruitment of PCT64 precursors to germinal centers and lowered their affinity threshold. We have thus developed additional models for HIV vaccine research, validated ApexGT immunogens for priming V2-apex bnAb precursors, and identified mRNA-LNP as a suitable approach to substantially improve the B cell response.
Keywords: B cell receptor; HIV vaccine; V2 apex; antibody; bnAbs; immunization; knockin; mRNA.
Copyright © 2022 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests J.R.W., K.M.M., J.M.S., and W.R.S. are named inventors on patent applications filed by Scripps and IAVI regarding ApexGT immunogens in this manuscript.
Figures






Similar articles
-
Human immunoglobulin repertoire analysis guides design of vaccine priming immunogens targeting HIV V2-apex broadly neutralizing antibody precursors.Immunity. 2022 Nov 8;55(11):2149-2167.e9. doi: 10.1016/j.immuni.2022.09.001. Epub 2022 Sep 29. Immunity. 2022. PMID: 36179689 Free PMC article.
-
Neutralizing Antibody Responses Induced by HIV-1 Envelope Glycoprotein SOSIP Trimers Derived from Elite Neutralizers.J Virol. 2020 Nov 23;94(24):e01214-20. doi: 10.1128/JVI.01214-20. Print 2020 Nov 23. J Virol. 2020. PMID: 32999024 Free PMC article.
-
Antibodies from Rabbits Immunized with HIV-1 Clade B SOSIP Trimers Can Neutralize Multiple Clade B Viruses by Destabilizing the Envelope Glycoprotein.J Virol. 2021 Aug 10;95(17):e0009421. doi: 10.1128/JVI.00094-21. Epub 2021 Aug 10. J Virol. 2021. PMID: 34076487 Free PMC article.
-
Depressing time: Waiting, melancholia, and the psychoanalytic practice of care.In: Kirtsoglou E, Simpson B, editors. The Time of Anthropology: Studies of Contemporary Chronopolitics. Abingdon: Routledge; 2020. Chapter 5. In: Kirtsoglou E, Simpson B, editors. The Time of Anthropology: Studies of Contemporary Chronopolitics. Abingdon: Routledge; 2020. Chapter 5. PMID: 36137063 Free Books & Documents. Review.
-
Env Exceptionalism: Why Are HIV-1 Env Glycoproteins Atypical Immunogens?Cell Host Microbe. 2020 Apr 8;27(4):507-518. doi: 10.1016/j.chom.2020.03.018. Cell Host Microbe. 2020. PMID: 32272076 Free PMC article. Review.
Cited by
-
mRNA-LNP prime boost evolves precursors toward VRC01-like broadly neutralizing antibodies in preclinical humanized mouse models.Sci Immunol. 2024 May 10;9(95):eadn0622. doi: 10.1126/sciimmunol.adn0622. Epub 2024 May 16. Sci Immunol. 2024. PMID: 38753808 Free PMC article.
-
Unmodified rabies mRNA vaccine elicits high cross-neutralizing antibody titers and diverse B cell memory responses.Nat Commun. 2023 Jun 22;14(1):3713. doi: 10.1038/s41467-023-39421-5. Nat Commun. 2023. PMID: 37349310 Free PMC article.
-
Structure and Dynamics Guiding Design of Antibody Therapeutics and Vaccines.Antibodies (Basel). 2023 Oct 18;12(4):67. doi: 10.3390/antib12040067. Antibodies (Basel). 2023. PMID: 37873864 Free PMC article. Review.
-
Nucleic Acid Vaccines Encoding Proteins and Virus-like Particles for HIV Prevention.Vaccines (Basel). 2024 Mar 12;12(3):298. doi: 10.3390/vaccines12030298. Vaccines (Basel). 2024. PMID: 38543932 Free PMC article. Review.
-
mRNA vaccines for infectious diseases - advances, challenges and opportunities.Nat Rev Drug Discov. 2024 Nov;23(11):838-861. doi: 10.1038/s41573-024-01042-y. Epub 2024 Oct 4. Nat Rev Drug Discov. 2024. PMID: 39367276 Review.
References
-
- Abbott R.K., Lee J.H., Menis S., Skog P., Rossi M., Ota T., Kulp D.W., Bhullar D., Kalyuzhniy O., Havenar-Daughton C., et al. Precursor frequency and affinity determine B cell competitive fitness in germinal centers, tested with germline-targeting HIV vaccine immunogens. Immunity. 2018;48:133–146. doi: 10.1016/j.immuni.2017.11.023. - DOI - PMC - PubMed
-
- Andrabi R., Voss J.E., Liang C.H., Briney B., McCoy L.E., Wu C.Y., Wong C.H., Poignard P., Burton D.R. Identification of common features in prototype broadly neutralizing antibodies to HIV envelope V2 apex to facilitate vaccine design. Immunity. 2015;43:959–973. doi: 10.1016/j.immuni.2015.10.014. - DOI - PMC - PubMed
-
- Andrabi R., Su C.Y., Liang C.H., Shivatare S.S., Briney B., Voss J.E., Nawazi S.K., Wu C.Y., Wong C.H., Burton D.R. Glycans function as anchors for antibodies and help drive HIV broadly neutralizing antibody development. Immunity. 2017;47:524–537.e3. doi: 10.1016/j.immuni.2017.08.006. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

