The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients
- PMID: 29302014
- PMCID: PMC6707353
- DOI: 10.1126/science.aao3290
The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients
Abstract
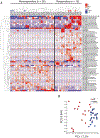
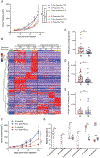
Anti-PD-1-based immunotherapy has had a major impact on cancer treatment but has only benefited a subset of patients. Among the variables that could contribute to interpatient heterogeneity is differential composition of the patients' microbiome, which has been shown to affect antitumor immunity and immunotherapy efficacy in preclinical mouse models. We analyzed baseline stool samples from metastatic melanoma patients before immunotherapy treatment, through an integration of 16S ribosomal RNA gene sequencing, metagenomic shotgun sequencing, and quantitative polymerase chain reaction for selected bacteria. A significant association was observed between commensal microbial composition and clinical response. Bacterial species more abundant in responders included Bifidobacterium longum, Collinsella aerofaciens, and Enterococcus faecium. Reconstitution of germ-free mice with fecal material from responding patients could lead to improved tumor control, augmented T cell responses, and greater efficacy of anti-PD-L1 therapy. Our results suggest that the commensal microbiome may have a mechanistic impact on antitumor immunity in human cancer patients.
Copyright © 2018, American Association for the Advancement of Science.
Figures

Comment in
-
Precision medicine using microbiota.Science. 2018 Jan 5;359(6371):32-34. doi: 10.1126/science.aar2946. Science. 2018. PMID: 29302001 No abstract available.
-
Anti-PD1 in the wonder-gut-land.Cell Res. 2018 Mar;28(3):263-264. doi: 10.1038/cr.2018.12. Epub 2018 Jan 16. Cell Res. 2018. PMID: 29336431 Free PMC article.
-
Microbiome: Gut microbiota sways response to cancer immunotherapy.Nat Rev Microbiol. 2018 Mar;16(3):121. doi: 10.1038/nrmicro.2018.12. Epub 2018 Jan 22. Nat Rev Microbiol. 2018. PMID: 29355853 No abstract available.
-
Checkpoint Checkmate: Microbiota Modulation of Cancer Immunotherapy.Clin Chem. 2018 Sep;64(9):1280-1283. doi: 10.1373/clinchem.2017.286229. Epub 2018 Apr 24. Clin Chem. 2018. PMID: 29691222 Free PMC article. No abstract available.
-
The microbiome influence.Nat Med. 2018 Dec;24(12):1782. doi: 10.1038/s41591-018-0285-2. Nat Med. 2018. PMID: 30523317 No abstract available.
-
[Gut microbiome influences efficacy of immunotherapy].Bull Cancer. 2019 Jan;106(1):10. doi: 10.1016/j.bulcan.2018.12.004. Epub 2019 Jan 11. Bull Cancer. 2019. PMID: 30642560 French. No abstract available.
Similar articles
-
Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy.Science. 2015 Nov 27;350(6264):1084-9. doi: 10.1126/science.aac4255. Epub 2015 Nov 5. Science. 2015. PMID: 26541606 Free PMC article.
-
Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients.Science. 2018 Jan 5;359(6371):97-103. doi: 10.1126/science.aan4236. Epub 2017 Nov 2. Science. 2018. PMID: 29097493 Free PMC article.
-
Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors.Science. 2018 Jan 5;359(6371):91-97. doi: 10.1126/science.aan3706. Epub 2017 Nov 2. Science. 2018. PMID: 29097494
-
Immunotherapy for the Treatment of Uveal Melanoma: Current Status and Emerging Therapies.Curr Oncol Rep. 2017 Jul;19(7):45. doi: 10.1007/s11912-017-0606-5. Curr Oncol Rep. 2017. PMID: 28508938 Review.
-
Systemic treatments for metastatic cutaneous melanoma.Cochrane Database Syst Rev. 2018 Feb 6;2(2):CD011123. doi: 10.1002/14651858.CD011123.pub2. Cochrane Database Syst Rev. 2018. PMID: 29405038 Free PMC article. Review.
Cited by
-
The Role of the Microbiome on the Pathogenesis and Treatment of Colorectal Cancer.Cancers (Basel). 2022 Nov 19;14(22):5685. doi: 10.3390/cancers14225685. Cancers (Basel). 2022. PMID: 36428777 Free PMC article. Review.
-
Human Microbiota and Breast Cancer-Is There Any Relevant Link?-A Literature Review and New Horizons Toward Personalised Medicine.Front Microbiol. 2021 Feb 25;12:584332. doi: 10.3389/fmicb.2021.584332. eCollection 2021. Front Microbiol. 2021. PMID: 33716996 Free PMC article. Review.
-
Next generation probiotics for human health: An emerging perspective.Heliyon. 2024 Aug 12;10(16):e35980. doi: 10.1016/j.heliyon.2024.e35980. eCollection 2024 Aug 30. Heliyon. 2024. PMID: 39229543 Free PMC article. Review.
-
Antibiotics impair immune checkpoint inhibitor effectiveness in Hispanic patients with non-small cell lung cancer (AB-CLICaP).Thorac Cancer. 2020 Sep;11(9):2552-2560. doi: 10.1111/1759-7714.13573. Epub 2020 Jul 24. Thorac Cancer. 2020. PMID: 32705787 Free PMC article.
-
Validation of a PD-1/CD28 chimeric switch receptor to augment CAR-T function in dogs with spontaneous B cell lymphoma.iScience. 2024 Aug 31;27(9):110863. doi: 10.1016/j.isci.2024.110863. eCollection 2024 Sep 20. iScience. 2024. PMID: 39314237 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

