Abstract
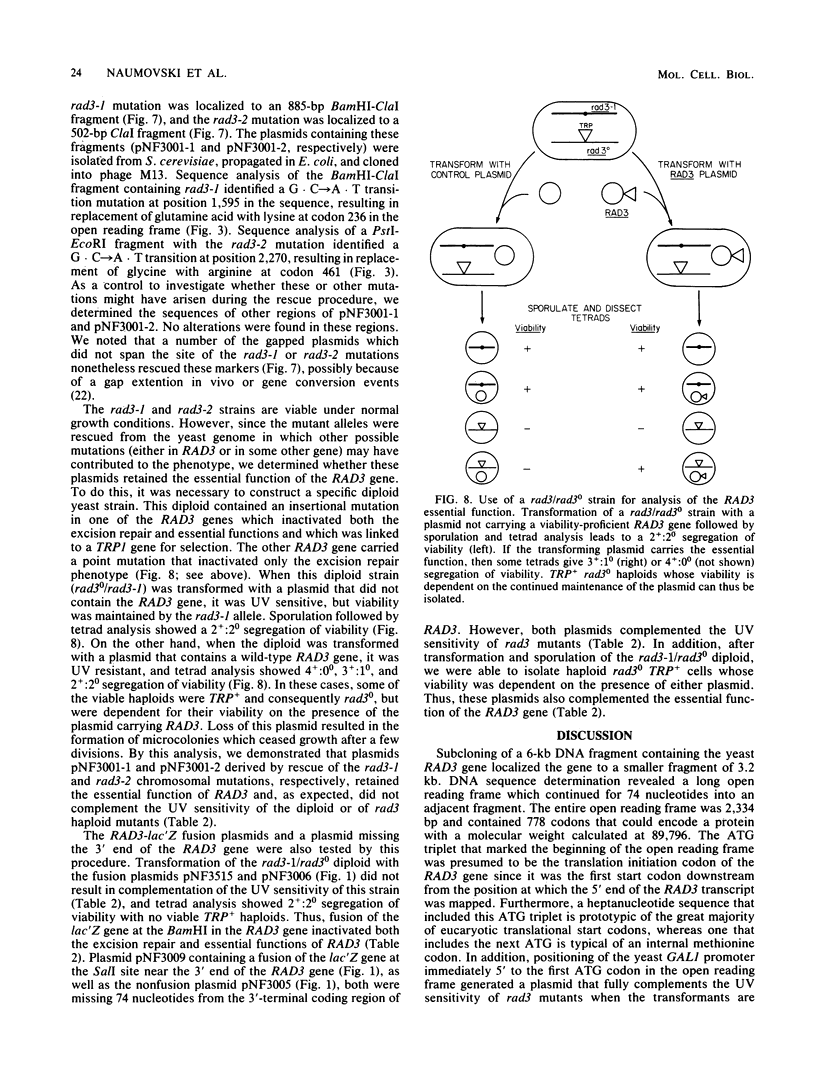
We determined the complete nucleotide sequence of the RAD3 gene of Saccharomyces cerevisiae. The coding region of the gene contained 2,334 base pairs that could encode a protein with a calculated molecular weight of 89,796. Analysis of RAD3 mRNA by Northern blots and by S1 nuclease mapping indicated that the transcript was approximately 2.5 kilobases and did not contain intervening sequences. Fusions between the RAD3 gene and the lac'Z gene of Escherichia coli were constructed and used to demonstrate that the RAD3 gene was not inducible by DNA damage caused by UV radiation or 4-nitroquinoline-1-oxide. Two UV-sensitive chromosomal mutant alleles of RAD3, rad3-1 and rad3-2, were rescued by gap repair of a centromeric plasmid, and their sequences were determined. The rad3-1 mutation changed a glutamic acid to lysine, and the rad3-2 mutation changed a glycine to arginine. Previous studies have shown that disruption of the RAD3 gene results in loss of an essential function and is associated with inviability of haploid cells. In the present experiments, plasmids carrying the rad3-1 and rad3-2 mutations were introduced into haploid cells containing a disrupted RAD3 gene. These plasmids expressed the essential function of RAD3 but not its DNA repair function. A 74-base-pair deletion at the 3' end of the RAD3 coding region or a fusion of this deletion to the E. coli lac'Z gene did not affect either function of RAD3.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Berk A. J., Sharp P. A. Sizing and mapping of early adenovirus mRNAs by gel electrophoresis of S1 endonuclease-digested hybrids. Cell. 1977 Nov;12(3):721–732. doi: 10.1016/0092-8674(77)90272-0. [DOI] [PubMed] [Google Scholar]
- Biggin M. D., Gibson T. J., Hong G. F. Buffer gradient gels and 35S label as an aid to rapid DNA sequence determination. Proc Natl Acad Sci U S A. 1983 Jul;80(13):3963–3965. doi: 10.1073/pnas.80.13.3963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casadaban M. J., Chou J., Cohen S. N. In vitro gene fusions that join an enzymatically active beta-galactosidase segment to amino-terminal fragments of exogenous proteins: Escherichia coli plasmid vectors for the detection and cloning of translational initiation signals. J Bacteriol. 1980 Aug;143(2):971–980. doi: 10.1128/jb.143.2.971-980.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Germino J., Gray J. G., Charbonneau H., Vanaman T., Bastia D. Use of gene fusions and protein-protein interaction in the isolation of a biologically active regulatory protein: the replication initiator protein of plasmid R6K. Proc Natl Acad Sci U S A. 1983 Nov;80(22):6848–6852. doi: 10.1073/pnas.80.22.6848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall M. N., Hereford L., Herskowitz I. Targeting of E. coli beta-galactosidase to the nucleus in yeast. Cell. 1984 Apr;36(4):1057–1065. doi: 10.1016/0092-8674(84)90055-2. [DOI] [PubMed] [Google Scholar]
- Higgins D. R., Prakash S., Reynolds P., Polakowska R., Weber S., Prakash L. Isolation and characterization of the RAD3 gene of Saccharomyces cerevisiae and inviability of rad3 deletion mutants. Proc Natl Acad Sci U S A. 1983 Sep;80(18):5680–5684. doi: 10.1073/pnas.80.18.5680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinnen A., Hicks J. B., Fink G. R. Transformation of yeast. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1929–1933. doi: 10.1073/pnas.75.4.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jensen R., Sprague G. F., Jr, Herskowitz I. Regulation of yeast mating-type interconversion: feedback control of HO gene expression by the mating-type locus. Proc Natl Acad Sci U S A. 1983 May;80(10):3035–3039. doi: 10.1073/pnas.80.10.3035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. Possible role of flanking nucleotides in recognition of the AUG initiator codon by eukaryotic ribosomes. Nucleic Acids Res. 1981 Oct 24;9(20):5233–5252. doi: 10.1093/nar/9.20.5233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langford C. J., Klinz F. J., Donath C., Gallwitz D. Point mutations identify the conserved, intron-contained TACTAAC box as an essential splicing signal sequence in yeast. Cell. 1984 Mar;36(3):645–653. doi: 10.1016/0092-8674(84)90344-1. [DOI] [PubMed] [Google Scholar]
- Müller-Hill B., Kania J. Lac repressor can be fused to beta-galactosidase. Nature. 1974 Jun 7;249(457):561–563. doi: 10.1038/249561a0. [DOI] [PubMed] [Google Scholar]
- Naumovski L., Friedberg E. C. A DNA repair gene required for the incision of damaged DNA is essential for viability in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1983 Aug;80(15):4818–4821. doi: 10.1073/pnas.80.15.4818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naumovski L., Friedberg E. C. Construction of plasmid vectors that facilitate subcloning and recovery of yeast and Escherichia coli DNA fragments. Gene. 1983 May-Jun;22(2-3):203–209. doi: 10.1016/0378-1119(83)90104-x. [DOI] [PubMed] [Google Scholar]
- Naumovski L., Friedberg E. C. Molecular cloning of eucaryotic genes required for excision repair of UV-irradiated DNA: isolation and partial characterization of the RAD3 gene of Saccharomyces cerevisiae. J Bacteriol. 1982 Oct;152(1):323–331. doi: 10.1128/jb.152.1.323-331.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naumovski L., Friedberg E. C. Saccharomyces cerevisiae RAD2 gene: isolation, subcloning, and partial characterization. Mol Cell Biol. 1984 Feb;4(2):290–295. doi: 10.1128/mcb.4.2.290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norrander J., Kempe T., Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983 Dec;26(1):101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- Okayama H., Berg P. High-efficiency cloning of full-length cDNA. Mol Cell Biol. 1982 Feb;2(2):161–170. doi: 10.1128/mcb.2.2.161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orr-Weaver T. L., Szostak J. W., Rothstein R. J. Genetic applications of yeast transformation with linear and gapped plasmids. Methods Enzymol. 1983;101:228–245. doi: 10.1016/0076-6879(83)01017-4. [DOI] [PubMed] [Google Scholar]
- Proudfoot N. J., Brownlee G. G. 3' non-coding region sequences in eukaryotic messenger RNA. Nature. 1976 Sep 16;263(5574):211–214. doi: 10.1038/263211a0. [DOI] [PubMed] [Google Scholar]
- Reynolds R. J., Friedberg E. C. Molecular mechanisms of pyrimidine dimer excision in Saccharomyces cerevisiae: incision of ultraviolet-irradiated deoxyribonucleic acid in vivo. J Bacteriol. 1981 May;146(2):692–704. doi: 10.1128/jb.146.2.692-704.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruby S. W., Szostak J. W., Murray A. W. Cloning regulated yeast genes from a pool of lacZ fusions. Methods Enzymol. 1983;101:253–269. doi: 10.1016/0076-6879(83)01019-8. [DOI] [PubMed] [Google Scholar]
- Rupp W. D., Sancar A., Sancar G. B. Properties and regulation of the UVRABC endonuclease. Biochimie. 1982 Aug-Sep;64(8-9):595–598. doi: 10.1016/s0300-9084(82)80094-1. [DOI] [PubMed] [Google Scholar]
- Sancar G. B., Sancar A., Rupp W. D. Sequences of the E. coli uvrC gene and protein. Nucleic Acids Res. 1984 Jun 11;12(11):4593–4608. doi: 10.1093/nar/12.11.4593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilcox D. R., Prakash L. Incision and postincision steps of pyrimidine dimer removal in excision-defective mutants of Saccharomyces cerevisiae. J Bacteriol. 1981 Nov;148(2):618–623. doi: 10.1128/jb.148.2.618-623.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang E., Friedberg E. C. Molecular cloning and nucleotide sequence analysis of the Saccharomyces cerevisiae RAD1 gene. Mol Cell Biol. 1984 Oct;4(10):2161–2169. doi: 10.1128/mcb.4.10.2161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaret K. S., Sherman F. DNA sequence required for efficient transcription termination in yeast. Cell. 1982 Mar;28(3):563–573. doi: 10.1016/0092-8674(82)90211-2. [DOI] [PubMed] [Google Scholar]