High-throughput clone library analysis of the mucosa-associated microbiota reveals dysbiosis and differences between inflamed and non-inflamed regions of the intestine in inflammatory bowel disease
- PMID: 21219646
- PMCID: PMC3032643
- DOI: 10.1186/1471-2180-11-7
High-throughput clone library analysis of the mucosa-associated microbiota reveals dysbiosis and differences between inflamed and non-inflamed regions of the intestine in inflammatory bowel disease
Abstract
Background: The gut microbiota is thought to play a key role in the development of the inflammatory bowel diseases Crohn's disease (CD) and ulcerative colitis (UC). Shifts in the composition of resident bacteria have been postulated to drive the chronic inflammation seen in both diseases (the "dysbiosis" hypothesis). We therefore specifically sought to compare the mucosa-associated microbiota from both inflamed and non-inflamed sites of the colon in CD and UC patients to that from non-IBD controls and to detect disease-specific profiles.
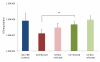
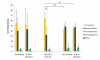
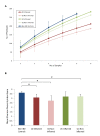
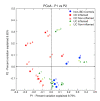
Results: Paired mucosal biopsies of inflamed and non-inflamed intestinal tissue from 6 CD (n = 12) and 6 UC (n = 12) patients were compared to biopsies from 5 healthy controls (n = 5) by in-depth sequencing of over 10,000 near full-length bacterial 16S rRNA genes. The results indicate that mucosal microbial diversity is reduced in IBD, particularly in CD, and that the species composition is disturbed. Firmicutes were reduced in IBD samples and there were concurrent increases in Bacteroidetes, and in CD only, Enterobacteriaceae. There were also significant differences in microbial community structure between inflamed and non-inflamed mucosal sites. However, these differences varied greatly between individuals, meaning there was no obvious bacterial signature that was positively associated with the inflamed gut.
Conclusions: These results may support the hypothesis that the overall dysbiosis observed in inflammatory bowel disease patients relative to non-IBD controls might to some extent be a result of the disturbed gut environment rather than the direct cause of disease. Nonetheless, the observed shifts in microbiota composition may be important factors in disease maintenance and severity.
Figures

Similar articles
-
The bacteriology of biopsies differs between newly diagnosed, untreated, Crohn's disease and ulcerative colitis patients.J Med Microbiol. 2006 Aug;55(Pt 8):1141-1149. doi: 10.1099/jmm.0.46498-0. J Med Microbiol. 2006. PMID: 16849736
-
Analysis of endoscopic brush samples identified mucosa-associated dysbiosis in inflammatory bowel disease.J Gastroenterol. 2018 Jan;53(1):95-106. doi: 10.1007/s00535-017-1384-4. Epub 2017 Aug 29. J Gastroenterol. 2018. PMID: 28852861
-
The Roles of Inflammation, Nutrient Availability and the Commensal Microbiota in Enteric Pathogen Infection.Microbiol Spectr. 2015 Jun;3(3). doi: 10.1128/microbiolspec.MBP-0008-2014. Microbiol Spectr. 2015. PMID: 26185088
-
Development, validation and implementation of an in vitro model for the study of metabolic and immune function in normal and inflamed human colonic epithelium.Dan Med J. 2015 Jan;62(1):B4973. Dan Med J. 2015. PMID: 25557335 Review.
-
Mucosal barrier, bacteria and inflammatory bowel disease: possibilities for therapy.Dig Dis. 2014;32(4):475-83. doi: 10.1159/000358156. Epub 2014 Jun 23. Dig Dis. 2014. PMID: 24969297 Review.
Cited by
-
Indigo naturalis as a potential drug in the treatment of ulcerative colitis: a comprehensive review of current evidence.Pharm Biol. 2024 Dec;62(1):818-832. doi: 10.1080/13880209.2024.2415652. Epub 2024 Oct 30. Pharm Biol. 2024. PMID: 39475104 Free PMC article. Review.
-
Co-Administration of Proton Pump Inhibitors May Negatively Affect the Outcome in Inflammatory Bowel Disease Treated with Vedolizumab.Biomedicines. 2024 Jan 11;12(1):158. doi: 10.3390/biomedicines12010158. Biomedicines. 2024. PMID: 38255263 Free PMC article.
-
Exercise attenuates PCB-induced changes in the mouse gut microbiome.Environ Health Perspect. 2013 Jun;121(6):725-30. doi: 10.1289/ehp.1306534. Epub 2013 Apr 26. Environ Health Perspect. 2013. PMID: 23632211 Free PMC article.
-
The cystic fibrosis airway microbiome.Cold Spring Harb Perspect Med. 2013 Mar 1;3(3):a009738. doi: 10.1101/cshperspect.a009738. Cold Spring Harb Perspect Med. 2013. PMID: 23457293 Free PMC article. Review.
-
Gut microbiota dysbiosis as an inflammaging condition that regulates obesity-related retinopathy and nephropathy.Front Microbiol. 2022 Nov 2;13:1040846. doi: 10.3389/fmicb.2022.1040846. eCollection 2022. Front Microbiol. 2022. PMID: 36406423 Free PMC article.
References
-
- Barrett JC, Hansoul S, Nicolae DL, Cho JH, Duerr RH, Rioux JD, Brant SR, Silverberg MS, Taylor KD, Barmada MM, Bitton A, Dassopoulos T, Datta LW, Green T, Griffiths AM, Kistner EO, Murtha MT, Regueiro MD, Rotter JI, Schumm LP, Steinhart AH, Targan SR, Xavier RJ. NIDDK IBD Genetics Consortium. Libioulle C, Sandor C, Lathrop M, Belaiche J, Dewit O, Gut I. et al.Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nat Genet. 2008;40:955–962. doi: 10.1038/ng.175. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

