Differential in vivo gene expression of major Leptospira proteins in resistant or susceptible animal models
- PMID: 22729538
- PMCID: PMC3416592
- DOI: 10.1128/AEM.00911-12
Differential in vivo gene expression of major Leptospira proteins in resistant or susceptible animal models
Abstract
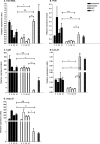
Transcripts of Leptospira 16S rRNA, FlaB, LigB, LipL21, LipL32, LipL36, LipL41, and OmpL37 were quantified in the blood of susceptible (hamsters) and resistant (mice) animal models of leptospirosis. We first validated adequate reference genes and then evaluated expression patterns in vivo compared to in vitro cultures. LipL32 expression was downregulated in vivo and differentially regulated in resistant and susceptible animals. FlaB expression was also repressed in mice but not in hamsters. In contrast, LigB and OmpL37 were upregulated in vivo. Thus, we demonstrated that a virulent strain of Leptospira differentially adapts its gene expression in the blood of infected animals.
Figures
Similar articles
-
[Genotyping and evaluation of infection dynamics in a Colombian isolate of Leptospira santarosai in hamster as an experimental model].Biomedica. 2014 Jul-Sep;34(3):460-72. doi: 10.1590/S0120-41572014000300015. Biomedica. 2014. PMID: 25504132 Spanish.
-
Evaluation of LipL32 and LigA/LigB Knockdown Mutants in Leptospira interrogans Serovar Copenhageni: Impacts to Proteome and Virulence.Front Microbiol. 2022 Feb 2;12:799012. doi: 10.3389/fmicb.2021.799012. eCollection 2021. Front Microbiol. 2022. PMID: 35185824 Free PMC article.
-
The OmpL37 surface-exposed protein is expressed by pathogenic Leptospira during infection and binds skin and vascular elastin.PLoS Negl Trop Dis. 2010 Sep 7;4(9):e815. doi: 10.1371/journal.pntd.0000815. PLoS Negl Trop Dis. 2010. PMID: 20844573 Free PMC article.
-
Leptospira as an emerging pathogen: a review of its biology, pathogenesis and host immune responses.Future Microbiol. 2010 Sep;5(9):1413-25. doi: 10.2217/fmb.10.102. Future Microbiol. 2010. PMID: 20860485 Free PMC article. Review.
-
Pathogenesis of leptospirosis: the influence of genomics.Vet Microbiol. 2011 Nov 21;153(1-2):73-81. doi: 10.1016/j.vetmic.2011.02.055. Epub 2011 Mar 5. Vet Microbiol. 2011. PMID: 21440384 Review.
Cited by
-
Draft genome of the Leptospira interrogans strains, Acegua, RCA, Prea, and Capivara, obtained from wildlife maintenance hosts and infected domestic animals.Mem Inst Oswaldo Cruz. 2016 Apr;111(4):280-3. doi: 10.1590/0074-02760160010. Mem Inst Oswaldo Cruz. 2016. PMID: 27074260 Free PMC article.
-
Leptospiral Immunoglobulin-Like Domain Proteins: Roles in Virulence and Immunity.Front Immunol. 2021 Jan 8;11:579907. doi: 10.3389/fimmu.2020.579907. eCollection 2020. Front Immunol. 2021. PMID: 33488581 Free PMC article. Review.
-
Efficient Detection of Pathogenic Leptospires Using 16S Ribosomal RNA.PLoS One. 2015 Jun 19;10(6):e0128913. doi: 10.1371/journal.pone.0128913. eCollection 2015. PLoS One. 2015. PMID: 26091292 Free PMC article.
-
Revisiting the Development of Vaccines Against Pathogenic Leptospira: Innovative Approaches, Present Challenges, and Future Perspectives.Front Immunol. 2022 Jan 3;12:760291. doi: 10.3389/fimmu.2021.760291. eCollection 2021. Front Immunol. 2022. PMID: 35046936 Free PMC article. Review.
-
The leptospiral outer membrane.Curr Top Microbiol Immunol. 2015;387:187-221. doi: 10.1007/978-3-662-45059-8_8. Curr Top Microbiol Immunol. 2015. PMID: 25388136 Free PMC article. Review.
References
-
- Adler B, Lo M, Seemann T, Murray GL. 2011. Pathogenesis of leptospirosis: the influence of genomics. Vet. Microbiol. 153: 73–81 - PubMed
-
- Andersen CL, Jensen JL, Orntoft TF. 2004. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64: 5245–5250 doi:10.1158/0008-5472.CAN-04-0496 - DOI - PubMed
-
- Carrillo-Casas EM, Hernandez-Castro R, Suarez-Guemes F, de la Pena-Moctezuma A. 2008. Selection of the internal control gene for real-time quantitative RT-PCR assays in temperature treated leptospira. Curr. Microbiol. 56: 539–546 - PubMed
-
- Chaemchuen S, Rungpragayphan S, Poovorawan Y, Patarakul K. 2011. Identification of candidate host proteins that interact with LipL32, the major outer membrane protein of pathogenic Leptospira, by random phage display peptide library. Vet. Microbiol. 153: 178–185 - PubMed
-
- Charon NW, Goldstein SF. 2002. Genetics of motility and chemotaxis of a fascinating group of bacteria: the Spirochetes. Annu. Rev. Genet. 36: 47–73 doi:10.1146/annurev.genet.36.041602.134359 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

