This is a preprint.
Distinct patterns of SARS-CoV-2 transmission in two nearby communities in Wisconsin, USA
- PMID: 32676620
- PMCID: PMC7359545
- DOI: 10.1101/2020.07.09.20149104
Distinct patterns of SARS-CoV-2 transmission in two nearby communities in Wisconsin, USA
Update in
-
Revealing fine-scale spatiotemporal differences in SARS-CoV-2 introduction and spread.Nat Commun. 2020 Nov 3;11(1):5558. doi: 10.1038/s41467-020-19346-z. Nat Commun. 2020. PMID: 33144575 Free PMC article.
Abstract
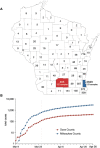
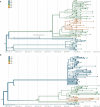
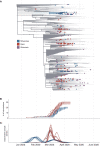
Evidence-based public health approaches that minimize the introduction and spread of new SARS-CoV-2 transmission clusters are urgently needed in the United States and other countries struggling with expanding epidemics. Here we analyze 247 full-genome SARS-CoV-2 sequences from two nearby communities in Wisconsin, USA, and find surprisingly distinct patterns of viral spread. Dane County had the 12th known introduction of SARS-CoV-2 in the United States, but this did not lead to descendant community spread. Instead, the Dane County outbreak was seeded by multiple later introductions, followed by limited community spread. In contrast, relatively few introductions in Milwaukee County led to extensive community spread. We present evidence for reduced viral spread in both counties, and limited viral transmission between counties, following the statewide Safer-at-Home public health order, which went into effect 25 March 2020. Our results suggest that early containment efforts suppressed the spread of SARS-CoV-2 within Wisconsin.
Figures

Similar articles
-
Revealing fine-scale spatiotemporal differences in SARS-CoV-2 introduction and spread.Nat Commun. 2020 Nov 3;11(1):5558. doi: 10.1038/s41467-020-19346-z. Nat Commun. 2020. PMID: 33144575 Free PMC article.
-
Genomic epidemiology of the Los Angeles COVID-19 outbreak and the early history of the B.1.43 strain in the USA.BMC Genomics. 2022 Apr 4;23(1):260. doi: 10.1186/s12864-022-08488-7. BMC Genomics. 2022. PMID: 35379194 Free PMC article.
-
A Genome Epidemiological Study of SARS-CoV-2 Introduction into Japan.mSphere. 2020 Nov 11;5(6):e00786-20. doi: 10.1128/mSphere.00786-20. mSphere. 2020. PMID: 33177213 Free PMC article.
-
Genomic surveillance identifies SARS-CoV-2 transmission patterns in local university populations, Wisconsin, USA, 2020-2022.Microb Genom. 2023 Mar;9(3):mgen000970. doi: 10.1099/mgen.0.000970. Microb Genom. 2023. PMID: 37000821 Free PMC article.
-
The Potential Role of Super Spread Events in SARS-COV-2 Pandemic; a Narrative Review.Arch Acad Emerg Med. 2020 Sep 21;8(1):e74. eCollection 2020. Arch Acad Emerg Med. 2020. PMID: 33134970 Free PMC article. Review.
References
-
- Coronavirus Locations: COVID-19 Map by County and State. (2020).
-
- Oude Munnink B. B. et al. Rapid SARS-CoV-2 whole genome sequencing for informed public health decision making in the Netherlands. Biorxiv (2020). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
