Cell type-specific long-range connections of basal forebrain circuit
- PMID: 27642784
- PMCID: PMC5095704
- DOI: 10.7554/eLife.13214
Cell type-specific long-range connections of basal forebrain circuit
Erratum in
-
Correction: Cell type-specific long-range connections of basal forebrain circuit.Elife. 2016 Oct 24;5:e22475. doi: 10.7554/eLife.22475. Elife. 2016. PMID: 27775508 Free PMC article. No abstract available.
Abstract
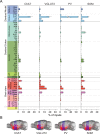
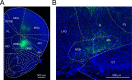
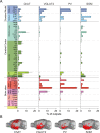
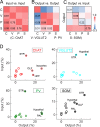
The basal forebrain (BF) plays key roles in multiple brain functions, including sleep-wake regulation, attention, and learning/memory, but the long-range connections mediating these functions remain poorly characterized. Here we performed whole-brain mapping of both inputs and outputs of four BF cell types - cholinergic, glutamatergic, and parvalbumin-positive (PV+) and somatostatin-positive (SOM+) GABAergic neurons - in the mouse brain. Using rabies virus -mediated monosynaptic retrograde tracing to label the inputs and adeno-associated virus to trace axonal projections, we identified numerous brain areas connected to the BF. The inputs to different cell types were qualitatively similar, but the output projections showed marked differences. The connections to glutamatergic and SOM+ neurons were strongly reciprocal, while those to cholinergic and PV+ neurons were more unidirectional. These results reveal the long-range wiring diagram of the BF circuit with highly convergent inputs and divergent outputs and point to both functional commonality and specialization of different BF cell types.
Keywords: anatomy; basal forebrain; cholinergic; mouse; neuroscience; parvalbumin; rabies virus; somatostatin.
Conflict of interest statement
LL: Reviewing editor, eLife. The other authors declare that no competing interests exist.
Figures

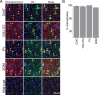

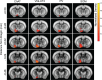
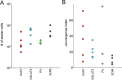
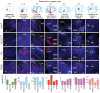



Similar articles
-
Basal forebrain subcortical projections.Brain Struct Funct. 2019 Apr;224(3):1097-1117. doi: 10.1007/s00429-018-01820-6. Epub 2019 Jan 5. Brain Struct Funct. 2019. PMID: 30612231 Free PMC article.
-
Whole-Brain Mapping of Direct Inputs to and Axonal Projections from GABAergic Neurons in the Parafacial Zone.Neurosci Bull. 2018 Jun;34(3):485-496. doi: 10.1007/s12264-018-0216-8. Epub 2018 Mar 20. Neurosci Bull. 2018. PMID: 29557546 Free PMC article.
-
Basal forebrain circuit for sleep-wake control.Nat Neurosci. 2015 Nov;18(11):1641-7. doi: 10.1038/nn.4143. Epub 2015 Oct 12. Nat Neurosci. 2015. PMID: 26457552 Free PMC article.
-
[Advances in the study of the effect of basal forebrain on sleep-wake regulation].Yao Xue Xue Bao. 2016 Aug;51(8):1196-201. Yao Xue Xue Bao. 2016. PMID: 29897712 Review. Chinese.
-
Optogenetic Dissection of the Basal Forebrain Neuromodulatory Control of Cortical Activation, Plasticity, and Cognition.J Neurosci. 2015 Oct 14;35(41):13896-903. doi: 10.1523/JNEUROSCI.2590-15.2015. J Neurosci. 2015. PMID: 26468190 Free PMC article. Review.
Cited by
-
Basal forebrain subcortical projections.Brain Struct Funct. 2019 Apr;224(3):1097-1117. doi: 10.1007/s00429-018-01820-6. Epub 2019 Jan 5. Brain Struct Funct. 2019. PMID: 30612231 Free PMC article.
-
Cell-Type-Specific Afferent Innervation of the Nucleus Accumbens Core and Shell.Front Neuroanat. 2018 Oct 16;12:84. doi: 10.3389/fnana.2018.00084. eCollection 2018. Front Neuroanat. 2018. PMID: 30459564 Free PMC article.
-
Specific Basal Forebrain-Cortical Cholinergic Circuits Coordinate Cognitive Operations.J Neurosci. 2018 Oct 31;38(44):9446-9458. doi: 10.1523/JNEUROSCI.1676-18.2018. J Neurosci. 2018. PMID: 30381436 Free PMC article. Review.
-
Neuromodulatory control of localized dendritic spiking in critical period cortex.Nature. 2019 Mar;567(7746):100-104. doi: 10.1038/s41586-019-0963-3. Epub 2019 Feb 20. Nature. 2019. PMID: 30787434 Free PMC article.
-
Development of a synchronous recording and photo-stimulating electrode in multiple brain neurons.Front Neurosci. 2023 Jun 13;17:1195095. doi: 10.3389/fnins.2023.1195095. eCollection 2023. Front Neurosci. 2023. PMID: 37383109 Free PMC article.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

