Abstract
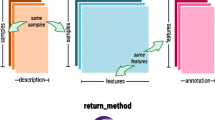
The Bioconductor project is an “open source and open development software project for the analysis and comprehension of genomic data” (1), primarily based on the R programming language. Infrastructure packages, such as Biobase, are maintained by Bioconductor core developers and serve several key roles to the broader community of Bioconductor software developers and users. In particular, Biobase introduces an S4 class, the eSet, for high-dimensional assay data. Encapsulating the assay data as well as meta-data on the samples, features, and experiment in the eSet class definition ensures propagation of the relevant sample and feature meta-data throughout an analysis. Extending the eSet class promotes code reuse through inheritance as well as interoperability with other R packages and is less error-prone. Recently proposed class definitions for high-throughput SNP arrays extend the eSet class. This chapter highlights the advantages of adopting and extending Biobase class definitions through a working example of one implementation of classes for the analysis of high-throughput SNP arrays.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JYH, Zhang J. (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol5(10):R80.
Di X, Matsuzaki H, Webster TA, Hubbell E, Liu G, Dong S, Bartell D, Huang J, Chiles R, Yang G, Mei Shen M, Kulp D, Kennedy GC, Mei R, Jones KW, Cawley S. (2005) Dynamic model based algorithms for screening and genotyping over 100 K SNPs on oligonucleotide microarrays. Bioinformatics 21(9):1958–1963.
Rabbee N, Speed TP. (2006) A genotype calling algorithm for Affymetrix SNP arrays. Bioinformatics 22(1):7–12.
Affymetrix. (2006) BRLMM: an improved genotype calling method for the genechip human mapping 500 k array set. Tech. rep., Affymetrix, Inc. White paper, Santa Clara, CA.
Carvalho B, Bengtsson H, Speed TP, Irizarry RA. (2007) Exploration, normalization, and genotype calls of high-density oligonucleotide SNP array data. Biostatistics 8(2):485–499.
Nannya Y, Sanada M, Nakazaki K, Hosoya N, Wang L, Hangaishi A, Kurokawa M, Chiba S, Bailey DK, Kennedy GC, Ogawa S. (2005) A robust algorithm for copy number detection using high-density oligonucleotide single nucleotide polymorphism genotyping arrays. Cancer Res 65(14):6071–6079.
Huang J, Wei W, Chen J, Zhang J, Liu G, Di X, Mei R, Ishikawa S, Aburatani H, Jones KW, Shapero MH. (2006) CARAT: a novel method for allelic detection of DNA copy number changes using high density oligonucleotide arrays. BMC Bioinformatics 7:83.
Laframboise T, Harrington D, Weir BA. (2006) PLASQ: a generalized linear model-based procedure to determine allelic dosage in cancer cells from SNP array data. Biostatistics 8(2):323–336.
Carter NP. (2007) Methods and strategies for analyzing copy number variation using DNA microarrays. Nat Genet 39(7 Suppl):S16–S21.
Chambers JM. (1998) Programming with Data: A Guide to the S Language, Springer-Verlag, New York.
Scharpf RB, Ting JC, Pevsner J, Ruczinski I. (2007) SNPchip: R classes and methods for SNP array data. Bioinformatics 23(5): 627–628.
Scharpf RB, Parmigiani G, Pevsner J, Ruczinski I. (2008) Hidden Markov models for the assessment of chromosomal alterations using high-throughput SNP arrays. Ann Appl Stat 2(2):687–713.
Leisch F. (2003) Sweave and beyond: Computations on text documents. In Kurt Hornik, Friedrich Leisch, and Achim Zeileis (eds). Proceedings of the 3rd International Workshop on Distributed Statistical Computing, Vienna, Austria, 2003.Sarkar D. (2008) Lattice: Multivariate Data Visualization with R. Springer, New York.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Appendix
Appendix
This document was created using Sweave (13).
-
R version 2.8.0 Under development (unstable) (2008-06-18 r45949), powerpc-apple-darwin8.11.0
-
Locale: C
-
Base packages: base, datasets, grDevices, graphics, methods, stats, tools, utils
-
Other packages: Biobase 2.1.0, DBI 0.2-4, RSQLite 0.6-4, SNPchip 1.5.2, VanillaICE 1.3.7, oligoClasses 1.1.22, pd.mapping50k.hind240 0.4.1, pd.mapping50k.xba240 0.4.1
Rights and permissions
Copyright information
© 2010 Humana Press, a part of Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Scharpf, R.B., Ruczinski, I. (2010). R Classes and Methods for SNP Array Data. In: Matthiesen, R. (eds) Bioinformatics Methods in Clinical Research. Methods in Molecular Biology, vol 593. Humana Press. https://doi.org/10.1007/978-1-60327-194-3_4
Download citation
DOI: https://doi.org/10.1007/978-1-60327-194-3_4
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-60327-193-6
Online ISBN: 978-1-60327-194-3
eBook Packages: Springer Protocols