Abstract
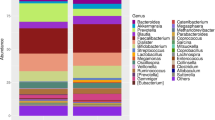
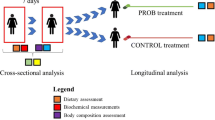
Gut microbiota is believed to play a crucial role in modulating obesity in humans, and probiotics affecting gut microbiota can alleviate some of the obesity-related health complications. The study was aimed to investigate changes in the composition of the gut microbiome in obese humans due to short-term (2 weeks) treatment of obese patients with a probiotic preparation containing Bifidobacterium longum. Faecal microbiome diversity was studied using the 16S amplicon sequencing by Illumina MiSeq. Bioinformatic analysis showed distribution across 14 phyla (with Firmicutes and Bacteroidetes dominating), 21 class, 125 genera and 973 OTUs. The probiotic treatment decreased relative abundance of Bacteroidetes (Prevotellaceae and Bacteroidaceae), while increasing that of Actinobacteria (Bifidobacteriaceae and Coriobacteriaceae), and Firmicutes (Negativicutes: Veillonellaceae and Clostridia: Peptostreptococcaceae). The probiotic treatment decreased total blood sugar and increased patients’ assessment of their physical and mental health. Thus even the short-term Bifidobacterium-based probiotic treatment brought significant compositional changes in the 16S rRNA gene diversity in faecal bacterial assemblages by increasing beneficial and decreasing pathogenic or opportunistic bacteria; the related shifts in life quality assessment necessitate further research into the causal relationships involved.


Similar content being viewed by others
Availability of Data and Material
The read data were deposited in GenBank under the sequence read archive (SRA) accession No. SRP194142.
References
Tsai YL, Lin TL, Chang CJ et al (2019) Probiotics, prebiotics and amelioration of diseases. J Biomed Sci 26:3. https://doi.org/10.1186/s12929-018-0493-6
Borovik TE, Ladodo KS, Semenova NN et al (2016) Skvashenniye molochnye produckty v pitanyi detey v Rossiskoy Federatsii: proshloye I nastoyashee [Fermented dairy products in the nutrition of infants in the Russian Federation: past and present]. Curr Pediatr 15:556–561. https://doi.org/10.15690/vsp.v15i6.1651(in Russian)
Uhr GT, Dohnalová L, Thaiss CA (2019) The dimension of time in host-microbiome interactions. mSystems 4:e00216–e00218. https://doi.org/10.1128/mSystems.00216-18
Tupikin AE, Kalmykova AI, Kabilov MR (2016) Draft genome sequence of the probiotic Bifidobacterium longum subsp. longum strain MC-42. Genome Announc 4:e01411–e01416. https://doi.org/10.1128/genomeA.01411-16
Ware JE, Kosinski M, Keller SD (1994) SF-36 physical and mental health summary scales: a user’s manual. The Health Institute, New England Medical Center, Boston
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998. https://doi.org/10.1038/nmeth.2604
Edgar RC (2016) SINTAX, a simple non-bayesian taxonomy classifier for 16S and ITS sequences. https://doi.org/10.1101/074161
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:1–9
Janssens Y, Nielandt J, Bronselaer A et al (2018) Disbiome database: linking the microbiome to disease. BMC Microbiol 18:50. https://doi.org/10.1186/s12866-018-1197-5
Braune A, Blaut M (2016) Bacterial species involved in the conversion of dietary flavonoids in the human gut. Gut Microbes 7:216–234. https://doi.org/10.1080/19490976.2016.1158395
Vermeire S, Joossens M, Verbeke K et al (2015) Donor species richness determines faecal microbiota transplantation success in inflammatory bowel disease. J Crohn’s Colitis 10:387–394. https://doi.org/10.1093/ecco-jcc/jjv203
Qin J, Li Y, Cai Z et al (2012) A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490:55–60. https://doi.org/10.1038/nature11450
Naseribafrouei A, Hestad K, Avershina E et al (2014) Correlation between the human faecal microbiota and depression. Neurogastroenterol Motil 26:1155–1162. https://doi.org/10.1111/nmo.12378
Wang Y, Song J, Zhai Y et al (2015) Romboutsia sedimentorum sp. nov., isolated from an alkaline-saline lake sediment and emended description of the genus Romboutsia. Int J Syst Evol Microbiol 65:1193–1198. https://doi.org/10.1099/ijs.0.000079
Petrov VA, Alifirova VM, Saltykova IV et al (2016) Sravnitelnoye isucheniye kushechnoi microbioty pri bolezni Parkinsona I drugikh nevrologicheskich zabolevaniyah [Comparison study of gut microbiota in case of Parkinson’s disease and other neurological disorders]. Bull Sib Med 15:113–125. https://doi.org/10.20538/1682-0363-2016-5-113-125(in Russian)
Tito RY, Cypers H, Joossens M et al (2017) Brief report: dialister as a microbial marker of disease activity in spondyloarthritis. Arthritis Rheumatol 69:114–121. https://doi.org/10.1002/art.39802
Ohara T (2019) Identification of the microbial diversity after fecal microbiota transplantation therapy for chronic intractable constipation using 16 s rRNA amplicon sequencing. PLoS ONE 14:e0214085. https://doi.org/10.1371/journal.pone.0214085
Strati F, Cavalieri D, Albanese D et al (2017) New evidences on the altered gut microbiota in autism spectrum disorders. Microbiome 5:24. https://doi.org/10.1186/s40168-017-0242-1
Park S-H, Kim K-A, Ahn Y-T et al (2015) Comparative analysis of gut microbiota in elderly people of urbanized towns and longevity villages. BMC Microbiol 15:49. https://doi.org/10.1186/s12866-015-0386-8
Lopetuso LR, Petito V, Graziani C et al (2018) Gut microbiota in health, diverticular disease, irritable bowel syndrome, and inflammatory bowel diseases: time for microbial marker of gastrointestinal disorders. Dig Disord 36:56–65. https://doi.org/10.1159/000477205
Iino C, Shimoyama T, Iino K et al (2019) Daidzein intake is associated with equol producing status through an increase in the intestinal bacteria responsible for equol production. Nutrients 11:pii: E433. https://doi.org/10.3390/nu11020433
Dillon SM, Lee EJ, Kotter CV et al (2015) Gut dendritic cell activation links an altered colonic microbiome to mucosal and systemic T-cell activation in untreated HIV-1 infection. Mucosal Immunol 9:24–37. https://doi.org/10.1038/mi.2015.33
Costea PI, Hildebrand F, Arumugam M et al (2017) Enterotypes in the landscape of gut microbial community composition. Nat Microbiol 3:8–16. https://doi.org/10.1038/s41564-017-0072-8
Leite GSF, Resende AS, West NP et al (2018) Probiotics and sports: a new magic bullet? Nutrition 60:152–160. https://doi.org/10.1016/j.nut.2018.09.023
Coqueiro AY, de Oliveira Garcia AB, Rogero MM et al (2017) Probiotic supplementation in sports and physical exercise: does it present any ergogenic effect? Nutr Health 23:239–249. https://doi.org/10.1177/0260106017721000
Funding
This research was funded by the Russian Ministry of Science and Higher Education, the projects (2018−2020) Nos. 0309-2018-0004 and AAAA-A17-117020210021-7.
Author information
Authors and Affiliations
Contributions
Conceptualization, G.S. and A.K.; methodology, T.A.; software, M.K.; validation, N.N.; formal analysis, M.K.; investigation, T.A. and G.S.; resources, V.V.; data curation, A.T. and N.N.; writing—original draft preparation, N.N.; writing−review and editing, V.V.; supervision, V.V.; project administration, M.K.; and funding acquisition, A.K. and V.V.
Corresponding author
Ethics declarations
Conflict of interest
Author A.K. is an employee of Bio-Vesta LLC. Any opinions or scientific interpretations expressed in this manuscript are those of the author and do not necessarily reflect the position or policy of Bio-Vesta LLC. Otherwise the authors declare that they have no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Consent to participate
All patients were duly informed and gave their consent to the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Naumova, N., Alikina, T., Tupikin, A. et al. Human Gut Microbiome Response to Short-Term Bifidobacterium-Based Probiotic Treatment. Indian J Microbiol 60, 451–457 (2020). https://doi.org/10.1007/s12088-020-00888-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-020-00888-1