Multi-species transcriptome meta-analysis of the response to retinoic acid in vertebrates and comparative analysis of the effects of retinol and retinoic acid on gene expression in LMH cells
- PMID: 33653267
- PMCID: PMC7923837
- DOI: 10.1186/s12864-021-07451-2
Multi-species transcriptome meta-analysis of the response to retinoic acid in vertebrates and comparative analysis of the effects of retinol and retinoic acid on gene expression in LMH cells
Abstract
Background: Retinol (RO) and its active metabolite retinoic acid (RA) are major regulators of gene expression in vertebrates and influence various processes like organ development, cell differentiation, and immune response. To characterize a general transcriptomic response to RA-exposure in vertebrates, independent of species- and tissue-specific effects, four publicly available RNA-Seq datasets from Homo sapiens, Mus musculus, and Xenopus laevis were analyzed. To increase species and cell-type diversity we generated RNA-seq data with chicken hepatocellular carcinoma (LMH) cells. Additionally, we compared the response of LMH cells to RA and RO at different time points.
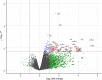
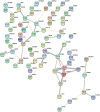
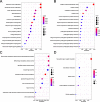
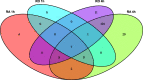
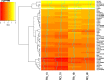
Results: By conducting a transcriptome meta-analysis, we identified three retinoic acid response core clusters (RARCCs) consisting of 27 interacting proteins, seven of which have not been associated with retinoids yet. Comparison of the transcriptional response of LMH cells to RO and RA exposure at different time points led to the identification of non-coding RNAs (ncRNAs) that are only differentially expressed (DE) during the early response.
Conclusions: We propose that these RARCCs stand on top of a common regulatory RA hierarchy among vertebrates. Based on the protein sets included in these clusters we were able to identify an RA-response cluster, a control center type cluster, and a cluster that directs cell proliferation. Concerning the comparison of the cellular response to RA and RO we conclude that ncRNAs play an underestimated role in retinoid-mediated gene regulation.
Keywords: Meta-analysis; RNA-seq; Retinoic acid; Retinoids; Retinol; Transcriptomics.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Retinoid signaling by all-trans retinoic acid and all-trans retinoyl-beta-D-glucuronide is attenuated by simultaneous exposure of human keratinocytes to retinol.J Invest Dermatol. 1999 Feb;112(2):157-64. doi: 10.1046/j.1523-1747.1999.00496.x. J Invest Dermatol. 1999. PMID: 9989790
-
In vitro antimalarial activity of retinoids and the influence of selective retinoic acid receptor antagonists.Acta Trop. 2003 Aug;87(3):345-53. doi: 10.1016/s0001-706x(03)00119-0. Acta Trop. 2003. PMID: 12875928
-
Regulation of retinoid-induced differentiation in embryonal carcinoma PCC4.aza1R cells: effects of retinoid-receptor selective ligands.Cell Growth Differ. 1996 Mar;7(3):327-37. Cell Growth Differ. 1996. PMID: 8838863
-
[Mechanism of action of retinoids in a new therapeutic approach to acute promyelocytic leukemia].Bull Cancer. 1992;79(7):697-704. Bull Cancer. 1992. PMID: 1334741 Review. French.
-
Cyclin proteolysis as a retinoid cancer prevention mechanism.Ann N Y Acad Sci. 2001 Dec;952:13-22. doi: 10.1111/j.1749-6632.2001.tb02724.x. Ann N Y Acad Sci. 2001. PMID: 11795432 Review.
Cited by
-
Proteomic Studies of Primary Acute Myeloid Leukemia Cells Derived from Patients Before and during Disease-Stabilizing Treatment Based on All-Trans Retinoic Acid and Valproic Acid.Cancers (Basel). 2021 Apr 29;13(9):2143. doi: 10.3390/cancers13092143. Cancers (Basel). 2021. PMID: 33946813 Free PMC article.
-
A Convergent Functional Genomics Analysis to Identify Biological Regulators Mediating Effects of Creatine Supplementation.Nutrients. 2021 Jul 23;13(8):2521. doi: 10.3390/nu13082521. Nutrients. 2021. PMID: 34444681 Free PMC article.
-
The retinoic acid response is a minor component of the cardiac phenotype in H9c2 myoblast differentiation.BMC Genomics. 2023 Aug 2;24(1):431. doi: 10.1186/s12864-023-09512-0. BMC Genomics. 2023. PMID: 37533008 Free PMC article.
-
Transcriptional profiling of the M. complexus in naked neck chickens suggest a direct pleiotropic effect of GDF7 on feathering and reduced hatchability.BMC Genomics. 2024 Nov 15;25(1):1092. doi: 10.1186/s12864-024-10965-0. BMC Genomics. 2024. PMID: 39548399 Free PMC article.
-
Erf Affects Commitment and Differentiation of Osteoprogenitor Cells in Cranial Sutures via the Retinoic Acid Pathway.Mol Cell Biol. 2021 Jul 23;41(8):e0014921. doi: 10.1128/MCB.00149-21. Epub 2021 Jul 23. Mol Cell Biol. 2021. PMID: 33972395 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

