Back to the origin: reconsidering replication, transcription, epigenetics, and cell cycle control
- PMID: 23634256
- PMCID: PMC3636748
- DOI: 10.1177/1947601912474891
Back to the origin: reconsidering replication, transcription, epigenetics, and cell cycle control
Abstract
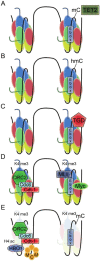
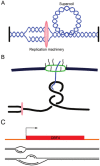
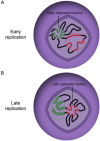
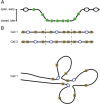
In bacteria, replication is a carefully orchestrated event that unfolds the same way for each bacterium and each cell division. The process of DNA replication in bacteria optimizes cell growth and coordinates high levels of simultaneous replication and transcription. In metazoans, the organization of replication is more enigmatic. The lack of a specific sequence that defines origins of replication has, until recently, severely limited our ability to define the organizing principles of DNA replication. This question is of particular importance as emerging data suggest that replication stress is an important contributor to inherited genetic damage and the genomic instability in tumors. We consider here the replication program in several different organisms including recent genome-wide analyses of replication origins in humans. We review recent studies on the role of cytosine methylation in replication origins, the role of transcriptional looping and gene gating in DNA replication, and the role of chromatin's 3-dimensional structure in DNA replication. We use these new findings to consider several questions surrounding DNA replication in metazoans: How are origins selected? What is the relationship between replication and transcription? How do checkpoints inhibit origin firing? Why are there early and late firing origins? We then discuss whether oncogenes promote cancer through a role in DNA replication and whether errors in DNA replication are important contributors to the genomic alterations and gene fusion events observed in cancer. We conclude with some important areas for future experimentation.
Keywords: checkpoints; epigenetics; origin; replication; replicon; transcription.
Conflict of interest statement
Figures
Similar articles
-
Replication origins and timing of temporal replication in budding yeast: how to solve the conundrum?Curr Genomics. 2010 May;11(3):199-211. doi: 10.2174/138920210791110942. Curr Genomics. 2010. PMID: 21037857 Free PMC article.
-
Genome-wide estimation of firing efficiencies of origins of DNA replication from time-course copy number variation data.BMC Bioinformatics. 2010 May 13;11:247. doi: 10.1186/1471-2105-11-247. BMC Bioinformatics. 2010. PMID: 20462459 Free PMC article.
-
Open chromatin encoded in DNA sequence is the signature of 'master' replication origins in human cells.Nucleic Acids Res. 2009 Oct;37(18):6064-75. doi: 10.1093/nar/gkp631. Epub 2009 Aug 10. Nucleic Acids Res. 2009. PMID: 19671527 Free PMC article.
-
Time to be versatile: regulation of the replication timing program in budding yeast.J Mol Biol. 2013 Nov 29;425(23):4696-705. doi: 10.1016/j.jmb.2013.09.020. Epub 2013 Sep 25. J Mol Biol. 2013. PMID: 24076190 Review.
-
Behavior of replication origins in Eukaryota - spatio-temporal dynamics of licensing and firing.Cell Cycle. 2015;14(14):2251-64. doi: 10.1080/15384101.2015.1056421. Epub 2015 Jun 1. Cell Cycle. 2015. PMID: 26030591 Free PMC article. Review.
Cited by
-
Genomic location of the major ribosomal protein gene locus determines Vibrio cholerae global growth and infectivity.PLoS Genet. 2015 Apr 13;11(4):e1005156. doi: 10.1371/journal.pgen.1005156. eCollection 2015 Apr. PLoS Genet. 2015. PMID: 25875621 Free PMC article.
-
Gene duplication and deletion caused by over-replication at a fork barrier.Nat Commun. 2023 Nov 25;14(1):7730. doi: 10.1038/s41467-023-43494-7. Nat Commun. 2023. PMID: 38007544 Free PMC article.
-
High-Resolution Chrono-Transcriptome of Lactococcus lactis Reveals That It Expresses Proteins with Adapted Size and pI upon Acidification and Nutrient Starvation.Appl Environ Microbiol. 2022 May 10;88(9):e0247621. doi: 10.1128/aem.02476-21. Epub 2022 Apr 13. Appl Environ Microbiol. 2022. PMID: 35416684 Free PMC article.
-
Replication Termination: Containing Fork Fusion-Mediated Pathologies in Escherichia coli.Genes (Basel). 2016 Jul 25;7(8):40. doi: 10.3390/genes7080040. Genes (Basel). 2016. PMID: 27463728 Free PMC article. Review.
-
Dynamic regulation of histone H3K9 is linked to the switch between replication and transcription at the Dbf4 origin-promoter locus.Cell Cycle. 2016 Sep;15(17):2321-35. doi: 10.1080/15384101.2016.1201254. Epub 2016 Jun 24. Cell Cycle. 2016. PMID: 27341472 Free PMC article.
References
-
- Jacob F, Brenner S. [On the regulation of DNA synthesis in bacteria: the hypothesis of the replicon]. C R Hebd Seances Acad Sci. 1963;256:298-300 - PubMed
-
- Mechali M. Eukaryotic DNA replication origins: many choices for appropriate answers. Nat Rev Mol Cell Biol. 2010;11:728-38 - PubMed
-
- Bell SP, Dutta A. DNA replication in eukaryotic cells. Annu Rev Biochem. 2002;71:333-74 - PubMed
-
- Todorovic V, Falaschi A, Giacca M. Replication origins of mammalian chromosomes: the happy few. Front Biosci. 1999;4:D859-68 - PubMed
-
- Kohzaki H, Murakami Y. Transcription factors and DNA replication origin selection. Bioessays. 2005;27:1107-16 - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
