Abstract
We have developed a new assay, ISET (isolation by size of epithelial tumor cells), which allows the counting and the immunomorphological and molecular characterization of circulating tumor cells in patients with carcinoma, using peripheral blood sample volumes as small as 1 ml. Using this assay, epithelial tumor cells can be isolated individually by filtration because of their larger size when compared to peripheral blood leukocytes. ISET parameters were defined using peripheral blood spiked with tumor cell lines (HepG2, Hep3B, MCF-7, HeLa, and LNCaP). ISET can detect a single, micropipetted tumor cell, added to 1 ml of blood. We also demonstrate that fluorescence in situ hybridization can be used to perform chromosomal analyses on tumor cells collected using ISET. Polymerase chain reaction-based genetic analyses can be applied to ISET-isolated cells, and, as an example, we demonstrate homozygous p53 deletion in single Hep3B cells after filtration and laser microdissection. Finally, we provide evidence for the in vivo feasibility of ISET in patients with hepatocellular carcinoma undergoing tumor resection. ISET, but not reverse transcriptase-polymerase chain reaction, allowed analysis of cell morphology, counting of tumor cells, and demonstration of tumor microemboli spread into peripheral blood during surgery. Overall, ISET constitutes a novel approach that should open new perpectives in molecular medicine.
Although cancer is being diagnosed at increasingly earlier stages, most patients continue to succumb to metastatic disease, 1 and suitable assays that efficiently recognize invasive tumor cells appearing in the peripheral blood circulation are lacking so far. In this context, animal studies demonstrating circulating tumorous cells (CTCs) spread from solid tumors 2 have been followed by similar efforts in humans, the major obstacle being, at least in patients without clinical signs of metastasis, their extremely low number, 3 in the range of one per milliliter.
Because of their high sensitivity, reverse transcriptase-polymerase chain reaction (RT-PCR)-based assays have been applied to this problem over the past few years. 3 However, cell integrity is destroyed during RNA extraction, thus preventing the analysis of cell morphology and phenotype. Moreover, RT-PCR tests are unsuitable, in most instances, for distinguishing between circulating nontumorous and tumorous cells. 4,5 These assays detect tissue-specific mRNAs in peripheral blood, and thus their specificity for tumor cells depends entirely on the assumption that normal tissue cells do not circulate, unless they become tumorous.
We describe here a new technology, ISET (isolation by size of epithelial tumor cells), which provides a morphological, immunocytological, and genetic characterization of individual CTCs. The test allows the collection of carcinoma cells by filtration, because of their larger size when compared to peripheral blood leukocytes. We demonstrate that tumor cell karyotype analysis can easily be performed with fluorescence in situ hybridization (FISH). Furthermore, selected individual cells can be recovered by laser microdissection, and their DNA amplified and screened for genetic abnormalities in target sequences. Additional advantages of this technique are its high sensitivity and its suitability for application to a broad range of carcinomas.
We validated this experimental approach in patients with hepatocellular carcinoma undergoing tumor resection and showed that ISET, but not α-fetoprotein (AFP) nested PCR, allowed the identification and counting of tumor liver cells that had been disseminated during surgery.
Patients and Methods
Patients
Seven patients with hepatocellular carcinoma undergoing surgical liver resection were included in this study. Peripheral blood samples (15 ml) were obtained, on EDTA buffer, before intervention and during surgery and processed up to 5 hours after collection. Nine milliliters of blood was used for RT-PCR, and 6 ml for ISET. Eight healthy volunteers and eight patients with chronic hepatitis were included as controls. Informed consent was obtained from all patients and volunteers participating in this study. Investigations were performed after approval of the local ethical committee.
Cell Lines
HepG2 (HB 8065; American Type Culture Collection (ATCC), Rockville, MD) and Hep3B (ATCC, HB 8064) cells (derived from human hepatocellular carcinomas), MCF-7 (ATCC, HTB-22, derived from human breast adenocarcinoma), HeLa (ATCC, CCL-2, derived from a human cervix epithelioid carcinoma), and LNCaP cells (ATCC, CRL 1740, derived from a human metastatic prostatic adenocarcinoma) were cultured as previously described. 6
Morphometric Analysis
Morphometric analysis of May-Grumwald-Giemsa-stained HepG2, Hep3B, MCF-7, HeLa, and LNCaP cells and peripheral blood leukocytes (PBLs) was performed with a transmission microscope and Visiolab 200 and Visiolab 1000 softwares. At least 200 cells were analyzed for each cell type. The morphometric instrument was carefully calibrated before each measurement. Only nonclustered isolated cells were analyzed to record the whole cell area, cytoplasm and nucleus areas, and the mean tumor cell/leukocyte area ratio.
ISET
ISET was carried out using a previously described 7,8 module of filtration (licenses EP513139, US5606351, JO5504405) kindly provided by the Biocom company (Les Ulis, France) and a polycarbonate Track-Etch-type membrane (Cyclotron Technology) with calibrated, 8-μm-diameter, cylindrical pores. The module of filtration has 12 wells, making it possible to load and filter 12 individual samples in parallel. Each sample is filtered through a 0.6-cm-diameter surface area in the membrane. Peripheral blood (6 ml) from patients with carcinoma or from healthy volunteers was collected on buffered EDTA, diluted 1:10 with the filtration buffer containing 0.175% saponin, 0.2% paraformaldehyde, 0.0372% EDTA, and 0.1% bovine serum albumin, left for 10 minutes at room temperature, and filtered. Ten milliters of diluted solution, corresponding to 1 ml of undiluted blood, was loaded on each well and filtered by gentle aspiration under vacuum (created by a vacuum pump). The membrane was then washed once by aspiration with phosphate-buffered saline (PBS), disassembled from the filtration module, and allowed to air-dry. One or more spots, each one corresponding to 1 ml of filtered blood, were then stained with hematoxylin and eosin (H&E) or May-Grumwald-Giemsa, washed once with PBS, and dried. For microscopic studies, the membrane was placed on a specifically designed stage 8 without aspiration.
Sensitivity Tests
Sensitivity tests were performed in parallel by ISET and RT-PCR. One and three cultured HepG2, LNCaP, and MCF-7 cells, in fivefold replicates, were individually collected under a microscope equipped with a magnifying glass and mixed to 1 ml of peripheral blood. These 1-ml samples were then analyzed by ISET or by RT-PCR. ISET was carried out by counting tumor cells after KL1 immunostaining. RT-PCR analyses were performed with AFP (α-fetoprotein)-specific primers for HepG2 cells and PSA (prostate specific antigen)-specific primers for LNCaP cells (see below) (Table 1) ▶ . Individually collected MCF-7 cells were tested using ISET only.
Table 1.
Primers Used for RT-PCR and PCR Assays
| Primers | Sequence | Position | Size PCR product |
|---|---|---|---|
| p53 | |||
| P53 F2.2 | 5′-GCCATTCTTTTCCTGCTCCA-3′ | Exon 2 | 302 |
| P53 RS2 | 5′-TTCGCTTCCCACAGGTCTCT-3′ | ||
| P53 F3.1 | 5′-CCCAGCCCCCTAGCAGAGAC-3′ | Exon 3 | 98 |
| P53 R3 | 5′-GGTCCCAGCCCAACCCTTGT-3′ | ||
| P53 4.1 | 5′-CTGGTCCTCTGACTGCTCTT-3′ | Exon 4 | 338 |
| P53 RS4 | 5′-AGGCATTGAAGTCTCATGGA-3′ | ||
| P53 F5 | 5′-TGTTTGTTTCTTTGCTGCCGTGT-3′ | Exon 5 | 238 |
| P53 R5 | 5′-CCCTGTCGTCTCTCCAGCCC-3′ | ||
| P53 F6.1 | 5′-TCCGCGCCATGGCCATCTAC-3′ | Exon 6 | 311 |
| P53 R6 | 5′-AACCACCCTTAACCCCTCCT-3′ | ||
| P53 F7 | 5′-CTCATCTTGGGCCTGTGTT-3′ | Exon 7 | 185 |
| P53 RS7 | 5′-GGGTCAGCGGCAAGCAGAG-3′ | ||
| P53 F8 | 5′-CTGCCTCTTGCTTCTCTTTT-3′ | Exon 8 | 235 |
| P53 RS8 | 5′-GAGGCAAGGAAAGGTGATAA-3′ | ||
| P53 F9 | 5′-GGAGGAGACCAAGGGTGCAGTT-3′ | Exon 9 | 215 |
| P53 R9.1 | 5′-ATGCCCCAATTGCAGGTAAAACA-3′ | ||
| P53 F10 | 5′-GCGACGAGAGTGAGACCCCATC-3′ | Exon 10 | 322 |
| P53 R10.1 | 5′-AGGGGAGTAGGGCCAGTAAG-3′ | ||
| P53 FS11 | 5′-CAGGGAAAAGGGGCACAGAC-3′ | Exon 11 | 323 |
| P53 R11 | 5′-CCTCCTCCCCACAACAAAACACC-3′ | ||
| HLA | |||
| HLA GH26 (F) | 5′-GTGCTGCAGGTGTAAACTTGTACCAG-3′ | Exon 4 | 250 |
| HLA GH27 (R) | 5′-CACGGATCCGGTAGCAGCGGTAGAGTT-3′ | ||
| α-Fetoprotein | |||
| AFP out 1 (F) | 5′-CTCTTCCAGAAACTAGGAGAA-3′ | Exon 11 | 537 |
| AFP out 2 (R) | 5′-CTCTTCAGCAAAGCAGACTT-3′ | Exon 13 | |
| AFP in 3 (F) | 5′-GCTGACATTATTATCGGACAC-3′ | Exon 12 | 282 |
| AFP in 4 (R) | 5′-AGCCTCAAGTTGTTCCTCTGT-3′ | Exon 13 | |
| Prostate-specific antigen | |||
| PSA Ex | 5′-GAACCAGAGGAGTGTCTTGCCCCAAAA-3′ | Exon 3–4 | 421 |
| PSA 7 | 5′-CACAGACACCCCATCCTATC-3′ | 3′NTR | |
| PDC2 | 5′-TGGACCTCACACCTAAGGACAAAG-3′ | 3′NTR | 343 |
| PSA 8 | 5′-TGCGCAAGTTCACCCTCA-3′ | Exon 4 |
Detection of CTC by RT-PCR
Peripheral blood (6 ml) was collected from patients in buffered EDTA. Nucleated cells were isolated by Ficoll (Ficoll Paque Plus; Pharmacia Biotech, Uppsala, Sweden) in preadapted tubes (Leucosep, Bernas, Paris) and resuspended in TRIzol B (Life Technologies). RNA was extracted and resuspended in 6 μl of diethylpyrocarbonate-treated water (1 μl of DEPC-treated water per milliliter of processed blood). RNA solution (6 μl) was reverse-transcribed in a final volume of 20 μl. Ten microliters of cDNA, corresponding to 3 ml of peripheral blood samples, was amplified with a nested protocol using liver-specific (AFP) and prostate-specific (PSA) primers, as already reported. 4,6 For sensitivity tests, the whole RNA extracted from 1 ml of blood spiked with tumor cells was reverse transcribed and then amplified as described.
Cell Staining and Immunostaining
To stain the cells, after rehydration with PBS, hematoxylin was applied to the membrane for 2 minutes, followed by 1% eosin for 3 minutes. The cells were permeabilized with 0.2% Triton for 10 minutes before immunostaining. Primary antibodies diluted 1:100 in 10% fetal calf serum were then applied to the spot for 1 hour at room temperature. These were KL1 (Cytokeratin gp 56 kd; Immunotech S. A., Marseille, France), a cytokeratin, broad spectrum monoclonal antibody; anti-AFP antibody from rabbit (Dako, Glostrup, Denmark); anti-albumin from rabbit (Dako); rabbit anti-glutamine synthetase 9 ; monoclonal anti-LCA antibody (Dako) reactive to lymphomonocytic cells; and goat anti-MMP3 (or stromelysin 1, which reacts with an enzyme catalyzing degradation of collagen types IV, IX, and XI and proteoglycans, fibronectin and laminin) (Binding Site, Birmingham, UK). The spots were then treated with a biotinylated mixture of anti-mouse and anti-rabbit secondary antibodies for 30 minutes (peroxidase/DAB ChemMate Detection Kit; Dako), followed by a complex, streptavidin-peroxidase, for 30 minutes and DAB (diaminobenzidine) for 15 minutes. For anti-MMP3, a universal biotinylated secondary antibody (Binding Site) was used. Cell nuclei were counterstained with Mayer’s hematoxylin (Dako). For double staining, we applied anti-AFP or KL1 antibodies, revealed with a peroxidase/DAB ChemMate Detection Kit (Dako) followed by LCA antibody, and an alkaline phosphatase ChemMate Detection Kit (Dako). For immunofluorescence, fluorescein-conjugated anti-mouse or anti-rabbit secondary antibodies (Amersham, Les Ulis, France) were applied for 45 minutes at room temperature, and the nuclei were counterstained with 7-aminoactinomycin D (Sigma Chemical Co., St. Louis, MO) for 5 minutes at room temperature.
Molecular Analysis
Laser microdissection of epithelial tumor cells was performed on H&E-stained filters. Single spots were cut from the membrane and mounted with nail polish on a 0.17-mm-thick slide. Slides were then placed on a robot-stage microscope equipped with a 337-nm pulsed laser microbeam (P.A.L.M., Munich, Germany). Individual cells were dissected with the laser microbeam and then catapulted with a single laser shot into the lid of a microfuge tube suitable for PCR. 10 The collected cell was then recovered in 15 μl of lysis buffer (100 mmol/L Tris-HCl (pH 8) and 400 μg/ml proteinase K) and spun down by centrifugation. After lysis at 37°C for 16 hours, proteinase K was inactivated at 90°C for 10 minutes and DNA was amplified using the PEP (primer-extension preamplification) protocol. 11 Five out of 60 μl of the extension product was coamplified in a final volume of 100 μl containing 10 mmol/L Tris-HCl, 50 mmol/L KCl, 2 mmol/L MgCl2, 0.01% gelatin, 250 mmol/L of each of four deoxynucleotides, 10 pmol of each HLA and p53-specific primer, and 2.5 U of Taq polymerase (Perkin-Elmer Cetus, Emeryville, CA) for 40 cycles. PCR products were then analyzed by electrophoresis in a 2% agarose gel.
For p53 gene mapping, 50 ng of DNA extracted from Hep3B or HepG2 cells was coamplified with HLA and p53 primers. Ten p53-specific primer sets were used to amplify p53 exons (Table 1) ▶ .
Fluorescence in Situ Hybridization
The membranes were pretreated with Triton 0.1% in PBS buffer at room temperature for 10 minutes, followed by proteinase K (0.1 μg/ml) digestion at 37°C for 8 minutes. Interphase nuclei were denatured in 70% formamide/2SSC (sodium saline citrate) at 72°C for 2 minutes and then dehydrated through an ethanol series (70, 80, 90, 100%). Hybridization and signal revelation of a chromosome 1 centromeric biotinylated (Oncor, Gaithersburg, MD) or fluorophore-labeled probe (Vysis, Downers Grove, IL) were performed in accordance with the manufacturer’s instructions.
Statistical Methods
Statistical analyses were performed using SPSS 7.5 statistical software (Chicago, IL).
Results
Morphometric Analyses
Mean whole-cell areas ranged from 796 μm 2 (Hep3B) to 396 μm 2 (MCF-7), and the mean tumor cell/leukocyte area ratio was 5.7 for Hep3B, 4.5 for HepG2 and HeLa, 4.1 for LNCaP, and 2.8 for MCF-7. A significant (P < 0.001) difference in whole-cell area was found between tumor cells from any of the carcinoma cell lines and PBLs (Figure 1A) ▶ .
Figure 1.
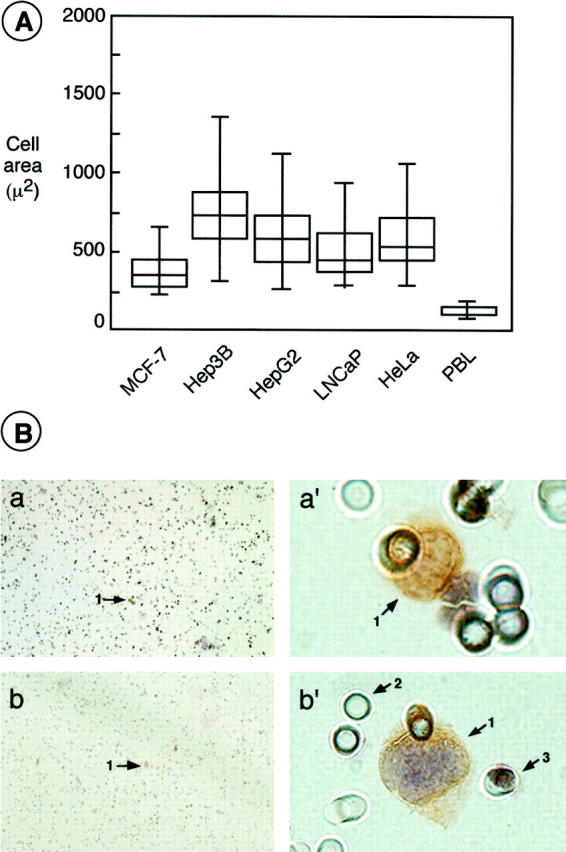
A: Box plots of cell areas according to cell type. Each box has its ends at the quartiles, and the median of distribution is marked by a line within the box. The whiskers at either end extend to the 10th and 90th percentiles. A significant difference (P < 0.001) in the whole cell area was found between tumor cells from any of the carcinoma cell lines and PBL. B: Sensitivity test. One MCF-7 (a and a′) or LNCaP (b and b′) cell was micropipetted under the microscope, mixed with 1 ml of blood, and recovered using ISET. a and b: Low magnification (×10) showing the area of filtration and the single cell (arrow). a′ and b′: The same cell as in a and b, but at a higher magnification (×100). Immunostaining was with KL1 antibody. Note the well-preserved morphological details. Arrows: 1, tumorous cell; 2, membrane pore; 3, leukocyte (trapped in a pore lumen).
Sensitivity Tests
Sensitivity tests performed on individually collected cultured cells showed that the sensitivity threshold of ISET is close to one carcinoma cell per milliliter. In fivefold replicate tests using one cell, we found one LNCaP cell five times, one HepG2 cell four times, and one MCF-7 cell three times. In fivefold replicate tests using three cells, we found three LNCaP cells three times (two cells and one cell once), three HepG2 cells four times (one cell once), and three MCF-7 cells four times (no cells one time), using ISET (Table 2) ▶ . In contrast, RT-nested PCR protocols were incapable of detecting either one or three individually collected HepG2 or LNCaP cells when mixed with such a small volume of blood (1 ml) (Figure 1B ▶ , Table 2 ▶ ).
Table 2.
In Vitro and in Vivo Sensitivity of ISET
| No. of cells detected by ISET | |||
|---|---|---|---|
| Hep G2 | LNC aP | MCF-7 | |
| a. In vitro tests | |||
| No. of micropipetted cells | |||
| 3 | 3 (−) | 1 (−) | 3 |
| 3 (−) | 3 (−) | 3 | |
| 3 (−) | 3 (−) | 0 | |
| 1 (−) | 3 (−) | 3 | |
| 3 (−) | 2 (−) | 3 | |
| 1 | 1 (−) | 1 (−) | 1 |
| 1 (−) | 1 (−) | 1 | |
| 0 (−) | 1 (−) | 0 | |
| 1 (−) | 1 (−) | 1 | |
| 1 (−) | 1 (−) | 0 | |
| b. In vivo tests in patients with liver cancer |
| No. of tumor cells (*) and tumor microemboli (°) detected by ISET | ||
|---|---|---|
| Patient | Before surgery | During tumor resection |
| 1 | 4° (−) | 3*+ 5° (−) |
| 2 | 0 (−) | 1* (−) |
| 3 | 0 (−) | 1*+ 12° (−) |
| 4 | 1° (−) | 0 (−) |
| 5 | 0 (−) | 2* (+) |
| 6 | 1° (−) | 2° (+) |
| 7 | 0 (−) | 2* (+) |
RT-PCR results are in parentheses. ISET tests were performed in 1 ml peripheral blood samples. RT-PCR in vitro and in vivo tests were performed in 1 and 3 ml of blood, respectively.
Immunomorphological Analysis of Tumor Cells Isolated by ISET
Morphological analysis of epithelial cells, after H&E staining, showed that cell morphology is undamaged after ISET. An average number of 0.0002% leukocytes is also retained on the membrane, often trapped in the pore lumen. However, they are easy to recognize because of their size and nuclear morphology. All carcinoma cells were positive for KL1 immunostaining. The fine granular cytoplasmic staining was particularly intense in MCF-7 cells and much weaker in LNCaP. No KL1 staining was detected in PBL, except in some leukocytes trapped in the membrane pores.
In double staining with anti-AFP or KL1 and anti-LCA antobodies (Figure 2c) ▶ , anti-LCA gives a clear cell membrane labeling and a weak cytoplasmic staining. With anti-human matrix metalloproteinase 3 we observed a cytoplasmic diffuse staining in LNCaP (Figure 2d) ▶ and HepG2 cells.
Figure 2.
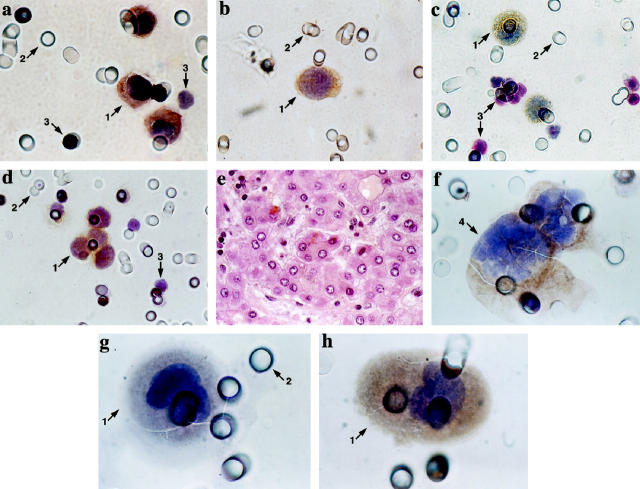
Immunocytochemical characterization of cells mixed with peripheral blood and recovered using ISET. a–d: Cells from tumorous cell lines. a: HepG2 cells stained with anti-KL1 (×60). b: Anti-albumin-positive Hep3B cell (×40). c: Double staining of HepG2 cells and leukocytes with anti-AFP and anti-LCA (×40). d: LNCaP cells positive to the anti-MMP3 antibody (×40). f–h: Anti-AFP-positive tumorous cells isolated from patients with hepatocellular carcinoma before tumor resection (f, ×100) and during surgery (g and h, ×100). A tumor embolus is shown in f (arrow 4) to reproduce the histological lobular structure observed in the tumorous tissue from the same patient (e, H&E staining, ×20). Cell nuclei were counterstained with Mayer’s hematoxylin. Arrows: 1, tumorous cells; 2, membrane pores; 3, leukocytes.
Molecular Analysis
We studied Hep3B cells, which carry a p53 deletion previously described by Southern blot, 12 and HepG2 cells, the p53 gene of which is nonrearranged. In a preliminary test, we mapped the Hep3B p53 deletion by PCR with 10 exon-specific primer sets and observed that the homozygous deletion spans the area from exon 7 to exon 11 (data not shown). We then applied primers specific to p53 exons 8 and 9, in coamplification with HLA primers, to DNA obtained from single Hep3B cells isolated using ISET and laser microdissection 13 (Figure 3, A and B) ▶ . Our results demonstrate the p53 deletion in individually collected Hep3B cells.
Figure 3.
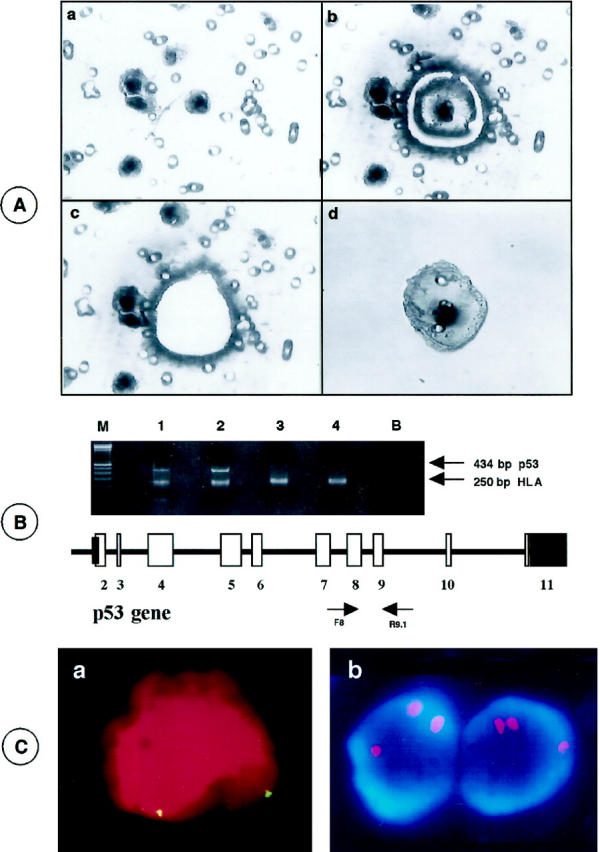
A: Laser microdissection of a Hep3B cell and p53 gene analysis. a: H&E-stained cells on the filter. The membrane around a single cell is precisely cut by the laser microbeam (b) and catapulted by laser pressure (c). d: Inversion of the optical device allows a check that the cell has now been collected into the microfuge lid. B: Gel electrophoresis and ethidium bromide staining of DNA from single HepG2 cells (lanes 1 and 2) and Hep3B cells (lanes 3 and 4) coamplified with p53 primers on exon 8 to 9 and HLA primers. Lane M: 100-bp molecular weight marker. Lane B: PCR-negative control (buffer only). The figure shows that only Hep3B cells carry an homozygous deletion of the p53 gene. C: FISH performed with centromeric chromosome 1-specific probes on HepG2 (a) and MCF-7 (b) cells. a: Biotinylated probe revealed using avidin-fluorescein isothiocyanate. The nucleus is counterstained with propidium iodide. Two spots are clearly visible, indicating two chromosomes one. b: Three red spots (fluorophore-labeled probe) reveal the presence of three chomosomes 1 in MCF-7 cells. Nuclei are counterstained using 4,6-diamidino-2-phenyl-indole (DAPl, 1 μg/ml).
Fluorescence in Situ Hybridization
We determined whether FISH could be applied to cells isolated using the ISET protocol. Figure 3C ▶ shows the data obtained using two, well characterized breast (MCF-7)- and liver (HepG2)-derived tumor cell lines. FISH, performed using a probe specific for the chromosome 1 centromere, clearly showed three and two chromosomes 1 in MCF-7 and HepG2, respectively, in line with previously reported results. 14
Patients with Hepatocellular Carcinoma
Liver-derived cells with cytological features characteristic of tumor cells, reacting or not with anti-AFP antibody, were identified in the peripheral blood of patients before and during surgery (Table 2 ▶ and Figure 2, f ▶ -h). Tumor emboli were also found, and in some instances these demonstrated features reproducing the histological lobular structure observed in tumorous tissue from the same patient (Figure 2, f and e) ▶ . The number of tumor emboli and tumor cells recovered before and during tumor resection is shown in Table 2 ▶ . In addition, a variable number of hepatoid cells was found in certain peripheral blood samples (data not shown). When compared with AFP RT-PCR tests, performed in parallel on the same peripheral blood samples, ISET produced more sensitive and specific results (Table 2) ▶ .
Finally, peripheral blood from eight healthy volunteers and eight patients with chronic hepatitis but without liver cancer scored negative with both ISET and AFP RT-PCR.
Discussion
We describe a new method, called ISET, for isolating by size circulating tumorous cells in patients with carcinoma. In fact, morphometric analyses of tumor cell lines derived from liver, prostate, breast, and cervix human carcinomas have confirmed the significantly larger cell size of these carcinoma cells as compared to peripheral blood leukocytes. This method is easy to perform, rapid, and inexpensive and makes it possible to directly isolate and count tumor cells in patients with different types of carcinoma. In previous studies 7,15 filtration analyses have been applied to tumor and/or epithelial cells obtained from peripheral blood by immunomagnetic selection. Filtration has been used to eliminate contaminating leukocytes and beads. ISET is the first test allowing direct filtration of peripheral blood that avoids immunomagnetic and flow cytometric cell separation. 16-19 Avoiding these steps, the efficiency of which often depends on the expression of selected antigens (eg, BerEP4) on the tumor cell membrane, ISET makes it possible minimize cell loss. Actually, one of the most valuable advantages of ISET is its ability to isolate epithelial cells without damaging cell morphology, a property that is connected to its high sensitivity. Thus we repeatedly detected by ISET a single carcinoma cell collected under a microscope and mixed with 1 ml of peripheral blood before filtration. In contrast, RT-PCR analyses failed to detect a single cell when diluted in only 1 ml of blood. ISET succeeded in isolating tumor cells from all tested human carcinoma-derived cell lines (HepG2, Hep3B, HeLa, LNCaP, MCF-7), including MCF-7, the size of which is only two times that of leukocytes. Our results therefore suggest that ISET is potentially suitable for application to tumor cells from a wide range of human carcinomas. Clearly, however, this methodology should not be suitable for studies on small neuroendocrine tumor cells. Further studies, focusing on additional cell lines and patients with a variety of carcinoma types, have to be performed to define the size threshold of ISET application.
Immunocytochemistry and immunofluorescence can detect tissue-specific antigens and markers of invasiveness, 20 angiogenesis, 21 apoptosis, and cell proliferation 22 in individual isolated CTCs. We also demonstrated the feasibility of cytogenetic studies in CTCs by applying the FISH protocol to this material. Furthermore, we performed PCR-based genetic analyses on individual laser-microdissected cells isolated with ISET.
In patients with liver cancer undergoing tumor resection, ISET (but not AFP RT-nested PCR) has demonstrated the iatrogenous spread of tumor cells and tumor microemboli, which is of potential importance to the prevention of tumor recurrence after surgery.
In conclusion, this new technique provides an unique opportunity for cytological analysis of peripheral blood in oncology and for combining immunomorphological studies with novel assays exploring genetic abnormalities in individual isolated cells. The potential uses for ISET go well beyond the field of oncology, because it also allows the isolation of large, circulating, nontumorous cells. For example, the isolation of trophoblastic cells from the peripheral blood of pregnant women has been initiated in our laboratory and may constitute an important step toward improving the prenatal diagnosis of genetic diseases.
Acknowledgments
We thank Dr. J.C. Bisconte for support and helpful discussion and Dr. B. Bressac for providing the p53 primers.
Footnotes
Address reprint requests to Dr. Patrizia Paterlini-Bréchot, INSERM U 370, Faculté Necker, 156 rue de Vaugirard, 75015 Paris, France. E-mail: paterlini@necker.fr.
Supported by grants from Biocom (Les Ulis, France), INSERM, Assistance Publique (contract no. CRC950191), the LNC (Ligue Nationale contre le Cancer), SIBioC (Società Italiana di Biochimica Clinica), and the CNR (Centro Nazionale delle Ricerche).
Dr. Sabile, Dr. Louha, and Dr. Sitruk contributed equally to this work.
References
- 1.Kohn E, Liotta L: Molecular insights into cancer invasion: strategies for prevention and intervention. Cancer Res 1995, 55:1856-1862 [PubMed] [Google Scholar]
- 2.Liotta L, Kleinerman J, Saidel G: Quantitative relationship of intravascular tumor cells, tumor vessels and pulmonary metastasis following tumor implantation. Cancer Res 1974, 34:997-1004 [PubMed] [Google Scholar]
- 3.Pelkey T, Frierson HJ, Bruns D: Molecular and immunological detection of circulating tumor cells and micrometastases from solid tumors. Clin Chem 1996, 42:1369-1381 [PubMed] [Google Scholar]
- 4.Louha M, Nicolet J, Zylberberg H, Sabile A, Vons C, Vona G, Poussin K, Tournebize M, Capron F, Pol S, Franco D, Lacour B, Bréchot C, Paterlini-Bréchot P: Liver resection and needle liver biopsy cause hematogenous dissemination of liver cells. Hepatology 1999, 29:879-882 [DOI] [PubMed] [Google Scholar]
- 5.Cama C, Olsson C, Buttyan R, De Vries G, Wise G, Katz A: Molecular staging of prostate cancer. III. Effects of cystoscopy and needle biopsy on the enhanced reverse transcriptase chain reaction assay. J Urol 1997, 157:1748-1751 [PubMed] [Google Scholar]
- 6.Sabile A, Louha M, Bonte E, Poussin K, Vona G, Mejean A, Chretien Y, Bougas L, Lacour B, Capron F, Roseto A, Bréchot C, Paterlini-Bréchot P: Efficiency of Ber-EP4 antibody in isolating circulating epithelial tumor cells before RT-PCR detection. Am J Clin Pathol 1999, 112:171-178 [DOI] [PubMed] [Google Scholar]
- 7.Rostagno P, Moll J, Bisconte J, Caldani C: Detection of rare circulating breast cancer cells by filtration cytometry and identification by DNA content: sensitivity in an experimental model. Anticancer Res 1997, 17:2481-2485 [PubMed] [Google Scholar]
- 8.Truchaud A, Caldani C, Bisconte J, Bergogne-Berezin E, Buisson D: Filtration cytometry: parallel real-time analysis for bacteria, cells, and particles. Clin Chem 1992, 38:1650-1651 [Google Scholar]
- 9.Christa L, Simon M, Flinois J, Gebhardt R, Bréchot C, Lasserre C: Overexpression of glutamine synthetase in human primary liver cancer. Gastroenterology 1994, 106:1312-1320 [DOI] [PubMed] [Google Scholar]
- 10.Becker I, Becker K, Rohrl M, Minkus G, Schutze K, Hofler H: Single-cell mutation analysis of tumors from stained histologic slides. Lab Invest 1996, 75:801-807 [PubMed] [Google Scholar]
- 11.Zhang L, Cui X, Schmitt K, Hubert R, W N, Arnheim N: Whole genomic amplification from a single cell: implications for genetic analysis. Proc Natl Acad Sci USA 1992, 89:5847–5851 [DOI] [PMC free article] [PubMed]
- 12.Bressac B, Galvin KM, Liang TJ, Isselbacher KJ, Wands JR, Ozturk M: Abnormal structure and expression of p53 gene in human hepatocellular carcinoma. Proc Natl Acad Sci USA 1990, 87:1973-1977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Schütze K, Posl H, Lahr G: Laser micromanipulation systems as universal tools in cellular and molecular biology and medicine. Cell Mol Biol 1998, 44:735-746 [PubMed] [Google Scholar]
- 14.Leger I, Thomas M, Ronot X, Brugal G: Detection of chromosome 1 aberrations by fluorescent in situ hybridization (FISH) in the human breast cancer cell line MCF-7. Anal Cell Pathol 1993, 5:299-309 [PubMed] [Google Scholar]
- 15.Rye P, Hoifodt H, Overli G, Fodstad O: Immunobead filtration: a novel approach for the isolation and propagation of tumor cells. Am J Pathol 1997, 150:99-106 [PMC free article] [PubMed] [Google Scholar]
- 16.Brandt B, Roetger A, Heidl S, Jackisch C, Lelle R, Assmann G, Zänker K: Isolation of blood-borne epithelium-derived c-erbB-2 oncoprotein-positive clustered cells from peripheral blood of breast cancer patients. Int J Cancer 1998, 76:824-828 [DOI] [PubMed] [Google Scholar]
- 17.Racila E, Euhus D, Weiss A, Rao C, McConnell J, Terstappen L, Uhr J: Detection and characterization of carcinoma cells in the blood. Proc Natl Acad Sci USA 1998, 95:4589-4594 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wong L, Bateman W, Morris A, Fraser I: Detection of circulating tumor cells with the magnetic activated cell sorter. Br J Surg 1995, 82:1333-1337 [DOI] [PubMed] [Google Scholar]
- 19.Melchior S, Corey E, Ellis W, Ross A, Layton T, Oswin M, Lange P, Vessella R: Early tumor cell dissemination in patients with clinically localized carcinoma of the prostate. Clin Cancer Res 1997, 3:249-256 [PubMed] [Google Scholar]
- 20.Woodhouse E, Chuaqui R, Liotta L: General mechanisms of metastasis. Cancer 1997, 80:1529-1537 [DOI] [PubMed] [Google Scholar]
- 21.Kumar R, Fidler I: Angiogenic molecules and cancer metastasis. In Vivo 1998, 12:27-34 [PubMed] [Google Scholar]
- 22.Luzzi K, MacDonald I, Schmidt E, Kerkvliet N, Morris V, Chambers A, Groom A: Multistep nature of metastatic inefficiency. Dormancy of solitary cells after successful extravasation and limited survival of early micrometastases. Am J Pathol 1998, 153:865-873 [DOI] [PMC free article] [PubMed] [Google Scholar]