Abstract
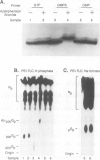
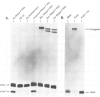
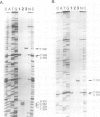
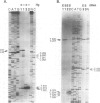
Ribonuclease P RNA is the catalytic moiety of the ribonucleoprotein enzyme that removes precursor sequences from 5'-ends of pre-tRNAs. A photoaffinity cross-linking agent was coupled to the substrate phosphate on which RNase P acts and used to map nucleotides in the vicinity of the catalytic site of this ribozyme. Mature tRNA(Phe) containing a 5'-thiophosphate was synthesized by transcription in vitro using phage T7 RNA polymerase in the presence of guanosine 5'-phosphorothioate. The photoagent (azidophenacyl) was coupled uniquely to the 5'-thiophosphate of the tRNA, the site of action by RNase P. The photoagent-containing tRNA binds to RNase P RNA and is cross-linked by UV irradiation to it at high efficiency (10-30%). Cross-linked conjugates are enzymatically inactive, consistent with the occupancy of the active site of the RNase P RNA by the tRNA. Reversal of the cross-link by phenylmercuric acetate restores activity. The sites of cross-linking in RNase P RNA were determined by primer extension. In order to identify generalities and detect idiosyncrasies, analyses were carried out using RNase P RNAs from three phylogenetically diverse organisms: Bacillus subtilis, Chromatium vinosum and Escherichia coli. In the context of a phylogenetic structure model, two regions of cross-linking are observed in all three RNAs. Two of the RNAs cross-link to a lesser extent at a third structural region and one of the RNAs is cross-linked to a small extent to a fourth region. All the sites of cross-linking between the substrate phosphate in tRNA and the RNase P RNAs are in the conserved core of the structure model, consistent with the importance of the cross-linked residues to the action of this RNA enzyme.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altman S. Ribonuclease P: an enzyme with a catalytic RNA subunit. Adv Enzymol Relat Areas Mol Biol. 1989;62:1–36. doi: 10.1002/9780470123089.ch1. [DOI] [PubMed] [Google Scholar]
- Barta A., Steiner G., Brosius J., Noller H. F., Kuechler E. Identification of a site on 23S ribosomal RNA located at the peptidyl transferase center. Proc Natl Acad Sci U S A. 1984 Jun;81(12):3607–3611. doi: 10.1073/pnas.81.12.3607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gardiner K. J., Marsh T. L., Pace N. R. Ion dependence of the Bacillus subtilis RNase P reaction. J Biol Chem. 1985 May 10;260(9):5415–5419. [PubMed] [Google Scholar]
- Gardiner K., Pace N. R. RNase P of Bacillus subtilis has a RNA component. J Biol Chem. 1980 Aug 25;255(16):7507–7509. [PubMed] [Google Scholar]
- Guerrier-Takada C., Altman S. M1 RNA with large terminal deletions retains its catalytic activity. Cell. 1986 Apr 25;45(2):177–183. doi: 10.1016/0092-8674(86)90381-8. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Gardiner K., Marsh T., Pace N., Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983 Dec;35(3 Pt 2):849–857. doi: 10.1016/0092-8674(83)90117-4. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Lumelsky N., Altman S. Specific interactions in RNA enzyme-substrate complexes. Science. 1989 Dec 22;246(4937):1578–1584. doi: 10.1126/science.2480641. [DOI] [PubMed] [Google Scholar]
- Hagenbüchle O., Santer M., Steitz J. A., Mans R. J. Conservation of the primary structure at the 3' end of 18S rRNA from eucaryotic cells. Cell. 1978 Mar;13(3):551–563. doi: 10.1016/0092-8674(78)90328-8. [DOI] [PubMed] [Google Scholar]
- Hanna M. M., Meares C. F. Synthesis of a cleavable dinucleotide photoaffinity probe of ribonucleic acid polymerase: application to trinucleotide labeling of an Escherichia coli transcription complex. Biochemistry. 1983 Jul 19;22(15):3546–3551. doi: 10.1021/bi00284a002. [DOI] [PubMed] [Google Scholar]
- Hassur S. M., Whitlock H. W., Jr UV shadowing--a new and convenient method for the location of ultraviolet-absorbing species in polyacrylamide gels. Anal Biochem. 1974 May;59(1):162–164. doi: 10.1016/0003-2697(74)90020-7. [DOI] [PubMed] [Google Scholar]
- Hixson S. H., Hixson S. S. P-Azidophenacyl bromide, a versatile photolabile bifunctional reagent. Reaction with glyceraldehyde-3-phosphate dehydrogenase. Biochemistry. 1975 Sep 23;14(19):4251–4254. doi: 10.1021/bi00690a016. [DOI] [PubMed] [Google Scholar]
- Inoue T., Cech T. R. Secondary structure of the circular form of the Tetrahymena rRNA intervening sequence: a technique for RNA structure analysis using chemical probes and reverse transcriptase. Proc Natl Acad Sci U S A. 1985 Feb;82(3):648–652. doi: 10.1073/pnas.82.3.648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- James B. D., Olsen G. J., Liu J. S., Pace N. R. The secondary structure of ribonuclease P RNA, the catalytic element of a ribonucleoprotein enzyme. Cell. 1988 Jan 15;52(1):19–26. doi: 10.1016/0092-8674(88)90527-2. [DOI] [PubMed] [Google Scholar]
- Kim S. H., Quigley G. J., Suddath F. L., McPherson A., Sneden D., Kim J. J., Weinzierl J., Rich A. Three-dimensional structure of yeast phenylalanine transfer RNA: folding of the polynucleotide chain. Science. 1973 Jan 19;179(4070):285–288. doi: 10.1126/science.179.4070.285. [DOI] [PubMed] [Google Scholar]
- Lane D. J., Pace B., Olsen G. J., Stahl D. A., Sogin M. L., Pace N. R. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci U S A. 1985 Oct;82(20):6955–6959. doi: 10.1073/pnas.82.20.6955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murray A. W., Atkinson M. R. Adenosine 5'-phosphorothioate. A nucleotide analog that is a substrate, competitive inhibitor, or regulator of some enzymes that interact with adenosine 5'-phosphate. Biochemistry. 1968 Nov;7(11):4023–4029. doi: 10.1021/bi00851a032. [DOI] [PubMed] [Google Scholar]
- Neumann H., Smith R. A. Cleavage of the disulfide bonds of cystine and oxidized glutathione by phosphorothioate. Arch Biochem Biophys. 1967 Nov;122(2):354–361. doi: 10.1016/0003-9861(67)90205-6. [DOI] [PubMed] [Google Scholar]
- Pace N. R., Smith D. K., Olsen G. J., James B. D. Phylogenetic comparative analysis and the secondary structure of ribonuclease P RNA--a review. Gene. 1989 Oct 15;82(1):65–75. doi: 10.1016/0378-1119(89)90031-0. [DOI] [PubMed] [Google Scholar]
- Pace N. R., Smith D. Ribonuclease P: function and variation. J Biol Chem. 1990 Mar 5;265(7):3587–3590. [PubMed] [Google Scholar]
- Reich C., Gardiner K. J., Olsen G. J., Pace B., Marsh T. L., Pace N. R. The RNA component of the Bacillus subtilis RNase P. Sequence, activity, and partial secondary structure. J Biol Chem. 1986 Jun 15;261(17):7888–7893. [PubMed] [Google Scholar]
- Reich C., Olsen G. J., Pace B., Pace N. R. Role of the protein moiety of ribonuclease P, a ribonucleoprotein enzyme. Science. 1988 Jan 8;239(4836):178–181. doi: 10.1126/science.3122322. [DOI] [PubMed] [Google Scholar]
- Sampson J. R., DiRenzo A. B., Behlen L. S., Uhlenbeck O. C. Nucleotides in yeast tRNAPhe required for the specific recognition by its cognate synthetase. Science. 1989 Mar 10;243(4896):1363–1366. doi: 10.1126/science.2646717. [DOI] [PubMed] [Google Scholar]
- Sampson J. R., Uhlenbeck O. C. Biochemical and physical characterization of an unmodified yeast phenylalanine transfer RNA transcribed in vitro. Proc Natl Acad Sci U S A. 1988 Feb;85(4):1033–1037. doi: 10.1073/pnas.85.4.1033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger H. L., Klotz G., Riesner D., Gross H. J., Kleinschmidt A. K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci U S A. 1976 Nov;73(11):3852–3856. doi: 10.1073/pnas.73.11.3852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stark B. C., Kole R., Bowman E. J., Altman S. Ribonuclease P: an enzyme with an essential RNA component. Proc Natl Acad Sci U S A. 1978 Aug;75(8):3717–3721. doi: 10.1073/pnas.75.8.3717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waugh D. S., Green C. J., Pace N. R. The design and catalytic properties of a simplified ribonuclease P RNA. Science. 1989 Jun 30;244(4912):1569–1571. doi: 10.1126/science.2472671. [DOI] [PubMed] [Google Scholar]
- Wower J., Hixson S. S., Zimmermann R. A. Labeling the peptidyltransferase center of the Escherichia coli ribosome with photoreactive tRNA(Phe) derivatives containing azidoadenosine at the 3' end of the acceptor arm: a model of the tRNA-ribosome complex. Proc Natl Acad Sci U S A. 1989 Jul;86(14):5232–5236. doi: 10.1073/pnas.86.14.5232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Youvan D. C., Hearst J. E. Sequencing psoralen photochemically reactive sites in Escherichia coli 16 S rRNA. Anal Biochem. 1982 Jan 1;119(1):86–89. doi: 10.1016/0003-2697(82)90669-8. [DOI] [PubMed] [Google Scholar]