HYPER RECOMBINATION1 of the THO/TREX complex plays a role in controlling transcription of the REVERSION-TO-ETHYLENE SENSITIVITY1 gene in Arabidopsis
- PMID: 25680185
- PMCID: PMC4334170
- DOI: 10.1371/journal.pgen.1004956
HYPER RECOMBINATION1 of the THO/TREX complex plays a role in controlling transcription of the REVERSION-TO-ETHYLENE SENSITIVITY1 gene in Arabidopsis
Abstract
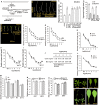
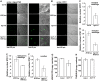
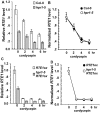
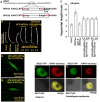
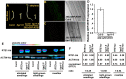
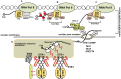
Arabidopsis REVERSION-TO-ETHYLENE SENSITIVITY1 (RTE1) represses ethylene hormone responses by promoting ethylene receptor ETHYLENE RESPONSE1 (ETR1) signaling, which negatively regulates ethylene responses. To investigate the regulation of RTE1, we performed a genetic screening for mutations that suppress ethylene insensitivity conferred by RTE1 overexpression in Arabidopsis. We isolated HYPER RECOMBINATION1 (HPR1), which is required for RTE1 overexpressor (RTE1ox) ethylene insensitivity at the seedling but not adult stage. HPR1 is a component of the THO complex, which, with other proteins, forms the TRanscription EXport (TREX) complex. In yeast, Drosophila, and humans, the THO/TREX complex is involved in transcription elongation and nucleocytoplasmic RNA export, but its role in plants is to be fully determined. We investigated how HPR1 is involved in RTE1ox ethylene insensitivity in Arabidopsis. The hpr1-5 mutation may affect nucleocytoplasmic mRNA export, as revealed by in vivo hybridization of fluorescein-labeled oligo(dT)45 with unidentified mRNA in the nucleus. The hpr1-5 mutation reduced the total and nuclear RTE1 transcript levels to a similar extent, and RTE1 transcript reduction rate was not affected by hpr1-5 with cordycepin treatment, which prematurely terminates transcription. The defect in the THO-interacting TEX1 protein of TREX but not the mRNA export factor SAC3B also reduced the total and nuclear RTE1 levels. SERINE-ARGININE-RICH (SR) proteins are involved mRNA splicing, and we found that SR protein SR33 co-localized with HPR1 in nuclear speckles, which agreed with the association of human TREX with the splicing machinery. We reveal a role for HPR1 in RTE1 expression during transcription elongation and less likely during export. Gene expression involved in ethylene signaling suppression was not reduced by the hpr1-5 mutation, which indicates selectivity of HPR1 for RTE1 expression affecting the consequent ethylene response. Thus, components of the THO/TREX complex appear to have specific roles in the transcription or export of selected genes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Similar articles
-
Association of cytochrome b5 with ETR1 ethylene receptor signaling through RTE1 in Arabidopsis.Plant J. 2014 Feb;77(4):558-67. doi: 10.1111/tpj.12401. Plant J. 2014. PMID: 24635651 Free PMC article.
-
HPR1, a component of the THO/TREX complex, plays an important role in disease resistance and senescence in Arabidopsis.Plant J. 2012 Mar;69(5):831-43. doi: 10.1111/j.1365-313X.2011.04835.x. Epub 2011 Dec 12. Plant J. 2012. PMID: 22035198
-
Arabidopsis RTE1 is essential to ethylene receptor ETR1 amino-terminal signaling independent of CTR1.Plant Physiol. 2012 Jul;159(3):1263-76. doi: 10.1104/pp.112.193979. Epub 2012 May 7. Plant Physiol. 2012. PMID: 22566492 Free PMC article.
-
Ethylene signaling: new levels of complexity and regulation.Curr Opin Plant Biol. 2008 Oct;11(5):479-85. doi: 10.1016/j.pbi.2008.06.011. Epub 2008 Aug 7. Curr Opin Plant Biol. 2008. PMID: 18692429 Free PMC article. Review.
-
New clues to understand the role of THO and other functionally related factors in mRNP biogenesis.Biochim Biophys Acta. 2012 Jun;1819(6):514-20. doi: 10.1016/j.bbagrm.2011.11.012. Epub 2011 Dec 20. Biochim Biophys Acta. 2012. PMID: 22207203 Review.
Cited by
-
Regulatory Network of Serine/Arginine-Rich (SR) Proteins: The Molecular Mechanism and Physiological Function in Plants.Int J Mol Sci. 2022 Sep 5;23(17):10147. doi: 10.3390/ijms231710147. Int J Mol Sci. 2022. PMID: 36077545 Free PMC article. Review.
-
Roles of Nuclear Pores and Nucleo-cytoplasmic Trafficking in Plant Stress Responses.Front Plant Sci. 2017 Apr 12;8:574. doi: 10.3389/fpls.2017.00574. eCollection 2017. Front Plant Sci. 2017. PMID: 28446921 Free PMC article. Review.
-
The THO/TREX Complex Active in miRNA Biogenesis Negatively Regulates Root-Associated Acid Phosphatase Activity Induced by Phosphate Starvation.Plant Physiol. 2016 Aug;171(4):2841-53. doi: 10.1104/pp.16.00680. Epub 2016 Jun 21. Plant Physiol. 2016. PMID: 27329222 Free PMC article.
-
Mechanistic Insights in Ethylene Perception and Signal Transduction.Plant Physiol. 2015 Sep;169(1):85-95. doi: 10.1104/pp.15.00845. Epub 2015 Aug 5. Plant Physiol. 2015. PMID: 26246449 Free PMC article. Review.
-
The role of TREX in gene expression and disease.Biochem J. 2016 Oct 1;473(19):2911-35. doi: 10.1042/BCJ20160010. Biochem J. 2016. PMID: 27679854 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

