Immunological Genome Project and systems immunology
- PMID: 23631936
- PMCID: PMC4615706
- DOI: 10.1016/j.it.2013.03.004
Immunological Genome Project and systems immunology
Abstract
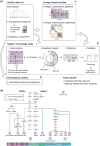
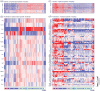
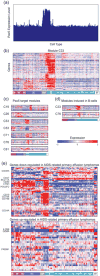
Immunological studies of single proteins in a single cell type have been complemented in recent years by larger studies, enabled by emerging high-throughput technologies. This trend has recently been exemplified by the discovery of gene networks controlling regulatory and effector αβ T cell subset development and human hematopoiesis. The Immunological Genome Project (ImmGen) aims to decipher the gene networks underpinning mouse hematopoiesis. The first phase, completed in 2012, profiled the transcriptome of 249 immune cell types. We discuss the utilities of the datasets in high-resolution mapping of the hematopoietic system. The immune transcriptome compendium has revealed unsuspected cell lineage relations and the network reconstruction has identified novel regulatory factors of hematopoiesis.
Keywords: hematopoiesis; systems immunology; transcriptional regulation.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures
Similar articles
-
Identification of transcriptional regulators in the mouse immune system.Nat Immunol. 2013 Jun;14(6):633-43. doi: 10.1038/ni.2587. Epub 2013 Apr 28. Nat Immunol. 2013. PMID: 23624555 Free PMC article.
-
Interactive Big Data Resource to Elucidate Human Immune Pathways and Diseases.Immunity. 2015 Sep 15;43(3):605-14. doi: 10.1016/j.immuni.2015.08.014. Epub 2015 Sep 8. Immunity. 2015. PMID: 26362267 Free PMC article.
-
Transcriptome profiling and sequencing of differentiated human hematopoietic stem cells reveal lineage-specific expression and alternative splicing of genes.Physiol Genomics. 2011 Oct 20;43(20):1117-34. doi: 10.1152/physiolgenomics.00099.2011. Epub 2011 Aug 9. Physiol Genomics. 2011. PMID: 21828245 Free PMC article.
-
Pleiotropic effects of prostaglandin E2 in hematopoiesis; prostaglandin E2 and other eicosanoids regulate hematopoietic stem and progenitor cell function.Prostaglandins Other Lipid Mediat. 2011 Nov;96(1-4):3-9. doi: 10.1016/j.prostaglandins.2011.06.004. Epub 2011 Jun 21. Prostaglandins Other Lipid Mediat. 2011. PMID: 21722751 Free PMC article. Review.
-
Gene regulatory networks directing myeloid and lymphoid cell fates within the immune system.Semin Immunol. 2008 Aug;20(4):228-35. doi: 10.1016/j.smim.2008.08.003. Epub 2008 Sep 3. Semin Immunol. 2008. PMID: 18771937 Review.
Cited by
-
Suture-anchored cutaneous tension induces persistent hypertrophic scarring in a novel murine model.Burns Trauma. 2024 Oct 20;12:tkae051. doi: 10.1093/burnst/tkae051. eCollection 2024. Burns Trauma. 2024. PMID: 39429643 Free PMC article.
-
A specific and portable gene expression program underlies antigen archiving by lymphatic endothelial cells.bioRxiv [Preprint]. 2024 Apr 2:2024.04.01.587647. doi: 10.1101/2024.04.01.587647. bioRxiv. 2024. PMID: 38617225 Free PMC article. Preprint.
-
Integrative temporal multi-omics reveals uncoupling of transcriptome and proteome during human T cell activation.NPJ Syst Biol Appl. 2024 Feb 28;10(1):21. doi: 10.1038/s41540-024-00346-4. NPJ Syst Biol Appl. 2024. PMID: 38418561 Free PMC article.
-
B cell receptor repertoire abnormalities in autoimmune disease.Front Immunol. 2024 Jan 31;15:1326823. doi: 10.3389/fimmu.2024.1326823. eCollection 2024. Front Immunol. 2024. PMID: 38361948 Free PMC article. Review.
-
INHBA(+) cancer-associated fibroblasts generate an immunosuppressive tumor microenvironment in ovarian cancer.NPJ Precis Oncol. 2024 Feb 15;8(1):35. doi: 10.1038/s41698-024-00523-y. NPJ Precis Oncol. 2024. PMID: 38360876 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

