Molecular Mechanism for Antibody-Dependent Enhancement of Coronavirus Entry
- PMID: 31826992
- PMCID: PMC7022351
- DOI: 10.1128/JVI.02015-19
Molecular Mechanism for Antibody-Dependent Enhancement of Coronavirus Entry
Abstract
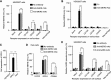
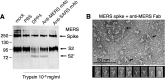
Antibody-dependent enhancement (ADE) of viral entry has been a major concern for epidemiology, vaccine development, and antibody-based drug therapy. However, the molecular mechanism behind ADE is still elusive. Coronavirus spike protein mediates viral entry into cells by first binding to a receptor on the host cell surface and then fusing viral and host membranes. In this study, we investigated how a neutralizing monoclonal antibody (MAb), which targets the receptor-binding domain (RBD) of Middle East respiratory syndrome (MERS) coronavirus spike, mediates viral entry using pseudovirus entry and biochemical assays. Our results showed that MAb binds to the virus surface spike, allowing it to undergo conformational changes and become prone to proteolytic activation. Meanwhile, MAb binds to cell surface IgG Fc receptor, guiding viral entry through canonical viral-receptor-dependent pathways. Our data suggest that the antibody/Fc-receptor complex functionally mimics viral receptor in mediating viral entry. Moreover, we characterized MAb dosages in viral-receptor-dependent, Fc-receptor-dependent, and both-receptors-dependent viral entry pathways, delineating guidelines on MAb usages in treating viral infections. Our study reveals a novel molecular mechanism for antibody-enhanced viral entry and can guide future vaccination and antiviral strategies.IMPORTANCE Antibody-dependent enhancement (ADE) of viral entry has been observed for many viruses. It was shown that antibodies target one serotype of viruses but only subneutralize another, leading to ADE of the latter viruses. Here we identify a novel mechanism for ADE: a neutralizing antibody binds to the surface spike protein of coronaviruses like a viral receptor, triggers a conformational change of the spike, and mediates viral entry into IgG Fc receptor-expressing cells through canonical viral-receptor-dependent pathways. We further evaluated how antibody dosages impacted viral entry into cells expressing viral receptor, Fc receptor, or both receptors. This study reveals complex roles of antibodies in viral entry and can guide future vaccine design and antibody-based drug therapy.
Keywords: IgG Fc receptor; MERS coronavirus; SARS coronavirus; antibody-dependent enhancement of viral entry; neutralizing antibody; spike protein; viral receptor.
Copyright © 2020 American Society for Microbiology.
Figures



Similar articles
-
Recombinant Receptor-Binding Domains of Multiple Middle East Respiratory Syndrome Coronaviruses (MERS-CoVs) Induce Cross-Neutralizing Antibodies against Divergent Human and Camel MERS-CoVs and Antibody Escape Mutants.J Virol. 2016 Dec 16;91(1):e01651-16. doi: 10.1128/JVI.01651-16. Print 2017 Jan 1. J Virol. 2016. PMID: 27795425 Free PMC article.
-
Mutations in the Spike Protein of Middle East Respiratory Syndrome Coronavirus Transmitted in Korea Increase Resistance to Antibody-Mediated Neutralization.J Virol. 2019 Jan 4;93(2):e01381-18. doi: 10.1128/JVI.01381-18. Print 2019 Jan 15. J Virol. 2019. PMID: 30404801 Free PMC article.
-
Structural definition of a neutralization epitope on the N-terminal domain of MERS-CoV spike glycoprotein.Nat Commun. 2019 Jul 11;10(1):3068. doi: 10.1038/s41467-019-10897-4. Nat Commun. 2019. PMID: 31296843 Free PMC article.
-
MERS-CoV spike protein: Targets for vaccines and therapeutics.Antiviral Res. 2016 Sep;133:165-77. doi: 10.1016/j.antiviral.2016.07.015. Epub 2016 Jul 26. Antiviral Res. 2016. PMID: 27468951 Free PMC article. Review.
-
Neutralizing Monoclonal Antibodies as Promising Therapeutics against Middle East Respiratory Syndrome Coronavirus Infection.Viruses. 2018 Nov 30;10(12):680. doi: 10.3390/v10120680. Viruses. 2018. PMID: 30513619 Free PMC article. Review.
Cited by
-
Design of customized coronavirus receptors.Nature. 2024 Oct 30. doi: 10.1038/s41586-024-08121-5. Online ahead of print. Nature. 2024. PMID: 39478224
-
Safety and Immunogenicity Study of a Bivalent Vaccine for Combined Prophylaxis of COVID-19 and Influenza in Non-Human Primates.Vaccines (Basel). 2024 Sep 26;12(10):1099. doi: 10.3390/vaccines12101099. Vaccines (Basel). 2024. PMID: 39460266 Free PMC article.
-
Enhanced RBD-Specific Antibody Responses and SARS-CoV-2-Relevant T-Cell Activity in Healthcare Workers Following Booster Vaccination.Curr Issues Mol Biol. 2024 Oct 2;46(10):11124-11135. doi: 10.3390/cimb46100660. Curr Issues Mol Biol. 2024. PMID: 39451540 Free PMC article.
-
Emerging and reemerging infectious diseases: global trends and new strategies for their prevention and control.Signal Transduct Target Ther. 2024 Sep 11;9(1):223. doi: 10.1038/s41392-024-01917-x. Signal Transduct Target Ther. 2024. PMID: 39256346 Free PMC article. Review.
-
Dual-role epitope on SARS-CoV-2 spike enhances and neutralizes viral entry across different variants.PLoS Pathog. 2024 Sep 5;20(9):e1012493. doi: 10.1371/journal.ppat.1012493. eCollection 2024 Sep. PLoS Pathog. 2024. PMID: 39236072 Free PMC article.
References
-
- Dejnirattisai W, Jumnainsong A, Onsirisakul N, Fitton P, Vasanawathana S, Limpitikul W, Puttikhunt C, Edwards C, Duangchinda T, Supasa S, Chawansuntati K, Malasit P, Mongkolsapaya J, Screaton G. 2010. Cross-reacting antibodies enhance dengue virus infection in humans. Science 328:745–748. doi:10.1126/science.1185181. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

