Starvation-associated genome restructuring can lead to reproductive isolation in yeast
- PMID: 23894280
- PMCID: PMC3722211
- DOI: 10.1371/journal.pone.0066414
Starvation-associated genome restructuring can lead to reproductive isolation in yeast
Abstract
Knowledge of the mechanisms that lead to reproductive isolation is essential for understanding population structure and speciation. While several models have been advanced to explain post-mating reproductive isolation, experimental data supporting most are indirect. Laboratory investigations of this phenomenon are typically carried out under benign conditions, which result in low rates of genetic change unlikely to initiate reproductive isolation. Previously, we described an experimental system using the yeast Saccharomyces cerevisiae where starvation served as a proxy to any stress that decreases reproduction and/or survivorship. We showed that novel lineages with restructured genomes quickly emerged in starved populations, and that these survivors were more fit than their ancestors when re-starved. Here we show that certain yeast lineages that survive starvation have become reproductively isolated from their ancestor. We further demonstrate that reproductive isolation arises from genomic rearrangements, whose frequency in starving yeast is several orders of magnitude greater than an unstarved control. By contrast, the frequency of point mutations is less than 2-fold greater. In a particular case, we observe that a starved lineage becomes reproductively isolated as a direct result of the stress-related accumulation of a single chromosome. We recapitulate this result by demonstrating that introducing an extra copy of one or several chromosomes into naïve, i.e. unstarved, yeast significantly diminishes their fertility. This type of reproductive barrier, whether arising spontaneously or via genetic manipulation, can be removed by making a lineage euploid for the altered chromosomes. Our model provides direct genetic evidence that reproductive isolation can arise frequently in stressed populations via genome restructuring without the precondition of geographic isolation.
Conflict of interest statement
Figures
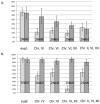
Similar articles
-
Negative epistasis: a route to intraspecific reproductive isolation in yeast?Curr Genet. 2016 Feb;62(1):25-9. doi: 10.1007/s00294-015-0505-y. Epub 2015 Jul 12. Curr Genet. 2016. PMID: 26164016 Free PMC article. Review.
-
Reciprocal gene loss following experimental whole-genome duplication causes reproductive isolation in yeast.Evolution. 2011 Apr;65(4):932-45. doi: 10.1111/j.1558-5646.2010.01171.x. Epub 2010 Nov 27. Evolution. 2011. PMID: 21114494
-
MAT heterozygosity and the second sterility barrier in the reproductive isolation of Saccharomyces species.Curr Genet. 2020 Oct;66(5):957-969. doi: 10.1007/s00294-020-01080-0. Epub 2020 Apr 30. Curr Genet. 2020. PMID: 32356035 Free PMC article.
-
A yeast living ancestor reveals the origin of genomic introgressions.Nature. 2020 Nov;587(7834):420-425. doi: 10.1038/s41586-020-2889-1. Epub 2020 Nov 11. Nature. 2020. PMID: 33177709
-
Yeast genome evolution--the origin of the species.Yeast. 2007 Nov;24(11):929-42. doi: 10.1002/yea.1515. Yeast. 2007. PMID: 17621376 Review.
Cited by
-
Genome-scale patterns in the loss of heterozygosity incidence in Saccharomyces cerevisiae.Genetics. 2022 May 5;221(1):iyac032. doi: 10.1093/genetics/iyac032. Genetics. 2022. PMID: 35212738 Free PMC article.
-
Chromosomal rearrangements as a major mechanism in the onset of reproductive isolation in Saccharomyces cerevisiae.Curr Biol. 2014 May 19;24(10):1153-9. doi: 10.1016/j.cub.2014.03.063. Epub 2014 May 8. Curr Biol. 2014. PMID: 24814147 Free PMC article.
-
Enforcement of Postzygotic Species Boundaries in the Fungal Kingdom.Microbiol Mol Biol Rev. 2022 Dec 21;86(4):e0009822. doi: 10.1128/mmbr.00098-22. Epub 2022 Sep 13. Microbiol Mol Biol Rev. 2022. PMID: 36098649 Free PMC article. Review.
References
-
- McClintock B (1984) The significance of responses of the genome to challenge. Science 226: 792. - PubMed
-
- Lin D, Gibson IB, Moore JM, Thornton PC, Leal SM, et al. (2011) Global chromosomal structural instability in a subpopulation of starving Escherichia coli cells. PLoS Genet 7: e1002223 doi:10.1371/journal.pgen.1002223 - DOI - PMC - PubMed
-
- Forche A, Abbey D, Pisithkul T, Weinzierl MA, Ringstrom T, et al. (2011) Stress alters rates and types of loss of heterozygosity in Candida albicans. MBio 2 Available: http://www.ncbi.nlm.nih.gov/pubmed/21791579. Accessed 2012 Feb 24. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

