iCLIP predicts the dual splicing effects of TIA-RNA interactions
- PMID: 21048981
- PMCID: PMC2964331
- DOI: 10.1371/journal.pbio.1000530
iCLIP predicts the dual splicing effects of TIA-RNA interactions
Abstract
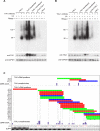
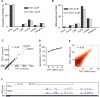
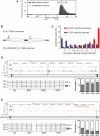
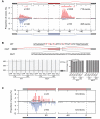
The regulation of alternative splicing involves interactions between RNA-binding proteins and pre-mRNA positions close to the splice sites. T-cell intracellular antigen 1 (TIA1) and TIA1-like 1 (TIAL1) locally enhance exon inclusion by recruiting U1 snRNP to 5' splice sites. However, effects of TIA proteins on splicing of distal exons have not yet been explored. We used UV-crosslinking and immunoprecipitation (iCLIP) to find that TIA1 and TIAL1 bind at the same positions on human RNAs. Binding downstream of 5' splice sites was used to predict the effects of TIA proteins in enhancing inclusion of proximal exons and silencing inclusion of distal exons. The predictions were validated in an unbiased manner using splice-junction microarrays, RT-PCR, and minigene constructs, which showed that TIA proteins maintain splicing fidelity and regulate alternative splicing by binding exclusively downstream of 5' splice sites. Surprisingly, TIA binding at 5' splice sites silenced distal cassette and variable-length exons without binding in proximity to the regulated alternative 3' splice sites. Using transcriptome-wide high-resolution mapping of TIA-RNA interactions we evaluated the distal splicing effects of TIA proteins. These data are consistent with a model where TIA proteins shorten the time available for definition of an alternative exon by enhancing recognition of the preceding 5' splice site. Thus, our findings indicate that changes in splicing kinetics could mediate the distal regulation of alternative splicing.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
A systematic analysis of intronic sequences downstream of 5' splice sites reveals a widespread role for U-rich motifs and TIA1/TIAL1 proteins in alternative splicing regulation.Genome Res. 2008 Aug;18(8):1247-58. doi: 10.1101/gr.073155.107. Epub 2008 May 2. Genome Res. 2008. PMID: 18456862 Free PMC article.
-
The RNA-binding protein TIA-1 is a novel mammalian splicing regulator acting through intron sequences adjacent to a 5' splice site.Mol Cell Biol. 2000 Sep;20(17):6287-99. doi: 10.1128/MCB.20.17.6287-6299.2000. Mol Cell Biol. 2000. PMID: 10938105 Free PMC article.
-
An intronic polypyrimidine-rich element downstream of the donor site modulates cystic fibrosis transmembrane conductance regulator exon 9 alternative splicing.J Biol Chem. 2004 Apr 23;279(17):16980-8. doi: 10.1074/jbc.M313439200. Epub 2004 Feb 13. J Biol Chem. 2004. PMID: 14966131
-
Hu antigen R (HuR) functions as an alternative pre-mRNA splicing regulator of Fas apoptosis-promoting receptor on exon definition.J Biol Chem. 2008 Jul 4;283(27):19077-84. doi: 10.1074/jbc.M800017200. Epub 2008 May 7. J Biol Chem. 2008. PMID: 18463097
-
TIA-1 and TIAR activate splicing of alternative exons with weak 5' splice sites followed by a U-rich stretch on their own pre-mRNAs.J Biol Chem. 2001 Nov 2;276(44):40638-46. doi: 10.1074/jbc.M105642200. Epub 2001 Aug 20. J Biol Chem. 2001. PMID: 11514562
Cited by
-
T-cell intracellular antigens in health and disease.Cell Cycle. 2015;14(13):2033-43. doi: 10.1080/15384101.2015.1053668. Cell Cycle. 2015. PMID: 26036275 Free PMC article. Review.
-
Control of alternative splicing in immune responses: many regulators, many predictions, much still to learn.Immunol Rev. 2013 May;253(1):216-36. doi: 10.1111/imr.12047. Immunol Rev. 2013. PMID: 23550649 Free PMC article. Review.
-
Nuclear matrix protein Matrin3 regulates alternative splicing and forms overlapping regulatory networks with PTB.EMBO J. 2015 Mar 4;34(5):653-68. doi: 10.15252/embj.201489852. Epub 2015 Jan 19. EMBO J. 2015. PMID: 25599992 Free PMC article.
-
Unravelling the RNA-Binding Properties of SAFB Proteins in Breast Cancer Cells.Biomed Res Int. 2015;2015:395816. doi: 10.1155/2015/395816. Epub 2015 Jul 26. Biomed Res Int. 2015. PMID: 26273616 Free PMC article.
-
Structure, dynamics and RNA binding of the multi-domain splicing factor TIA-1.Nucleic Acids Res. 2014 May;42(9):5949-66. doi: 10.1093/nar/gku193. Epub 2014 Mar 25. Nucleic Acids Res. 2014. PMID: 24682828 Free PMC article.
References
-
- Reed R. Mechanisms of fidelity in pre-mRNA splicing. Curr Opin Cell Biol. 2000;12:340–345. - PubMed
-
- Blencowe B. J. Alternative splicing: new insights from global analyses. Cell. 2006;126:37–47. - PubMed
-
- Ule J, Stefani G, Mele A, Ruggiu M, Wang X, et al. An RNA map predicting Nova-dependent splicing regulation. Nature. 2006;444:580–586. - PubMed
-
- Tian Q, Streuli M, Saito H, Schlossman S. F, Anderson P. A polyadenylate binding protein localized to the granules of cytolytic lymphocytes induces DNA fragmentation in target cells. Cell. 1991;67:629–639. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

