Synergistic and independent actions of multiple terminal nucleotidyl transferases in the 3' tailing of small RNAs in Arabidopsis
- PMID: 25928341
- PMCID: PMC4415790
- DOI: 10.1371/journal.pgen.1005091
Synergistic and independent actions of multiple terminal nucleotidyl transferases in the 3' tailing of small RNAs in Arabidopsis
Abstract
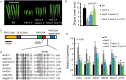
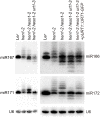
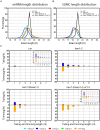
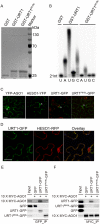
All types of small RNAs in plants, piwi-interacting RNAs (piRNAs) in animals and a subset of siRNAs in Drosophila and C. elegans are subject to HEN1 mediated 3' terminal 2'-O-methylation. This modification plays a pivotal role in protecting small RNAs from 3' uridylation, trimming and degradation. In Arabidopsis, HESO1 is a major enzyme that uridylates small RNAs to trigger their degradation. However, U-tail is still present in null hen1 heso1 mutants, suggesting the existence of (an) enzymatic activities redundant with HESO1. Here, we report that UTP: RNA uridylyltransferase (URT1) is a functional paralog of HESO1. URT1 interacts with AGO1 and plays a predominant role in miRNA uridylation when HESO1 is absent. Uridylation of miRNA is globally abolished in a hen1 heso1 urt1 triple mutant, accompanied by an extensive increase of 3'-to-5' trimming. In contrast, disruption of URT1 appears not to affect the heterochromatic siRNA uridylation. This indicates the involvement of additional nucleotidyl transferases in the siRNA pathway. Analysis of miRNA tailings in the hen1 heso1 urt1 triple mutant also reveals the existence of previously unknown enzymatic activities that can add non-uridine nucleotides. Importantly, we show HESO1 may also act redundantly with URT1 in miRNA uridylation when HEN1 is fully competent. Taken together, our data not only reveal a synergistic action of HESO1 and URT1 in the 3' uridylation of miRNAs, but also independent activities of multiple terminal nucleotidyl transferases in the 3' tailing of small RNAs and an antagonistic relationship between uridylation and trimming. Our results may provide further insight into the mechanisms of small RNA 3' end modification and stability control.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Distinct and cooperative activities of HESO1 and URT1 nucleotidyl transferases in microRNA turnover in Arabidopsis.PLoS Genet. 2015 Apr 30;11(4):e1005119. doi: 10.1371/journal.pgen.1005119. eCollection 2015 Apr. PLoS Genet. 2015. PMID: 25928405 Free PMC article.
-
The Arabidopsis nucleotidyl transferase HESO1 uridylates unmethylated small RNAs to trigger their degradation.Curr Biol. 2012 Apr 24;22(8):689-94. doi: 10.1016/j.cub.2012.02.051. Epub 2012 Mar 29. Curr Biol. 2012. PMID: 22464194 Free PMC article.
-
Uridylation and the SKI complex orchestrate the Calvin cycle of photosynthesis through RNA surveillance of TKL1 in Arabidopsis.Proc Natl Acad Sci U S A. 2022 Sep 20;119(38):e2205842119. doi: 10.1073/pnas.2205842119. Epub 2022 Sep 12. Proc Natl Acad Sci U S A. 2022. PMID: 36095196 Free PMC article.
-
Functions and mechanisms of RNA tailing by nucleotidyl transferase proteins in plants.Front Plant Sci. 2024 Oct 15;15:1452347. doi: 10.3389/fpls.2024.1452347. eCollection 2024. Front Plant Sci. 2024. PMID: 39474218 Free PMC article. Review.
-
RNA decay via 3' uridylation.Biochim Biophys Acta. 2013 Jun-Jul;1829(6-7):654-65. doi: 10.1016/j.bbagrm.2013.01.009. Epub 2013 Feb 4. Biochim Biophys Acta. 2013. PMID: 23385389 Review.
Cited by
-
Biogenesis of a 22-nt microRNA in Phaseoleae species by precursor-programmed uridylation.Proc Natl Acad Sci U S A. 2018 Jul 31;115(31):8037-8042. doi: 10.1073/pnas.1807403115. Epub 2018 Jul 16. Proc Natl Acad Sci U S A. 2018. PMID: 30012624 Free PMC article.
-
Prevalent cytidylation and uridylation of precursor miRNAs in Arabidopsis.Nat Plants. 2019 Dec;5(12):1260-1272. doi: 10.1038/s41477-019-0562-1. Epub 2019 Dec 2. Nat Plants. 2019. PMID: 31792392
-
Biomolecular condensates in plant RNA silencing: insights into formation, function, and stress responses.Plant Cell. 2024 Jan 30;36(2):227-245. doi: 10.1093/plcell/koad254. Plant Cell. 2024. PMID: 37772963 Free PMC article. Review.
-
RISC-interacting clearing 3'- 5' exoribonucleases (RICEs) degrade uridylated cleavage fragments to maintain functional RISC in Arabidopsis thaliana.Elife. 2017 May 2;6:e24466. doi: 10.7554/eLife.24466. Elife. 2017. PMID: 28463111 Free PMC article.
-
Molecular mechanism underlying the di-uridylation activity of Arabidopsis TUTase URT1.Nucleic Acids Res. 2022 Oct 14;50(18):10614-10625. doi: 10.1093/nar/gkac839. Nucleic Acids Res. 2022. PMID: 36177876 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

