Abstract
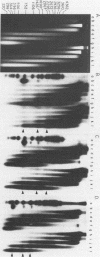
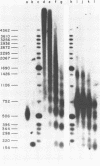
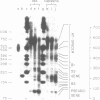
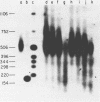
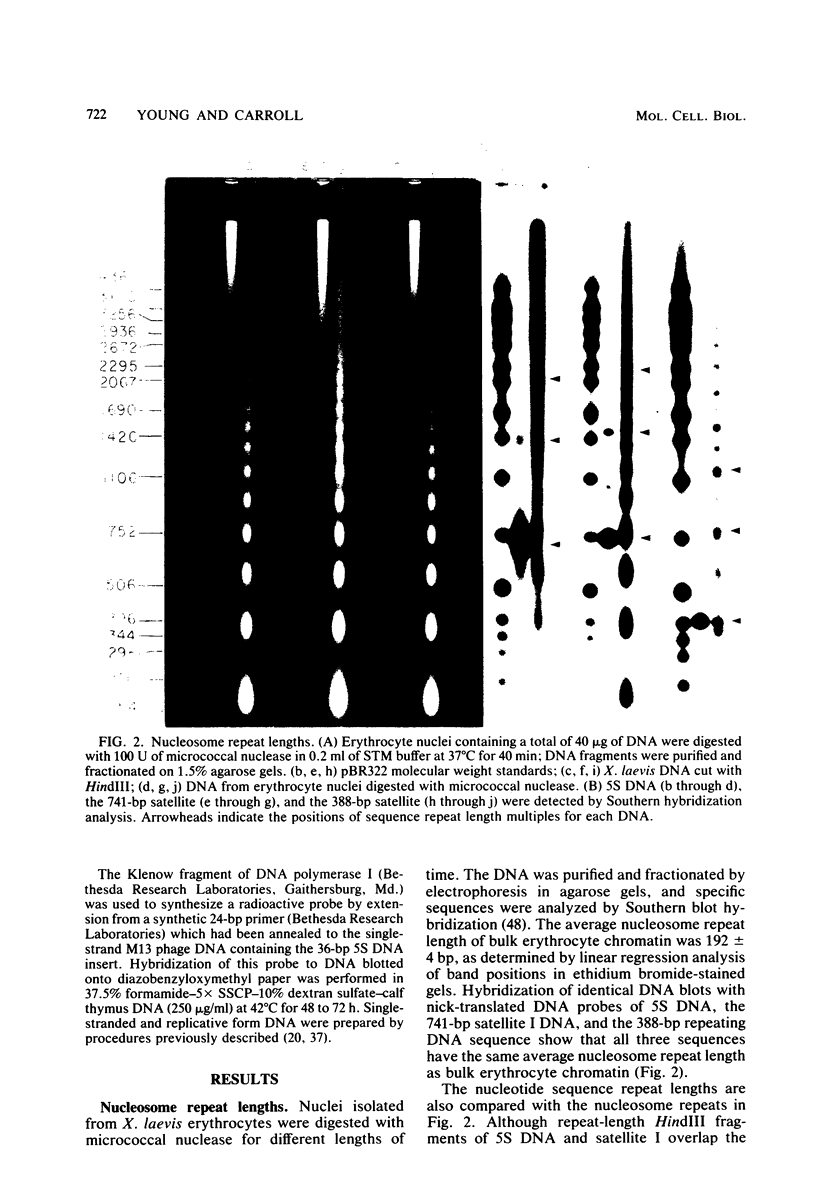
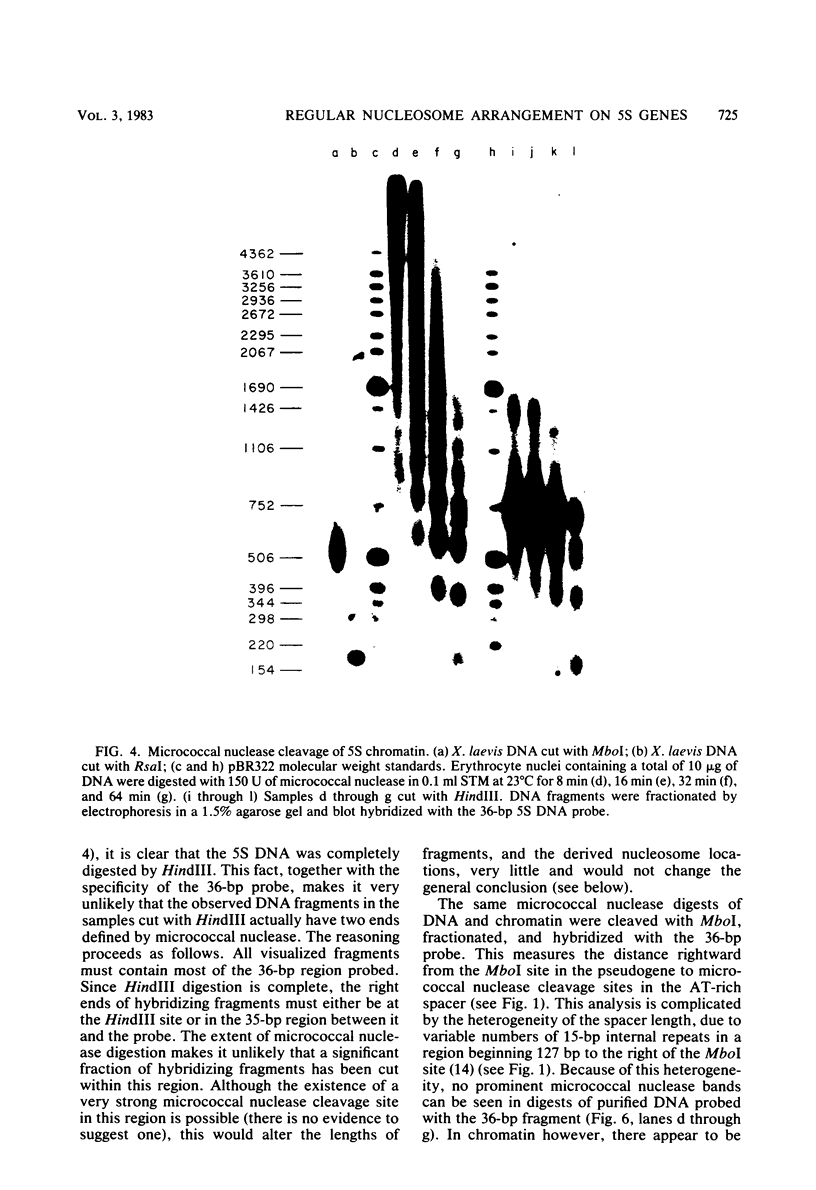
The chromatin structure of the oocyte-type 5S RNA genes in Xenopus laevis was investigated. Blot hybridization analysis of DNA from micrococcal nuclease digests of erythrocyte nuclei showed that 5S DNA has the same average nucleosome repeat length, 192 +/- 4 base pairs, as two Xenopus satellite DNAs and bulk erythrocyte chromatin. The positions of nuclease-sensitive regions in the 5S DNA repeats of purified DNA and chromatin from erythrocytes were mapped by using an indirect end-labeling technique. Although most of the sites cleaved in purified DNA were also cleaved in chromatin, the patterns of intensities were strikingly different in the two cases. In 5S chromatin, three nuclease-sensitive regions were spaced approximately a nucleosome length apart, suggesting a single, regular arrangement of nucleosomes on most of the 5S DNA repeats. The observed nucleosome locations are discussed with respect to nucleotide sequences known to be important for expression of 5S RNA. Because the preferred locations appear to be reestablished in each repeating unit, despite spacer length heterogeneity, we suggest that the regular chromatin structure reflects the presence of a sequence-specific DNA-binding component on inactive 5S RNA genes.
Full text
PDF








Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baer B. W., Kornberg R. D. Random location of nucleosomes on genes for 5 S rRNA. J Biol Chem. 1979 Oct 10;254(19):9678–9681. [PubMed] [Google Scholar]
- Birkenmeier E. H., Brown D. D., Jordan E. A nuclear extract of Xenopus laevis oocytes that accurately transcribes 5S RNA genes. Cell. 1978 Nov;15(3):1077–1086. doi: 10.1016/0092-8674(78)90291-x. [DOI] [PubMed] [Google Scholar]
- Bogenhagen D. F., Sakonju S., Brown D. D. A control region in the center of the 5S RNA gene directs specific initiation of transcription: II. The 3' border of the region. Cell. 1980 Jan;19(1):27–35. doi: 10.1016/0092-8674(80)90385-2. [DOI] [PubMed] [Google Scholar]
- Bogenhagen D. F., Wormington W. M., Brown D. D. Stable transcription complexes of Xenopus 5S RNA genes: a means to maintain the differentiated state. Cell. 1982 Feb;28(2):413–421. doi: 10.1016/0092-8674(82)90359-2. [DOI] [PubMed] [Google Scholar]
- Brown D. D., Gurdon J. B. Cloned single repeating units of 5S DNA direct accurate transcription of 5S RNA when injected into Xenopus oocytes. Proc Natl Acad Sci U S A. 1978 Jun;75(6):2849–2853. doi: 10.1073/pnas.75.6.2849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown D. D., Gurdon J. B. High-fidelity transcription of 5S DNA injected into Xenopus oocytes. Proc Natl Acad Sci U S A. 1977 May;74(5):2064–2068. doi: 10.1073/pnas.74.5.2064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown D. D., Sugimoto K. 5 S DNAs of Xenopus laevis and Xenopus mulleri: evolution of a gene family. J Mol Biol. 1973 Aug 15;78(3):397–415. doi: 10.1016/0022-2836(73)90464-6. [DOI] [PubMed] [Google Scholar]
- Brown D. D., Weber C. S. Gene linkage by RNA-DNA hybridization. I. Unique DNA sequences homologous to 4 s RNA, 5 s RNA and ribosomal RNA. J Mol Biol. 1968 Jun 28;34(3):661–680. doi: 10.1016/0022-2836(68)90188-5. [DOI] [PubMed] [Google Scholar]
- Brown D. D., Wensink P. C., Jordan E. Purification and some characteristics of 5S DNA from Xenopus laevis. Proc Natl Acad Sci U S A. 1971 Dec;68(12):3175–3179. doi: 10.1073/pnas.68.12.3175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bryan P. N., Hofstetter H., Birnstiel M. L. Nucleosome arrangement on tRNA genes of Xenopus laevis. Cell. 1981 Dec;27(3 Pt 2):459–466. doi: 10.1016/0092-8674(81)90387-1. [DOI] [PubMed] [Google Scholar]
- Carroll D., Brown D. D. Repeating units of Xenopus laevis oocyte-type 5S DNA are heterogeneous in length. Cell. 1976 Apr;7(4):467–475. doi: 10.1016/0092-8674(76)90198-7. [DOI] [PubMed] [Google Scholar]
- Dingwall C., Lomonossoff G. P., Laskey R. A. High sequence specificity of micrococcal nuclease. Nucleic Acids Res. 1981 Jun 25;9(12):2659–2673. doi: 10.1093/nar/9.12.2659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engelke D. R., Ng S. Y., Shastry B. S., Roeder R. G. Specific interaction of a purified transcription factor with an internal control region of 5S RNA genes. Cell. 1980 Mar;19(3):717–728. doi: 10.1016/s0092-8674(80)80048-1. [DOI] [PubMed] [Google Scholar]
- Fedoroff N. V., Brown D. D. The nucleotide sequence of oocyte 5S DNA in Xenopus laevis. I. The AT-rich spacer. Cell. 1978 Apr;13(4):701–716. doi: 10.1016/0092-8674(78)90220-9. [DOI] [PubMed] [Google Scholar]
- Fittler F., Zachau H. G. Subunit structure of alpha-satellite DNA containing chromatin from African green monkey cells. Nucleic Acids Res. 1979 Sep 11;7(1):1–13. doi: 10.1093/nar/7.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ford P. J., Southern E. M. Different sequences for 5S RNA in kidney cells and ovaries of Xenopus laevis. Nat New Biol. 1973 Jan 3;241(105):7–12. doi: 10.1038/newbio241007a0. [DOI] [PubMed] [Google Scholar]
- Gottesfeld J. M., Bloomer L. S. Nonrandom alignment of nucleosomes on 5S RNA genes of X. laevis. Cell. 1980 Oct;21(3):751–760. doi: 10.1016/0092-8674(80)90438-9. [DOI] [PubMed] [Google Scholar]
- Gottesfeld J. M. Organization of 5S genes in chromatin of Xenopus laevis. Nucleic Acids Res. 1980 Feb 25;8(4):905–922. [PMC free article] [PubMed] [Google Scholar]
- Gurdon J. B., Brown D. D. The transcription of 5 S DNA injected into Xenopus oocytes. Dev Biol. 1978 Dec;67(2):346–356. doi: 10.1016/0012-1606(78)90205-1. [DOI] [PubMed] [Google Scholar]
- Heidecker G., Messing J., Gronenborn B. A versatile primer for DNA sequencing in the M13mp2 cloning system. Gene. 1980 Jun;10(1):69–73. doi: 10.1016/0378-1119(80)90145-6. [DOI] [PubMed] [Google Scholar]
- Hewish D. R., Burgoyne L. A. Chromatin sub-structure. The digestion of chromatin DNA at regularly spaced sites by a nuclear deoxyribonuclease. Biochem Biophys Res Commun. 1973 May 15;52(2):504–510. doi: 10.1016/0006-291x(73)90740-7. [DOI] [PubMed] [Google Scholar]
- Humphries S. E., Young D., Carroll D. Chromatin structure of the 5S ribonucleic acid genes of Xenopus laevis. Biochemistry. 1979 Jul 24;18(15):3223–3231. doi: 10.1021/bi00582a006. [DOI] [PubMed] [Google Scholar]
- Hörz W., Altenburger W. Sequence specific cleavage of DNA by micrococcal nuclease. Nucleic Acids Res. 1981 Jun 25;9(12):2643–2658. doi: 10.1093/nar/9.12.2643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Igo-Kemenes T., Hörz W., Zachau H. G. Chromatin. Annu Rev Biochem. 1982;51:89–121. doi: 10.1146/annurev.bi.51.070182.000513. [DOI] [PubMed] [Google Scholar]
- Jacq C., Miller J. R., Brownlee G. G. A pseudogene structure in 5S DNA of Xenopus laevis. Cell. 1977 Sep;12(1):109–120. doi: 10.1016/0092-8674(77)90189-1. [DOI] [PubMed] [Google Scholar]
- Keene M. A., Elgin S. C. Micrococcal nuclease as a probe of DNA sequence organization and chromatin structure. Cell. 1981 Nov;27(1 Pt 2):57–64. doi: 10.1016/0092-8674(81)90360-3. [DOI] [PubMed] [Google Scholar]
- Korn L. J., Birkenmeier E. H., Brown D. D. Transcription initiation of Xenopus 5S ribosomal RNA genes in vitro. Nucleic Acids Res. 1979 Oct 25;7(4):947–958. doi: 10.1093/nar/7.4.947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korn L. J., Gurdon J. B. The reactivation of developmentally inert 5S genes in somatic nuclei injected into Xenopus oocytes. Nature. 1981 Feb 5;289(5797):461–465. doi: 10.1038/289461a0. [DOI] [PubMed] [Google Scholar]
- Korn L. J. Transcription of Xenopus 5S ribosomal RNA genes. Nature. 1982 Jan 14;295(5845):101–105. doi: 10.1038/295101a0. [DOI] [PubMed] [Google Scholar]
- Kornberg R. The location of nucleosomes in chromatin: specific or statistical. Nature. 1981 Aug 13;292(5824):579–580. doi: 10.1038/292579a0. [DOI] [PubMed] [Google Scholar]
- Louis C., Schedl P., Samal B., Worcel A. Chromatin structure of the 5S RNA genes of D. melanogaster. Cell. 1980 Nov;22(2 Pt 2):387–392. doi: 10.1016/0092-8674(80)90349-9. [DOI] [PubMed] [Google Scholar]
- Maio J. J., Brown F. L., Musich P. R. Subunit structure of chromatin and the organization of eukaryotic highly repetitive DNA: recurrent periodicities and models for the evolutionary origins of repetitive DNA. J Mol Biol. 1977 Dec 15;117(3):637–655. doi: 10.1016/0022-2836(77)90062-6. [DOI] [PubMed] [Google Scholar]
- Maniatis T., Jeffrey A., Kleid D. G. Nucleotide sequence of the rightward operator of phage lambda. Proc Natl Acad Sci U S A. 1975 Mar;72(3):1184–1188. doi: 10.1073/pnas.72.3.1184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGhee J. D., Felsenfeld G. Nucleosome structure. Annu Rev Biochem. 1980;49:1115–1156. doi: 10.1146/annurev.bi.49.070180.005343. [DOI] [PubMed] [Google Scholar]
- Messing J., Crea R., Seeburg P. H. A system for shotgun DNA sequencing. Nucleic Acids Res. 1981 Jan 24;9(2):309–321. doi: 10.1093/nar/9.2.309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messing J., Gronenborn B., Müller-Hill B., Hans Hopschneider P. Filamentous coliphage M13 as a cloning vehicle: insertion of a HindII fragment of the lac regulatory region in M13 replicative form in vitro. Proc Natl Acad Sci U S A. 1977 Sep;74(9):3642–3646. doi: 10.1073/pnas.74.9.3642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller J. R., Cartwright E. M., Brownlee G. G., Fedoroff N. V., Brown D. D. The nucleotide sequence of oocyte 5S DNA in Xenopus laevis. II. The GC-rich region. Cell. 1978 Apr;13(4):717–725. doi: 10.1016/0092-8674(78)90221-0. [DOI] [PubMed] [Google Scholar]
- Nedospasov S. A., Georgiev G. P. Non-random cleavage of SV40 DNA in the compact minichromosome and free in solution by micrococcal nuclease. Biochem Biophys Res Commun. 1980 Jan 29;92(2):532–539. doi: 10.1016/0006-291x(80)90366-6. [DOI] [PubMed] [Google Scholar]
- Ng S. Y., Parker C. S., Roeder R. G. Transcription of cloned Xenopus 5S RNA genes by X. laevis RNA polymerase III in reconstituted systems. Proc Natl Acad Sci U S A. 1979 Jan;76(1):136–140. doi: 10.1073/pnas.76.1.136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker C. S., Roeder R. G. Selective and accurate transcription of the Xenopus laevis 5S RNA genes in isolated chromatin by purified RNA polymerase III. Proc Natl Acad Sci U S A. 1977 Jan;74(1):44–48. doi: 10.1073/pnas.74.1.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pelham H. R., Wormington W. M., Brown D. D. Related 5S RNA transcription factors in Xenopus oocytes and somatic cells. Proc Natl Acad Sci U S A. 1981 Mar;78(3):1760–1764. doi: 10.1073/pnas.78.3.1760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ROBERTS W. K., DEKKER C. A., RUSHIZKY G. W., KNIGHT C. A. Studies on the mechanism of action of micrococcal nuclease. 1. Degradation of thymus deoxyribonucleic acid. Biochim Biophys Acta. 1962 May 14;55:664–673. doi: 10.1016/0006-3002(62)90844-2. [DOI] [PubMed] [Google Scholar]
- Sakonju S., Bogenhagen D. F., Brown D. D. A control region in the center of the 5S RNA gene directs specific initiation of transcription: I. The 5' border of the region. Cell. 1980 Jan;19(1):13–25. doi: 10.1016/0092-8674(80)90384-0. [DOI] [PubMed] [Google Scholar]
- Sakonju S., Brown D. D., Engelke D., Ng S. Y., Shastry B. S., Roeder R. G. The binding of a transcription factor to deletion mutants of a 5S ribosomal RNA gene. Cell. 1981 Mar;23(3):665–669. doi: 10.1016/0092-8674(81)90429-3. [DOI] [PubMed] [Google Scholar]
- Segall J., Matsui T., Roeder R. G. Multiple factors are required for the accurate transcription of purified genes by RNA polymerase III. J Biol Chem. 1980 Dec 25;255(24):11986–11991. [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Thomas J. O., Thompson R. J. Variation in chromatin structure in two cell types from the same tissue: a short DNA repeat length in cerebral cortex neurons. Cell. 1977 Apr;10(4):633–640. doi: 10.1016/0092-8674(77)90096-4. [DOI] [PubMed] [Google Scholar]
- Wahl G. M., Stern M., Stark G. R. Efficient transfer of large DNA fragments from agarose gels to diazobenzyloxymethyl-paper and rapid hybridization by using dextran sulfate. Proc Natl Acad Sci U S A. 1979 Aug;76(8):3683–3687. doi: 10.1073/pnas.76.8.3683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wegnez M., Monier R., Denis H. Sequence heterogeneity of 5 S RNA in Xenopus laevis. FEBS Lett. 1972 Sep 1;25(1):13–20. doi: 10.1016/0014-5793(72)80443-5. [DOI] [PubMed] [Google Scholar]
- Weil P. A., Segall J., Harris B., Ng S. Y., Roeder R. G. Faithful transcription of eukaryotic genes by RNA polymerase III in systems reconstituted with purified DNA templates. J Biol Chem. 1979 Jul 10;254(13):6163–6173. [PubMed] [Google Scholar]
- Wu C. The 5' ends of Drosophila heat shock genes in chromatin are hypersensitive to DNase I. Nature. 1980 Aug 28;286(5776):854–860. doi: 10.1038/286854a0. [DOI] [PubMed] [Google Scholar]