Abstract
Background
Cowpea (Vigna unguiculata) is an important crop in arid and semi-arid regions and is a good model for studying drought tolerance. MicroRNAs (miRNAs) are known to play critical roles in plant stress responses, but drought-associated miRNAs have not been identified in cowpea. In addition, it is not understood how miRNAs might contribute to different capacities of drought tolerance in different cowpea genotypes.
Results
We generated deep sequencing small RNA reads from two cowpea genotypes (CB46, drought-sensitive, and IT93K503-1, drought-tolerant) that grew under well-watered and drought stress conditions. We mapped small RNA reads to cowpea genomic sequences and identified 157 miRNA genes that belong to 89 families. Among 44 drought-associated miRNAs, 30 were upregulated in drought condition and 14 were downregulated. Although miRNA expression was in general consistent in two genotypes, we found that nine miRNAs were predominantly or exclusively expressed in one of the two genotypes and that 11 miRNAs were drought-regulated in only one genotype, but not the other.
Conclusions
These results suggest that miRNAs may play important roles in drought tolerance in cowpea and may be a key factor in determining the level of drought tolerance in different cowpea genotypes.
Background
Drought is one of the main abiotic factors that cause reduction or total loss of crop production. Because water is becoming limited for agriculture in many areas of the world, the investigation of natural mechanisms of drought tolerance is an important strategy for understanding the biological basis of response to drought stress and for selection of plants with improved drought tolerance [1,2]. Cowpea [Vigna unguiculata (L.) Walp.] is an economically important crop in semi-arid and arid tropical regions in Africa, Asia, and Central and South America, where cowpea is consumed as human food and nutritious fodder to livestock [3,4]. As a leguminous species, cowpea belongs to the same tribe (Phaseoleae) as common bean and soybean. Compared to these close relatives and most other crops, cowpea is well adapted to these regions because of its ability to fix nitrogen in poor soil and greater drought tolerance [4,5]. Therefore, cowpea is an excellent system for investigating the genetic basis of drought tolerance.
Efforts have been made to identify genetic elements that are involved in drought stress response in cowpea. For example, over a dozen genes have been shown to be associated with drought stress response through cloning and characterization of cDNAs [6-12]. In addition, ten drought tolerance quantitative trait loci (QTL) associated with tolerance in seedlings have been mapped in cowpea [13]. However, it is largely unknown how the expression of drought-associated cowpea genes or loci is regulated and how small RNAs are involved in the regulation.
MicroRNAs (miRNAs) are 20-24 nt single-stranded RNA molecules that are processed from RNA precursors that fold into stem-loop structures [14]. MiRNAs regulate gene expression of target mRNAs at the posttranscriptional level, which are recognized by nearly perfect base complementarity. Upon miRNA-target recognition, typically the target is negatively regulated via mRNA cleavage or translational repression [15]. Functional analyses have demonstrated that miRNAs are involved in a variety of developmental processes in plants [16]. In addition, miRNAs play critical roles in plant resistance to various abiotic and biotic stresses [17-19]. In particular, several approaches have been employed to study miRNAs that are involved in drought stress tolerance in plants. In one of the pioneering studies on stress-responsive miRNAs, Sunkar and Zhu [20] used small RNA cloning techniques to identify 26 novel miRNAs, among which miR393, miR397b, and miR402 were upregulated by dehydration and miR389a downregulated. Another miRNA family, miR169, was found to be downregulated by drought stress in an ABA-dependent pathway. The repression of miR169 leads to higher expression of its target gene NFYA5, which in turn enhances the drought resistance of the plant [21]. Many more miRNAs that are up- or down-regulated in drought condition were discovered by global miRNA expression profiling experiments with either microarray hybridization or small RNA deep sequencing [22-25].
Although numerous miRNAs have been identified in many plant species, including leguminous plants Medicago truncatula [26,27], soybean [28], and common bean [29], only two sequences have been reported for cowpea in the miRBase registry. Recently, 47 potential miRNAs belonging to 13 miRNA families were predicted in cowpea [30]. In another study, 18 conserved miRNAs belonging to 16 families were identified [31]. Both studies used a homology search approach to identify cowpea miRNAs that are conserved in other plants. In this study, we used Illumina deep sequencing technology to generate small RNA reads and used these reads to identify miRNAs in cowpea, especially cowpea-specific miRNAs and those associated with drought tolerance. To our knowledge, this is the first report of miRNAs identified through direct small RNA cloning in cowpea.
Despite inherent drought tolerance, cowpea varieties display significantly different levels of drought tolerance [32-34]. The study and comparison of plant genotypes differing in sensitivity to drought is a promising approach to discover natural tolerance mechanisms [35]. In order to gain insight into the role of miRNAs in tolerance to drought, we used two representative cowpea genotypes: California Blackeye No. 46 (CB46) and IT93K503-1. The drought-sensitive CB46 is the most widely grown blackeye-type cultivar in the United States and was developed at the University of California, Davis [36]. IT93K503-1 is a drought-tolerant breeding line developed by the International Institute of Tropical Agriculture (IITA) in Ibadan, Nigeria. We grew these two genotypes in well-watered and drought stress conditions and used leaves from the vegetative stage to construct four small RNA libraries. Using small RNA reads from these libraries, we identified 157 candidate miRNAs. Comparison of the expression pattern of miRNAs among libraries indicates that some miRNAs display different levels of expression in different genotypes, and thus may be a key factor to their different levels of drought tolerance.
Results
Identification of miRNAs in cowpea
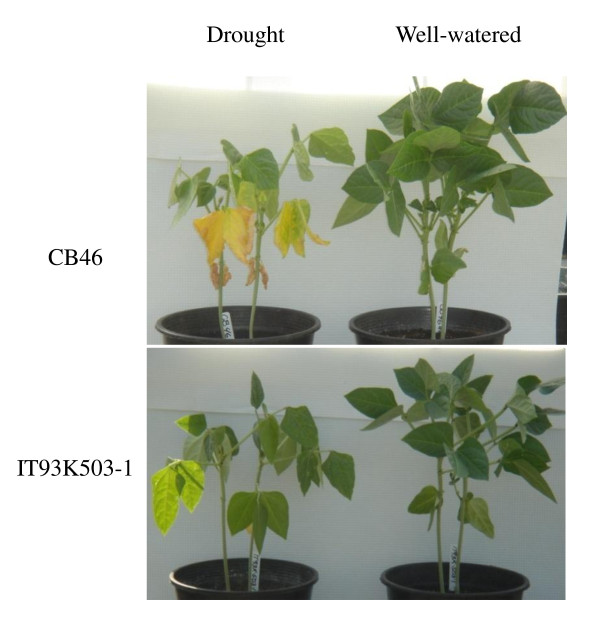
In order to study the role of miRNAs in drought tolerance, we grew cowpea plants (CB46 and IT93K503-1) in green house under well-watered and drought stress conditions. Drought stress was applied to 30-day-old plants. After 10 to 15 days of moderate drought stress (ψw = -1.5 MPa), the two genotypes showed apparent differences in drought tolerance. While IT93K503-1 plants continued to grow relatively well, CB46 plants displayed severe drought stress symptoms such as chlorotic leaves (Figure 1).
Figure 1.
Different drought tolerance of two cowpea genotypes. After treated with moderate drought stress (ψw = -1.5 MPa), IT93K503-1 plants continued to grow relatively well, but CB46 plants showed apparent symptoms of drought stress (chlorotic leaves).
We constructed four small RNA libraries (2 genotypes × 2 growth conditions) and obtained on average 6.9 million (range: 6.5 - 7.3 million) clean small RNA reads from each library (deep-sequencing data have been deposited into the NCBI/GEO database with accession number GSE26402). The average number of unique reads per library is 4.3 million (range: 3.9 - 4.6 million). Using the procedure and criteria described in the materials and methods section, we mapped unique small RNA reads to a cowpea EST assembly, BAC end sequences and methylation filtration sequences, GSS sequences in dbGSS, and a draft cowpea genome assembly, and predicted 14, 78, 6, and 125 miRNA precursors, respectively. These four sets of putative miRNA precursors were then compared with each other to remove redundancy, and we obtained 157 candidate miRNA genes (for detailed information, see Additional file 1). Based on similarity of mature miRNA sequences, these miRNA genes were clustered into 89 families. Whereas 27 families (93 miRNAs) have match to miRNAs from other plants in the miRBase (release 16) [37], 62 families (64 miRNAs) appear to be cowpea-specific. Using a cowpea EST assembly, we have also identified putative target protein-coding genes for 112 (71%) miRNAs.
Genotype-specific expression of miRNAs
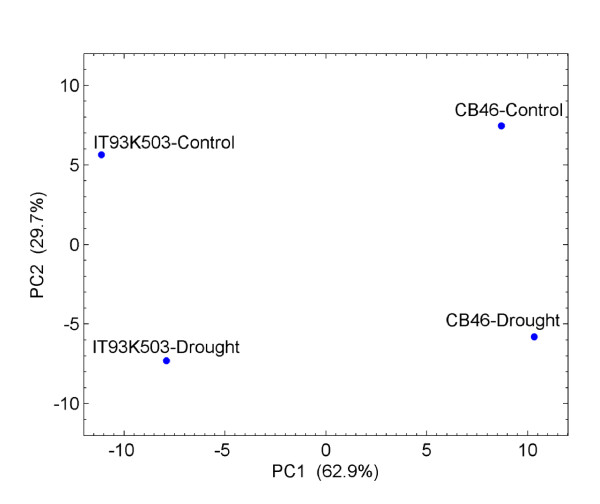
Because small RNA libraries were sequenced to great depth, counts of mature miRNAs can be used to evaluate their relative expression levels in different genotypes and growth conditions. We first applied Principal Component Analysis (PCA) to the log2 normalized counts (transcripts per ten million, TPTM) of 91 unique mature miRNAs that had combined expression of at least 50 TPTM in four libraries. As shown in Figure 2, the first two components account for over 93% of variation in the data set, with the first component accounting for 63%. The first component (PC1) separates two samples of one genotype from two samples of the other genotype, indicating genotype is the main factor that determines miRNA expression levels. Indeed, nine miRNAs account for 75% of variation in PC1 and they show clear genotype-specific expressions (Table 1, for predicted hairpin structures and mapping of small RNA reads to precursors, see Additional files 2 and 3). Whereas two miRNAs (vun_cand014 and vun_cand054) are predominantly expressed in CB46, the other seven miRNAs are exclusively or predominantly expressed in IT93K503-1 plants. The expression pattern of IT93K503-1 specific miRNA, vun_cand058, was confirmed with northern blot assay (Figure 3b).
Figure 2.
Principal component analysis (PCA) of log2 miRNA normalized counts of two cowpea genotypes in two growth conditions.
Table 1.
MiRNAs that showed genotype-specific expression
| Normalized Expression Level (TPTM)* | ||||||
|---|---|---|---|---|---|---|
| Family | Mature miRNA | IT93K503-Control | IT93K503-Drought | CB46-Control | CB46- Drought |
Putative Target |
| vun_cand058 | UUAAGCAGAAUGAUCAAAUUG | 942 | 1546 | 3 | 0 | hydroxyproline-rich glycoprotein |
| vun_cand048 | UGGUCUCUAAACUUUAGAAAUGAA | 746 | 263 | 0 | 2 | |
| vun_cand036 | UCAGAGGAAACAACACUUGUAC | 59 | 23 | 0 | 0 | |
| vun_cand045 | CGUGCUGAGAAAGUUGCUUCU | 52 | 79 | 14 | 5 | VTC2 (vitamin c defective 2) |
| vun_cand053 | GUAAUUGAGUUAAAAGGACUAUAU | 43 | 6 | 0 | 2 | cellulose synthase/transferase |
| vun_cand052 | CGAGAGCCACUCGCCUAAGCGA | 34 | 55 | 0 | 0 | |
| vun_cand055 | CCACUGUAGUAGCUCUCGCUCA | 30 | 40 | 0 | 0 | |
| vun_cand054 | AGCAAGUUGAGGAUGGAGCUU | 9 | 48 | 231 | 252 | CKA1 (casein kinase alpha 1) |
| vun_cand014 | UUCGGGAGUGAGAGCCAGUGA | 3 | 0 | 56 | 5 | UBP18 (ubiquitin-specific protease 18) |
*TPTM: transcripts per ten million
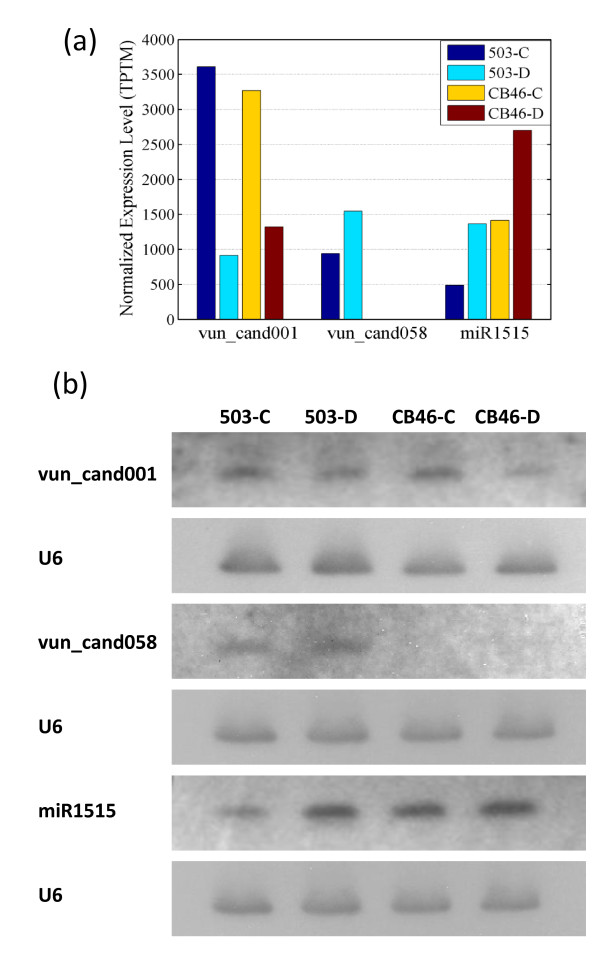
Figure 3.
Expression of selected miRNAs in two cowpea genotypes under two growth conditions. Vun_cand001 and vun_cand058 are two cowpea-specific miRNAs, and miR1515 is a conserved miRNA that is also found in other plants. A. Expression level based on normalized miRNA counts (transcripts per ten million, TPTM). B. Northern blots with mature miRNAs. U6 snRNA was used to show equal loading of total RNA in all lanes.
Because perfect matches were required when small RNA reads were mapped to cowpea sequences for miRNA prediction, genotype-specific expression of miRNAs could be caused by inter-genotype single nucleotide polymorphisms (SNPs) in mature miRNAs. To address this possibility, we re-mapped clean small RNA reads from each library to the precursors of nine miRNAs in Table 1, allowing up to one mismatch. The normalized counts of these mature miRNAs were essentially unchanged (data not shown). Therefore, genotype-specific expression of these miRNAs was genuine and was not an artifact of the reads mapping process.
Drought-associated miRNAs
To identify drought-associated miRNAs, we tested for differential expression of miRNAs in drought-stressed and corresponding control samples in each genotype using the statistical method developed by Audic and Claverie [38]. We used the following criteria to identify drought-associated miRNAs: (1) adjusted p-value was less than 0.01 in at least one of the two comparisons; (2) normalized counts (TPTM) was at least 100 in one of the four libraries; (3) log2 ratio of normalized counts between drought and control libraries was greater than 1 or less than -1 in one of the two genotypes. For differential expression analysis, we considered only unique mature miRNAs as they are the active form of the miRNA and in some cases, identical mature miRNA can be generated from two or more homologous miRNA genes. We found 44 drought-associated unique mature miRNAs that belong to 28 families (Additional file 4). The direction of statistically significant change was the same in both genotypes for all 44 miRNAs, indicating that miRNA gene expression in IT93K503-1 and CB46 had similar overall response to drought stress. Whereas thirty of 44 miRNAs were upregulated in the drought-stressed condition, fourteen were downregulated in one or both genotypes.
Among 44 drought-associated miRNAs, the expression of 22 miRNAs (17 families) in drought condition changed at least two-fold compared to the control in both genotypes (Additional file 4). Some of these miRNA families have been found to be associated with drought stress in previous studies, including miR156 and miR166 [39], miR159 [40], miR167 [24], miR169 [22], miR171 [25,41], miR395 [40], miR396 [24,39], and miR482 [29]. Most of the predicted targets encode transcription factors (Additional file 4). Other miRNA families, miR162, miR164, miR319, miR403, miR828, and four cowpea-specific miRNA (vun_cand001, vun_cand010, vun_cand041, and vun_cand057) were found to be associated with drought stress for the first time (northern blot confirmation of vun_cand001 was shown in Figure 3b).
We also found that 12 miRNAs showed at least two-fold change only in IT93K503-1 (Table 2), and 10 only in CB46 (Table 3). Although statistical tests indicated that some of these miRNAs (e.g. miR1515 which was validated by northern blot as shown in Figure 3b) were up- or down-regulated under drought stress in both genotypes without having two-fold change, 11 miRNAs were clearly regulated in only one genotype. Whereas miR160a, miR160b, miR171e, vun_cand015, vun_cand033, and vun_cand048 were significantly regulated by drought stress in IT93K503-1 plants only, miR171b, miR171d, miR2111b, miR390b, and miR393 were regulated only in CB46.
Table 2.
MiRNAs that displayed at least two fold change under drought stress only in IT93K503-1
| Normalized Expression Level (TPTM)* | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| miRNA ID | 503-C | 503-D | CB46-C | CB46-D | Log2(503-D/503-C) | Log2(CB46-D/CB46-C) | Adjusted p-value (503-D vs. 503-C) | Adjusted p-value (CB46-D vs. CB46-C) | Putative target |
| miR1515 | 489 | 1366 | 1415 | 2700 | 1.48 | 0.93 | 5e-63 | 4e-60 | |
| miR160a | 437 | 877 | 1048 | 1102 | 1.01 | 0.07 | 3e-21 | 1 | ARF10 |
| miR160b | 13 | 244 | 139 | 178 | 4.18 | 0.35 | 9e-36 | 1 | ARF10 |
| miR167b | 1649 | 4488 | 7539 | 13930 | 1.44 | 0.89 | 2e-201 | 1e-288 | ARF8 |
| miR171e | 25 | 137 | 59 | 43 | 2.44 | -0.45 | 4e-11 | 1 | |
| miR319b | 1019 | 2638 | 685 | 1215 | 1.37 | 0.83 | 5e-109 | 2e-21 | transferase family protein |
| miR390a | 2141 | 7242 | 3586 | 5308 | 1.76 | 0.57 | 0 | 3e-49 | leucine-rich repeat transmembrane protein kinase |
| vun_cand009 | 582 | 1519 | 1025 | 1903 | 1.38 | 0.89 | 4e-63 | 4e-39 | |
| vun_cand015 | 62 | 297 | 61 | 96 | 2.25 | 0.66 | 2e-23 | 1 | RPL6A |
| vun_cand020 | 4462 | 12349 | 6115 | 10535 | 1.47 | 0.78 | 0 | 6e-177 | pentatricopeptide repeat-containing protein |
| vun_cand033 | 478 | 172 | 96 | 113 | -1.48 | 0.23 | 0 | 1 | |
| vun_cand048 | 746 | 263 | 0 | 2 | -1.51 | N/A | 0 | 1 | |
*TPTM: transcripts per ten million; 503-D and 503-C: IT93K503-1 under drought and control condition, respectively; CB46-D and CB46-C: CB46 under drought and control condition, respectively.
Table 3.
MiRNAs that displayed at least two fold change under drought stress only in CB46
| Normalized Expression Level (TPTM)* | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| miRNA ID | 503-C | 503-D | CB46-C | CB46-D | Log2(503-D/503-C) | Log2(CB46-D/CB46-C) | Adjusted p-value (503-D vs. 503-C) | Adjusted p-value (CB46-D vs. CB46-C) | Putative target |
| miR166a | 7796 | 12734 | 7334 | 21341 | 0.71 | 1.54 | 4e-177 | 0 | REV |
| miR171b | 406 | 441 | 195 | 397 | 0.12 | 1.02 | 1 | 3e-09 | mRNA guanylyltransferase |
| miR171d | 58 | 55 | 85 | 215 | -0.08 | 1.33 | 1 | 2e-07 | |
| miR2111a | 458 | 678 | 191 | 1105 | 0.57 | 2.53 | 5e-05 | 1e-106 | kelch repeat-containing F-box protein |
| miR2111b | 241 | 340 | 107 | 333 | 0.50 | 1.64 | 0.48 | 4e-17 | kelch repeat-containing F-box protein |
| miR390b | 52 | 33 | 110 | 34 | -0.65 | -1.69 | 1 | 8e-05 | serine/threonine protein kinase |
| miR393 | 392 | 400 | 1120 | 258 | 0.03 | -2.12 | 1 | 0 | AFB3; AFB2 |
| miR396b | 4517 | 2958 | 5165 | 2411 | -0.61 | -1.10 | 0 | 0 | growth-regulating factor 3 |
| miR482 | 19518 | 10487 | 49339 | 13487 | -0.90 | -1.87 | 0 | 0 | ARA12; serine-type endopeptidase |
| vun_cand030 | 848 | 431 | 531 | 196 | -0.98 | -1.44 | 0 | 0 | zinc finger family protein |
*TPTM: transcripts per ten million; 503-D and 503-C: IT93K503-1 under drought and control condition, respectively; CB46-D and CB46-C: CB46 under drought and control condition, respectively.
Discussion
Regulation of gene expression through sequence-specific interaction between miRNAs and their target mRNAs offers an accurate and inheritable mechanism for plants to respond to environment stimuli [18]. Due to water limitations, drought is a major stress that limits the geographic distribution and yield of many crops. Therefore, extensive effort has been made for discovering genetic elements and mechanisms of drought tolerance, including the discovery of drought-associated miRNAs. As an important drought-tolerant crop in semi-arid and arid areas, cowpea offers a good system for the study of drought tolerance. Here we used deep sequencing of small RNA libraries from two cowpea genotypes and identified 157 miRNAs. By comparing the expression level of miRNAs in drought-stressed sample to control sample, we also identified 30 miRNAs that were upregulated in drought condition and 14 downregulated. This list of drought-associated miRNAs includes miRNA families that were known to be associated with drought in other plant species, indicating that they are involved in conserved drought response pathways. Some miRNA families, including some cowpea-specific miRNAs, were found to be associated with drought for the first time, suggesting that they may be involved in lineage- or species-specific stress response pathways and functions.
We predicted target genes for 32 out of 44 drought-associated miRNAs. The predicted target mRNAs encode proteins of diverse function, most of them being transcription factors (Additional file 4). For most of the conserved miRNAs, it is expected that their targets are also conserved. For example, our results showed that miR156 was upregulated in response to drought in cowpea. MiR156 has been known to be responsive to abiotic stresses and targets SPB transcription factors in Arabidopsis, maize, rice and wheat [24,39,41-43]. This miRNA is also involved in the regulation of development during vegetative phase change [42], indicating that reprogramming of development is a crucial step in plants to cope with drought stress. Another miRNA, miR169, was downregulated in both cowpea genotypes. In Arabidopsis, miR169 was downregulated and its target, a Nuclear Factor Y transcription factor NFYA5, was induced by drought stress [21]. MiR169 most likely functions in a similar way in cowpea to enhance drought tolerance by inducing the expression of NFYA5 orthologs.
The cowpea genotypes studied in this work have different abilities of drought tolerance. Because the two genotypes are highly similar to each other in their genetic composition, their phenotypic variations such as drought tolerance are most likely caused by changes in regulatory processes, rather than changes in proteins [44]. Due to their different geographical origins, the two genotypes are adapted to the particular environmental conditions in their natural habitats. It is thus expected to find constitutive differences, which could be related to metabolism, use of energetic resources, mobilization of biomass, structure of radical system, wax deposition in leaves, membrane stability or density of stomata, among other characteristics. We found that nine miRNAs were predominantly or exclusively expressed in only one genotype, regardless of the treatments. On the other hand, 11 miRNAs were found to be differentially expressed under drought stress in one genotype, but not the other. Changes in miRNA expression are expected to cause changes in the expression of target genes between the two genotypes.
Among miRNAs that had genotype-specific regulation, miR160a and miR160b were upregulated in response to drought in the tolerant, but not in the sensitive cultivar (Table 2). Their putative targets are members of the family of Auxin Response Factors (ARFs). ARFs are key elements in regulation of physiological and morphological mechanisms mediated by auxins that may contribute to stress adaptation [45]. Moreover, negative regulation of ARF10 by miR160 was demonstrated to be critical during seed germination in Arabidopsis thaliana through the crosstalk between auxin and ABA-dependent pathways [46]. On the other hand, two members of the miR2111 family were upregulated by drought in the sensitive, but not in the tolerant cultivar (Table 3). Their putative targets are Kelch repeat-containing F-box proteins that belong to a large family with members known to be involved in response to biotic and abiotic stresses [47]. Furthermore, F-box proteins containing Kelch repeats have been found to be responsive to drought in chickpea, a close relative of cowpea [48]. This suggests that genotype-specific regulation of miRNAs might be part of the reason why some cowpea genotypes have stronger drought tolerance than others.
Among the new miRNA candidates that were identified in this study, ten were regulated by drought stress and target genes were predicted for five of them. For instance, vun_cand030 was downregulated by drought and putatively targets a zinc finger protein. Zinc finger proteins are known to be involved in a variety of functions in development and stress response [49]. Moreover, vun_cand015 was upregulated by drought in the tolerant cultivar and putatively targets a basic-helix-loop-helix (bHLH) transcription factor. These proteins have roles in response to abiotic stresses, such as iron deficiency [50], freezing, and salt stress [51]. This suggests these new miRNAs may be indeed an integral component of drought response in cowpea.
For many miRNAs there were more than one target predicted. The possibility of a miRNA to have multiple targets is commonly observed. To confirm these predicted targets, we need to perform detailed analysis of cleavage of mRNA targets at the miRNA recognition site by experimental approaches, such as RACE and degradome analysis [52-54]. Once we validate the targets of drought-associated miRNAs, we will be in a better position to link the expression changes of miRNAs and their targets to differences of drought tolerance in cowpea.
Because we do not have the complete cowpea genome sequence, some miRNA genes were not identified, even though they had significant expression in our small RNA libraries. To find out how many miRNA families have been missed, we mapped unique small RNA reads to plant miRNA precursors in the miRBase, allowing up to 2 mismatches. Although we did not miss a large number of miRNAs, we did find that miR2118, miR2911, and miR529 had significant expression in our libraries (Additional file 5). The latter two were also induced by drought stress. MiR529 was identified as drought-associated miRNA in rice [25]. However, contrary to the pattern that we found in cowpea, it was downregulated under drought stress in rice. It is not clear whether it was caused by different sampling time or tissue, or species-specific stress response mechanisms.
Like protein coding genes, many miRNA families possess more than one miRNA gene and miRNA genes from the same family may have either identical or similar but different mature miRNA sequences. During evolutionary process, homologous miRNA genes may functionally diverge from each other. In the set of miRNAs that we identified in cowpea, members from miR166 and miR167 families showed clear evidence for functional diversification. While one member miRNA gene (miR166a, miR167b) was induced by drought stress, another miRNA from the same family (miR166b, miR167a) was significantly downregulated (Additional file 4).
Conclusions
Using deep sequencing technology, we identified 157 miRNAs in cowpea, including 44 miRNAs that are drought-associated. By comparing mature miRNA counts in different genotypes and growth conditions, we found 9 miRNAs that were almost exclusively expressed in only one genotype and 11 miRNAs that were regulated by drought stress in one genotype, but not the other. Our study demonstrated that deep sequencing of small RNAs is a cost-effective way for miRNA discovery and expression analysis. Compared to the homology search method, deep sequencing allowed the detection of species-specific miRNAs and digital expression analysis. Our findings demonstrate that expression patterns of some miRNAs may be very different even between two genotypes of the same species. Further characterization of the targets of drought-associated miRNAs will help understand the details of response and tolerance to drought in cowpea.
Methods
Plant materials
CB46 and IT93K503-1 plants were grown in a greenhouse at the University of California Riverside campus in Spring 2009. The temperature was 35°C during the day and 18°C at night with no artificial control of day length. Four seeds were germinated in 2 gallon-pots filled with steam-sterilized UC Riverside soil mix UCMIX-3 and thinned to two plants per pot two weeks after planting. Three replicate pots per treatment were arranged in a completely randomized block design. When plants were 30 days old, corresponding to late vegetative stage, deficit irrigation treatments were applied by withholding watering on the stressed pots while controlled pots were water daily to soil capacity. Third leaf water potential was monitored using a pressure chamber (Cornallis, PMS instruments, USA) [55] as the indicator of the stress level. Fresh leaves (second from apex) of three replicates were sampled and frozen in liquid nitrogen from control plants (well watered, ψw = -0.5 MPa) and moderately stressed plants (ψw = -1.5 MPa) for RNA extraction.
Small RNA library construction and sequencing
Total RNA was extracted with the TRIzol reagent (Invitrogen) according to the manufacturer's instructions. Small RNA libraries were constructed from cowpea leaves using the procedure used by Sunkar and Zhu [20] with minor modifications [56]. Briefly, for each treatment/genotype group, equal amount of total RNA was pooled from three replicates to generate ~700 μg of RNA. Pooled total RNA was resolved in a 15% denaturing polyacrylamide gel and the 20-30 nt small RNA fraction was extracted and eluted. A preadenylated adaptor (linker 1, IDT) was ligated to the 3' end of small RNAs with the use of T4 RNA ligase. Ligation products were then gel purified and subsequently ligated to an RNA adaptor at the 5'end. After ligation and purification, the products were used as template for RT-PCR. After synthesis and purification, the PCR products were quantified and sequenced using an Illumina Genome Analyzer.
miRNA identification
Only small RNA reads that passed the Illumina quality control and contained clear adaptor sequences were considered good reads for further processing. After adaptor sequence was trimmed, clean small RNA reads of 18nt or more were combined into unique sequences. Reads that match known plant repeats, rRNAs, tRNAs, snRNAs, and snoRNAs were removed. Unique small RNA reads were mapped to four genomic sequence resources with SOAP2 [57]: cowpea EST assembly available in HarvEST:Cowpea [58] (http://harvest.ucr.edu, version 1.17, 18,745 sequences, but we excluded those appear to be protein-coding genes), a combination of 260,642 cowpea gene-space random shotgun sequences [59] and 30,527 BAC end sequences (obtained from M.-C. Luo, UC Davis, http://phymap.ucdavis.edu:8080/cowpea), 54,123 cowpea Genome Survey Sequences (GSS) from dbGSS of GenBank http://www.ncbi.nlm.nih.gov/dbGSS/, and a draft cowpea genome assembly from 63× coverage Illumina pair-ended reads (296,868 contigs with total length of ~186 MB, available at http://www.harvest-blast.org). Perfect match was required.
We used the updated annotation criteria for plant miRNAs [60] and built an in-house pipeline for miRNA prediction. Unique reads with a redundancy of at least 10 copies are used as anchor sequences. With one end anchored at 10 bp from the mapped position, DNA segments of 100 - 300 bp that cover each anchor sequence were sampled with 20 bp as step size. Secondary structure of each segment was predicted with UNAFold [61]. We then examined the structures and only those met the following criteria were considered genuine miRNA candidates: (1) free energy is lower than or equal to -35 kcal/mol; (2) number of mismatches between putative miRNA and miRNA* is 4 or less; (3) number of asymmetrical bulges in the stem region is not greater than 1 and the size of each asymmetrical bulge is 2 or less; (4) strand bias - small RNA reads that map to the positive strand of the hairpin DNA segment account for at least 80% of all mapped reads; (5) precise cleavage - reads that map to the miRNA and miRNA* regions (defined as miRNA or miRNA* plus 2nt on 5' and 3' ends) account for at least 75% of all reads that map to the precursor. If two or more candidate hairpins were predicted from the same region, we compared these hairpins and chose a hairpin that has highest putative mature miRNA expression, lowest free energy, or shortest length.
In order to classify miRNAs into families, all predicted mature miRNAs were compared with themselves using the ssearch35 program in the FASTA package (version 3.5) [62]. Using a single-linkage algorithm, mature miRNAs with up to two mismatches were included in same clusters. Mature miRNAs were then compared with the mature miRNAs in the miRBase (Release 16) [37] using ssearch35. If a member in a cowpea miRNA cluster had a match (allowing up to two mismatches) in the miRBase, the family number of the known miRNA was assigned to the cluster, otherwise the cluster was annotated as a new family.
miRNA Target prediction
Mature miRNA sequences were used as query to search the cowpea EST assembly for potential target sites using miRanda [63]. The alignments between miRNAs and potential targets were extracted from the miRanda output and scored using a position-dependent, mispair penalty system [64-66]. Briefly, miRNA-target duplexes were divided into two regions: a core region that includes positions 2-13 from the 5' end of the miRNA, and a general region that contains other positions. In the general region, a penalty score of 1 was given to a mismatch or a single-nucleotide bulge or gap, and 0.5 to a G:U pair. Scores were doubled in the core region. A match was considered positive if the alignment between miRNA and target meets two conditions: (1) the penalty score is 4 or less; (2) total number of bulges and gaps is less than 2.
Principal component analysis
Counts of each mature miRNA were first normalized to transcripts per ten million (TPTM) according to the total number of clean small RNA reads in each of the four libraries. MiRNAs with combined expression of at least 50 TPTM were chosen for principal component analysis (PCA). We used the log2 values of miRNA normalized counts to build an expression matrix and used the princomp function in MATLAB (MathWorks Inc., Natick, MA) for PCA.
Statistical test for differential expression of miRNAs
Because deep sequencing of small RNAs provides a random sampling of mature miRNAs in the original small RNA pools, counts of miRNAs can be modeled by a Poisson distribution. We applied an established method [38,67] to calculate the p-value for differential expression of miRNAs between a drought-stressed sample and a control sample. The first step was to calculate a conditional probability using the formula:
Where N1 is total number of clean reads in the control library, N2 is total number of clean reads in the drought-stressed library, x is number of a mature miRNA in the control library, and y is number of the same mature miRNA in the drought-stressed library. A two-tailed p-value for differential expression was then calculated as p = 2q, where q was the accumulated probability:
Due to the x↔y symmetry of p(y|x), if q was greater than 0.5, p-value could be calculated as p = 2*(1-q). Bonferroni method was used to adjust p-values for multiple comparisons.
Northern blot analysis
~40 μg of total RNA were resolved in 15% denaturing polyacrylamide gels and transferred to neutral nylon membranes (Hybond NX). The RNA was transferred and fixed to the membranes by chemical cross-linking [68] and then hybridized to probes complementary to mature miRNA sequences at 38°C, overnight. After hybridization, the blots were washed twice, 5 minutes each at 38°C with washing solution (2X SSC, 0.1% SDS) and exposed to X-ray film to reveal the signals. Results obtained in Northern blot assays were verified in three replicated samples.
Authors' contributions
BEB-F, J-KZ, TJC and RL conceived the study. BEB-F, ZW, NND, JDE, and PAR carried out the experiments. BEB-F, LG, J-KZ, and RL analyzed the data, LG contributed new analysis tools, RL, BEB-F, TJC, and J-KZ wrote the paper. All authors read and approved the final manuscript.
Supplementary Material
MiRNAs that were identified in cowpea. Detailed information of the predicted cowpea miRNAs and their targets.
Predicted hairpin structures of nine genotype-specific miRNAs. Predicted structures of nine genotype-specific miRNAs with mature miRNAs marked in green.
Mapping of small RNA reads from four libraries to the precursors of nine genotype-specific miRNAs. Each figure shows the precursor sequence, predicted hairpin structure, and how each unique read was mapped to the precursor.
Drought-associated miRNAs in cowpea. Detailed information of drought-associated miRNAs and their targets.
Other conserved miRNAs that were expressed in cowpea. Three conserved miRNAs and their expression values in two cowpea genotypes under two growth conditions.
Contributor Information
Blanca E Barrera-Figueroa, Email: bbarrera@unpa.edu.mx.
Lei Gao, Email: leigao@ucr.edu.
Ndeye N Diop, Email: nn.diop@gmail.com.
Zhigang Wu, Email: zhigang.wu@email.ucr.edu.
Jeffrey D Ehlers, Email: jeff.ehlers@ucr.edu.
Philip A Roberts, Email: philip.roberts@ucr.edu.
Timothy J Close, Email: timothy.close@ucr.edu.
Jian-Kang Zhu, Email: zhu132@purdue.edu.
Renyi Liu, Email: renyi.liu@ucr.edu.
Acknowledgements and Funding
This work was supported by the UC Riverside Initial Complement Fund and a USDA Hatch Fund (CA-R-BPS-7754H) to RL, UCR Agricultural Experiment Station funds to TJC, NIH grants R01GM070795 and R01GM059138 to J-KZ, and UC-MEXUS and CONACYT-Mexico fellowships to BEB-F.
References
- Tuberosa R, Salvi S. Genomics-based approaches to improve drought tolerance of crops. Trends Plant Sci. 2006;11:405–412. doi: 10.1016/j.tplants.2006.06.003. [DOI] [PubMed] [Google Scholar]
- Ashraf M. Inducing drought tolerance in plants: Recent advances. Biotechnol Adv. 2010;28:169–183. doi: 10.1016/j.biotechadv.2009.11.005. [DOI] [PubMed] [Google Scholar]
- Singh BB, Ajeigbe HA, Tarawali SA, Fernandez-Rivera S, Abubakar M. Improving the production and utilization of cowpea as food and fodder. Field Crops Res. 2003;84:169–177. doi: 10.1016/S0378-4290(03)00148-5. [DOI] [Google Scholar]
- Ehlers JD, Hall AE. Cowpea (Vigna unguiculata L Walp) Field Crops Res. 1997;53:187–204. doi: 10.1016/S0378-4290(97)00031-2. [DOI] [Google Scholar]
- Sanginga N, Lyasse O, Singh BB. Phosphorus use efficiency and nitrogen balance of cowpea breeding lines in a low P soil of the derived savanna zone in West Africa. Plant Soil. 2000;220:119–128. doi: 10.1023/A:1004785720047. [DOI] [Google Scholar]
- Iuchi S, YamaguchiShinozaki K, Urao T, Terao T, Shinozaki K. Novel drought-inducible genes in the highly drought-tolerant cowpea: Cloning of cDNAs and analysis of the expression of the corresponding genes. Plant Cell Physiol. 1996;37:1073–1082. doi: 10.1093/oxfordjournals.pcp.a029056. [DOI] [PubMed] [Google Scholar]
- Iuchi S, YamaguchiShinozaki K, Urao T, Shinozaki K. Characterization of two cDNAs for novel drought-inducible genes in the highly drought-tolerant cowpea. J Plant Res. 1996;109:415–424. doi: 10.1007/BF02344557. [DOI] [PubMed] [Google Scholar]
- Iuchi S, Kobayashi M, Yamaguchi-Shinozaki K, Shinozaki K. A stress-inducible gene for 9-cis-epoxycarotenoid dioxygenase involved in abscisic acid biosynthesis under water stress in drought-tolerant cowpea. Plant Physiol. 2000;123:553–562. doi: 10.1104/pp.123.2.553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diop NN, Kidric M, Repellin A, Gareil M, d'Arcy-Lameta A, Thi ATP, Zuily-Fodil Y. A multicystatin is induced by drought-stress in cowpea (Vigna unguiculata (L.) Walp.) leaves. FEBS Lett. 2004;577:545–550. doi: 10.1016/j.febslet.2004.10.014. [DOI] [PubMed] [Google Scholar]
- El-Maarouf H, d'Arcy-Lameta A, Gareil M, Zuily-Fodil Y, Pham-Thi AT. Cloning and expression under drought of cDNAs coding for two PI-PLCs in cowpea leaves. Plant Physiol Biochem. 2001;39:167–172. doi: 10.1016/S0981-9428(00)01226-2. [DOI] [Google Scholar]
- D'Arcy-Lameta A, Ferrari-Iliou R, Contour-Ansel D, Pham-Thi AT, Zuily-Fodil Y. Isolation and characterization of four ascorbate peroxidase cDNAs responsive to water deficit in cowpea leaves. Ann Bot. 2006;97:133–140. doi: 10.1093/aob/mcj010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Contour-Ansel D, Torres-Franklin ML, De Carvalho MHC, D'Arcy-Lameta A. Glutathione reductase in leaves of cowpea: Cloning of two cDNAs, expression and enzymatic activity under progressive drought stress, desiccation and abscisic acid treatment. Ann Bot. 2006;98:1279–1287. doi: 10.1093/aob/mcl217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muchero W, Ehlers JD, Close TJ, Roberts PA. Mapping QTL for drought stress-induced premature senescence and maturity in cowpea [Vigna unguiculata (L.) Walp.] Theor Appl Genet. 2009;118:849–863. doi: 10.1007/s00122-008-0944-7. [DOI] [PubMed] [Google Scholar]
- Bartel DP. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. doi: 10.1016/S0092-8674(04)00045-5. [DOI] [PubMed] [Google Scholar]
- Brodersen P, Sakvarelidze-Achard L, Bruun-Rasmussen M, Dunoyer P, Yamamoto YY, Sieburth L, Voinnet O. Widespread translational inhibition by plant miRNAs and siRNAs. Science. 2008;320:1185–1190. doi: 10.1126/science.1159151. [DOI] [PubMed] [Google Scholar]
- Jover-Gil S, Candela H, Ponce MR. Plant microRNAs and development. Int J Dev Biol. 2005;49:733–744. doi: 10.1387/ijdb.052015sj. [DOI] [PubMed] [Google Scholar]
- Phillips JR, Dalmay T, Bartels D. The role of small RNAs in abiotic stress. FEBS Lett. 2007;581:3592–3597. doi: 10.1016/j.febslet.2007.04.007. [DOI] [PubMed] [Google Scholar]
- Sunkar R, Chinnusamy V, Zhu J, Zhu JK. Small RNAs as big players in plant abiotic stress responses and nutrient deprivation. Trends Plant Sci. 2007;12:301–309. doi: 10.1016/j.tplants.2007.05.001. [DOI] [PubMed] [Google Scholar]
- Navarro L, Dunoyer P, Jay F, Arnold B, Dharmasiri N, Estelle M, Voinnet O, Jones JDG. A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science. 2006;312:436–439. doi: 10.1126/science.1126088. [DOI] [PubMed] [Google Scholar]
- Sunkar R, Zhu JK. Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell. 2004;16:2001–2019. doi: 10.1105/tpc.104.022830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li WX, Oono Y, Zhu JH, He XJ, Wu JM, Iida K, Lu XY, Cui XP, Jin HL, Zhu JK. The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell. 2008;20:2238–2251. doi: 10.1105/tpc.108.059444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao BT, Liang RQ, Ge LF, Li W, Xiao HS, Lin HX, Ruan KC, Jin YX. Identification of drought-induced microRNAs in rice. Biochem Biophys Res Commun. 2007;354:585–590. doi: 10.1016/j.bbrc.2007.01.022. [DOI] [PubMed] [Google Scholar]
- Sunkar R, Zhou XF, Zheng Y, Zhang WX, Zhu JK. Identification of novel and candidate miRNAs in rice by high throughput sequencing. BMC Plant Biol. 2008;8:25. doi: 10.1186/1471-2229-8-25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu HH, Tian X, Li YJ, Wu CA, Zheng CC. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA. 2008;14:836–843. doi: 10.1261/rna.895308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou L, Liu Y, Liu Z, Kong D, Duan M, Luo L. Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J Exp Bot. 2010;61:4157–4168. doi: 10.1093/jxb/erq237. [DOI] [PubMed] [Google Scholar]
- Szittya G, Moxon S, Santos DM, Jing R, Fevereiro MP, Moulton V, Dalmay T. High-throughput sequencing of Medicago truncatula short RNAs identifies eight new miRNA families. BMC Genomics. 2008;9:593. doi: 10.1186/1471-2164-9-593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lelandais-Briere C, Naya L, Sallet E, Calenge F, Frugier F, Hartmann C, Gouzy J, Crespi M. Genome-Wide Medicago truncatula small RNA analysis revealed novel microRNAs and isoforms differentially regulated in roots and nodules. Plant Cell. 2009;21:2780–2796. doi: 10.1105/tpc.109.068130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian S, Fu Y, Sunkar R, Barbazuk WB, Zhu JK, Yu O. Novel and nodulation-regulated microRNAs in soybean roots. BMC Genomics. 2008;9:160. doi: 10.1186/1471-2164-9-160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arenas-Huertero C, Perez B, Rabanal F, Blanco-Melo D, De la Rosa C, Estrada-Navarrete G, Sanchez F, Covarrubias AA, Reyes JL. Conserved and novel miRNAs in the legume Phaseolus vulgaris in response to stress. Plant Mol Biol. 2009;70:385–401. doi: 10.1007/s11103-009-9480-3. [DOI] [PubMed] [Google Scholar]
- Lu YZ, Yang XY. Computational identification of novel microRNAs and their targets in Vigna unguiculata. Comp Funct Genomics. 2010. pii: 128297. [DOI] [PMC free article] [PubMed]
- Paul S, Kundu A, Pal A. Identification and validation of conserved microRNAs along with their differential expression in roots of Vigna unguiculata grown under salt stress. Plant Cell Tiss Org Cult. 2011;105:233–242. doi: 10.1007/s11240-010-9857-7. [DOI] [Google Scholar]
- Hall AE, Cisse N, Thiaw S, Elawad HOA, Ehlers JD, Ismail AM, Fery RL, Roberts PA, Kitch LW, Murdock LL, Boukar O, Phillips RD, McWatters KH. Development of cowpea cultivars and germplasm by the Bean/Cowpea CRSP. Field Crops Res. 2003;82:103–134. doi: 10.1016/S0378-4290(03)00033-9. [DOI] [Google Scholar]
- Hall A. Breeding for adaptation to drought and heat in cowpea. Eur J Agron. 2004;21:447–454. doi: 10.1016/j.eja.2004.07.005. [DOI] [Google Scholar]
- Muchero W, Ehlers JD, Roberts PA. Seedling stage drought-induced phenotypes and drought-responsive genes in diverse cowpea genotypes. Crop Sci. 2008;48:541–552. doi: 10.2135/cropsci2007.07.0397. [DOI] [Google Scholar]
- Zheng J, Fu J, Gou M, Huai J, Liu Y, Jian M, Huang Q, Guo X, Dong Z, Wang H, Wang G. Genome-wide transcriptome analysis of two maize inbred lines under drought stress. Plant Mol Biol. 2010;72:407–421. doi: 10.1007/s11103-009-9579-6. [DOI] [PubMed] [Google Scholar]
- Helms D, Panella L, Buddenhagen IW, Tucker CL, Gepts PL. Registration of California Blackeye 46 Cowpea. Crop Sci. 1991;31:1703–1703. [Google Scholar]
- Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ. miRBase: tools for microRNA genomics. Nucleic Acids Res. 2008;36:D154–D158. doi: 10.1093/nar/gkn221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Audic S, Claverie JM. The significance of digital gene expression profiles. Genome Res. 1997;7:986–995. doi: 10.1101/gr.7.10.986. [DOI] [PubMed] [Google Scholar]
- Kantar M, Lucas SJ, Budak H. miRNA expression patterns of Triticum dicoccoides in response to shock drought stress. Planta. 2011;233:471–484. doi: 10.1007/s00425-010-1309-4. [DOI] [PubMed] [Google Scholar]
- Frazier TP, Sun G, Burklew CE, Zhang B. Salt and Drought stresses induce the aberrant expression of microRNA genes in tobacco. Mol Biotechnol. 2011. Epub Feb.26,2011. [DOI] [PubMed]
- Shen J, Xie K, Xiong L. Global expression profiling of rice microRNAs by one-tube stem-loop reverse transcription quantitative PCR revealed important roles of microRNAs in abiotic stress responses. Mol Genet Genomics. 2010;284:477–488. doi: 10.1007/s00438-010-0581-0. [DOI] [PubMed] [Google Scholar]
- Wu G, Poethig RS. Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development. 2006;133:3539–3547. doi: 10.1242/dev.02521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ding D, Zhang LF, Wang H, Liu ZJ, Zhang ZX, Zheng YL. Differential expression of miRNAs in response to salt stress in maize roots. Ann Bot. 2009;103:29–38. doi: 10.1093/aob/mcn205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King MC, Wilson AC. Evolution at two levels in humans and chimpanzees. Science. 1975;188:107–116. doi: 10.1126/science.1090005. [DOI] [PubMed] [Google Scholar]
- Guilfoylea TJ, Hagena G. Auxin response factors. Curr Opin Plant Biol. 2007;10:453–460. doi: 10.1016/j.pbi.2007.08.014. [DOI] [PubMed] [Google Scholar]
- Liu PP, Montgomery TA, Fahlgren N, Kasschau KD, Nonogaki H, Carrington JC. Represssion of Auxin Responsive Factor10 by microRNA160 is critical for seed germination and post-germination stages. Plant J. 2007;52:133–146. doi: 10.1111/j.1365-313X.2007.03218.x. [DOI] [PubMed] [Google Scholar]
- Yan YS, Chen XY, Yang K, Sun ZX, Fu YP, Zhang YM, Fang RX. Overexpression of an B-box protein gene reduces abiotic stress tolerance and promotes root growth in rice. Mol Plant. 2011;4:190–197. doi: 10.1093/mp/ssq066. [DOI] [PubMed] [Google Scholar]
- Jia Y, Gu H, Wang X, Chen Q, Shi S, Zhang J, Ma L, Zhang H, Ma H. Molecular cloning and characterization of an F-box family gene carF-box1 from chickpea (Cicer arietinum L.) Mol Biol Rep. 2011. Epub ahead, 12 June 2011. [DOI] [PubMed]
- Golldack D, Lüking I, Yang O. Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep. 2011;30:1383–1391. doi: 10.1007/s00299-011-1068-0. [DOI] [PubMed] [Google Scholar]
- Long TA, Tsukagoshi H, Busch W, Lahner B, Salt DE, Benfey PN. The bHLH transcription factor POPEYE regulates response to iron deficiency in Arabidopsis roots. Plant Cell. 2010;22:2219–2236. doi: 10.1105/tpc.110.074096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li F, Guo S, Zhao Y, Chen D, Chong K, Xu Y. Overexpression of a homopeptide repeat-containing bHLH protein gene (OrbHLH001) from Dongxiang wild rice confers freezing and salt tolerance in transgenic Arabidopsis. Plant Cell Rep. 2010;29:977–986. doi: 10.1007/s00299-010-0883-z. [DOI] [PubMed] [Google Scholar]
- German MA, Pillay M, Jeong DH, Hetawal A, Luo SJ, Janardhanan P, Kannan V, Rymarquis LA, Nobuta K, German R, De Paoli E, Lu C, Schroth G, Meyers BC, Green PJ. Global identification of microRNA-target RNA pairs by parallel analysis of RNA ends. Nat Biotechnol. 2008;26:941–946. doi: 10.1038/nbt1417. [DOI] [PubMed] [Google Scholar]
- Addo-Quaye C, Eshoo TW, Bartel DP, Axtell MJ. Endogenous siRNA and miRNA targets identified by sequencing of the Arabidopsis degradome. Curr Biol. 2008;18:758–762. doi: 10.1016/j.cub.2008.04.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu L, Zhang QQ, Zhou HY, Ni FR, Wu XY, Qi YJ. Rice MicroRNA effector complexes and targets. Plant Cell. 2009;21:3421–3435. doi: 10.1105/tpc.109.070938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scholander PF, Bradstreet ED, Hemmingsen EA, Hammel HT. Sap pressure in vascular plants - Negative hydrostatic pressure can be measured in plants. Science. 1965;148:339–346. doi: 10.1126/science.148.3668.339. [DOI] [PubMed] [Google Scholar]
- Ghildiyal M, Seitz H, Horwich MD, Li CJ, Du TT, Lee S, Xu J, Kittler ELW, Zapp ML, Weng ZP, Zamore PD. Endogenous siRNAs derived from transposons and mRNAs in Drosophila somatic cells. Science. 2008;320:1077–1081. doi: 10.1126/science.1157396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li R, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K, Wang J. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics. 2009;25:1966–1967. doi: 10.1093/bioinformatics/btp336. [DOI] [PubMed] [Google Scholar]
- Close TJ, Wanamaker S, Roose ML, Lyon M. HarvEST. Methods Mol Biol. 2007;406:161–177. doi: 10.1007/978-1-59745-535-0_7. [DOI] [PubMed] [Google Scholar]
- Timko MP, Rushton PJ, Laudeman TW, Bokowiec MT, Chipumuro E, Cheung F, Town CD, Chen X. Sequencing and analysis of the gene-rich space of cowpea. BMC Genomics. 2008;9:103. doi: 10.1186/1471-2164-9-103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyers BC, Axtell MJ, Bartel B, Bartel DP, Baulcombe D, Bowman JL, Cao X, Carrington JC, Chen X, Green PJ, Griffiths-Jones S, Jacobsen SE, Mallory AC, Martienssen RA, Poethig RS, Qi Y, Vaucheret H, Voinnet O, Watanabe Y, Weigel D, Zhu JK. Criteria for annotation of plant MicroRNAs. Plant Cell. 2008;20:3186–3190. doi: 10.1105/tpc.108.064311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Markham NR, Zuker M. In: Bioinformatics, Volume II Structure, Function and Applications. Keith JM, editor. Totowa, NJ: Humana Press; 2008. UNAFold: software for nucleic acid folding and hybriziation; pp. 3–31. [Google Scholar]
- Pearson WR. Flexible sequence similarity searching with the FASTA3 program package. Methods Mol Biol. 2000;132:185–219. doi: 10.1385/1-59259-192-2:185. [DOI] [PubMed] [Google Scholar]
- Enright AJ, John B, Gaul U, Tuschl T, Sander C, Marks DS. MicroRNA targets in Drosophila. Genome Biol. 2004;5:R1. doi: 10.1186/gb-2003-5-1-r1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Allen E, Xie ZX, Gustafson AM, Carrington JC. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell. 2005;121:207–221. doi: 10.1016/j.cell.2005.04.004. [DOI] [PubMed] [Google Scholar]
- Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006;34:D140–D144. doi: 10.1093/nar/gkj112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fahlgren N, Howell MD, Kasschau KD, Chapman EJ, Sullivan CM, Cumbie JS, Givan SA, Law TF, Grant SR, Dangl JL, Carrington JC. High-throughput sequencing of Arabidopsis microRNAs: evidence for frequent birth and death of MIRNA genes. PLoS One. 2007;2:e219. doi: 10.1371/journal.pone.0000219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez-Ballester D, Casero D, Cokus S, Pellegrini M, Merchant SS, Grossman AR. RNA-Seq analysis of sulfur-deprived Chlamydomonas cells reveals aspects of acclimation critical for cell survival. Plant Cell. 2010;22:2058–2084. doi: 10.1105/tpc.109.071167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pall GS, Hamilton AJ. Improved northern blot method for enhanced detection of small RNA. Nature Protocols. 2008;3:1077–1084. doi: 10.1038/nprot.2008.67. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
MiRNAs that were identified in cowpea. Detailed information of the predicted cowpea miRNAs and their targets.
Predicted hairpin structures of nine genotype-specific miRNAs. Predicted structures of nine genotype-specific miRNAs with mature miRNAs marked in green.
Mapping of small RNA reads from four libraries to the precursors of nine genotype-specific miRNAs. Each figure shows the precursor sequence, predicted hairpin structure, and how each unique read was mapped to the precursor.
Drought-associated miRNAs in cowpea. Detailed information of drought-associated miRNAs and their targets.
Other conserved miRNAs that were expressed in cowpea. Three conserved miRNAs and their expression values in two cowpea genotypes under two growth conditions.