TDRD3, a novel Tudor domain-containing protein, localizes to cytoplasmic stress granules
- PMID: 18632687
- PMCID: PMC2536506
- DOI: 10.1093/hmg/ddn203
TDRD3, a novel Tudor domain-containing protein, localizes to cytoplasmic stress granules
Abstract
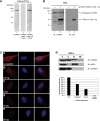
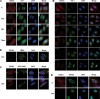
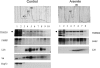
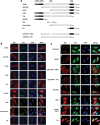
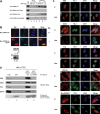
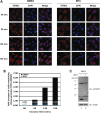
Our previous work has demonstrated that the Tudor domain of the 'survival of motor neuron' protein and the Tudor domain-containing protein 3 (TDRD3) are highly similar and that they both have the ability to interact with arginine-methylated polypeptides. TDRD3 has been identified among genes whose overexpression has a strong predictive value for poor prognosis of estrogen receptor-negative breast cancers, although its precise function remains unknown. TDRD3 is a modular protein, and in addition to its Tudor domain, it harbors a putative nucleic acid recognition motif and a ubiquitin-associated domain. We report here that TDRD3 localizes predominantly to the cytoplasm, where it co-sediments with the fragile X mental retardation protein on actively translating polyribosomes. We also demonstrate that TDRD3 accumulates into stress granules (SGs) in response to various cellular stresses. Strikingly, the Tudor domain of TDRD3 was found to be both required and sufficient for its recruitment to SGs, and the methyl-binding surface in the Tudor domain is important for this process. Pull down experiments identified five novel TDRD3 interacting partners, most of which are potentially methylated RNA-binding proteins. Our findings revealed that two of these proteins, SERPINE1 mRNA-binding protein 1 and DEAD/H box-3 (a gene often deleted in Sertoli-cell-only syndrome), are also novel constituents of cytoplasmic SGs. Taken together, we report the first characterization of TDRD3 and its functional interaction with at least two proteins implicated in human genetic diseases and present evidence supporting a role for arginine methylation in the regulation of SG dynamics.
Figures


Similar articles
-
Arginine methylation of USP9X promotes its interaction with TDRD3 and its anti-apoptotic activities in breast cancer cells.Cell Discov. 2017 Jan 3;3:16048. doi: 10.1038/celldisc.2016.48. eCollection 2017. Cell Discov. 2017. PMID: 28101374 Free PMC article.
-
Tdrd3 is a novel stress granule-associated protein interacting with the Fragile-X syndrome protein FMRP.Hum Mol Genet. 2008 Oct 15;17(20):3236-46. doi: 10.1093/hmg/ddn219. Epub 2008 Jul 28. Hum Mol Genet. 2008. PMID: 18664458
-
Structural plasticity of the TDRD3 Tudor domain probed by a fragment screening hit.FEBS J. 2018 Jun;285(11):2091-2103. doi: 10.1111/febs.14469. Epub 2018 Apr 24. FEBS J. 2018. PMID: 29645362
-
Type IA topoisomerases can be "magicians" for both DNA and RNA in all domains of life.RNA Biol. 2017 Jul 3;14(7):854-864. doi: 10.1080/15476286.2017.1330741. Epub 2017 May 23. RNA Biol. 2017. PMID: 28534707 Free PMC article. Review.
-
Tudor domain-containing proteins of Drosophila melanogaster.Dev Growth Differ. 2012 Jan;54(1):32-43. doi: 10.1111/j.1440-169x.2011.01308.x. Dev Growth Differ. 2012. PMID: 23741747 Review.
Cited by
-
A gene signature for predicting outcome in patients with basal-like breast cancer.Sci Rep. 2012;2:227. doi: 10.1038/srep00227. Epub 2012 Jan 17. Sci Rep. 2012. PMID: 22355741 Free PMC article.
-
Molecular mechanisms of stress granule assembly and disassembly.Biochim Biophys Acta Mol Cell Res. 2021 Jan;1868(1):118876. doi: 10.1016/j.bbamcr.2020.118876. Epub 2020 Sep 29. Biochim Biophys Acta Mol Cell Res. 2021. PMID: 33007331 Free PMC article. Review.
-
Molecular Evolution of DNA Topoisomerase III Beta (TOP3B) in Metazoa.J Mol Evol. 2021 Jul;89(6):384-395. doi: 10.1007/s00239-021-10011-7. Epub 2021 May 17. J Mol Evol. 2021. PMID: 33999213
-
KDM4C (GASC1) lysine demethylase is associated with mitotic chromatin and regulates chromosome segregation during mitosis.Nucleic Acids Res. 2014 Jun;42(10):6168-82. doi: 10.1093/nar/gku253. Epub 2014 Apr 11. Nucleic Acids Res. 2014. PMID: 24728997 Free PMC article.
-
Arginine methylation of USP9X promotes its interaction with TDRD3 and its anti-apoptotic activities in breast cancer cells.Cell Discov. 2017 Jan 3;3:16048. doi: 10.1038/celldisc.2016.48. eCollection 2017. Cell Discov. 2017. PMID: 28101374 Free PMC article.
References
-
- Pawson T. Specificity in signal transduction: from phosphotyrosine-SH2 domain interactions to complex cellular systems. Cell. 2004;116:191–203. - PubMed
-
- Lee D.Y., Teyssier C., Strahl B.D., Stallcup M.R. Role of protein methylation in regulation of transcription. Endocr. Rev. 2005;26:147–170. - PubMed
-
- Bedford M.T., Richard S. Arginine methylation an emerging regulator of protein function. Mol. Cell. 2005;18:263–272. - PubMed
-
- Boisvert F.M., Chenard C.A., Richard S. Protein interfaces in signaling regulated by arginine methylation. Sci. STKE. 2005;2005:re2. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

