Abstract
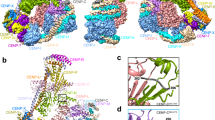
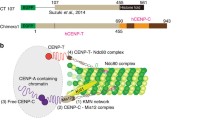
Equal distribution of DNA in mitosis requires the assembly of a large proteinaceous ensemble onto the centromeric DNA, called the kinetochore. With few exceptions, kinetochore specification is independent of the DNA sequence and is determined epigenetically by deposition at the centromeric chromatin of special nucleosomes containing an H3-related histone, CENP-A. Onto centromeric CENP-A chromatin is assembled the so-called constitutive centromere-associated network (CCAN) of 16 proteins distributed in several functional groups as follows: CENP-C, CENP-H/CENP-I/CENP-K/, CENP-L/CENP-M/CENP-N, CENP-O/CENP-P/CENP-Q/CENP-R/CENP-U(50), CENP-T/CENP-W, and CENP-S/CENP-X. One role of the CCAN is to recruit outer kinetochore components further, such as KNL1, the Mis12 complex, and the Ndc80 complex (KMN network) to which attach the spindle microtubules with their structural and regulatory proteins. Among the CENPs in CCAN, CENP-C and CENP-T are required in parallel for operational kinetochore specification and spindle attachment. This review presents discussion of the latest structural and functional data on CENP-A and CENPs from the CCAN as well as their interaction with the KMN network.




Similar content being viewed by others
References
Akiyoshi B, Sarangapani KK, Powers AF, Nelson CR, Reichow SL, Arellano-Santoyo H, Gonen T, Ranish JA, Asbury CL, Biggins S (2010) Tension directly stabilizes reconstituted kinetochore-microtubule attachments. Nature 468:576–579
Alonso A, Fritz B, Hasson D, Abrusan G, Cheung F, Yoda K, Radlwimmer B, Ladurner AG, Warburton PE (2007) Co-localization of CENP-C and CENP-H to discontinuous domains of CENP-A chromatin at human neocentromeres. Genome Biol 8:R148. doi:10.1186/gb-2007-8-7-r148
Amano M, Suzuki A, Hori T, Backer C, Okawa K, Cheeseman IM, Fukagawa T (2009) The CENP-S complex is essential for the stable assembly of outer kinetochore structure. J Cell Biol 186:173–182
Amaro AC, Samora CP, Holtackers R, Wang E, Kingston IJ, Alonso M, Lampson M, McAinsh AD, Meraldi P (2010) Molecular control of kinetochore-microtubule dynamics and chromosome oscillations. Nat Cell Biol 12:319–329
Amor DJ, Bentley K, Ryan J, Perry J, Wong L, Slater H, Choo KH (2004) Human centromere repositioning “in progress”. Proc Natl Acad Sci USA 101:6542–6547
Ando S, Yang H, Nozaki N, Okazaki T, Yoda K (2002) CENP-A, -B, and -C chromatin complex that contains the I-type alpha-satellite array constitutes the prekinetochore in HeLa cells. Mol Cell Biol 22:2229–2241
Bergmann JH, Rodríguez MG, Martins NM, Kimura H, Kelly DA, Masumoto H, Larionov V, Jansen LE, Earnshaw WC (2011) Epigenetic engineering shows H3K4me2 is required for HJURP targeting and CENP-A assembly on a synthetic human kinetochore. EMBO J 30:328–340
Bharadwaj R, Qi W, Yu H (2004) Identification of two novel components of the human NDC80 kinetochore complex. J Biol Chem 279:13076–13085
Bierie B, Edwin M, Melenhorst JJ, Hennighausen L (2004) The proliferation associated nuclear element (PANE1) is conserved between mammals and fish and preferentially expressed in activated lymphoid cells. Gene Expr Patterns 4:389–395
Black BE, Foltz DR, Chakravarthy S, Luger K, Woods VL Jr, Cleveland DW (2004) Structure determinants for generating centromeric chromatin. Nature 430:578–582
Black BE, Jensen LE, Maddox PS, Foltz DR, Desai AB, Shah JV, Cleveland DW (2007) Centromere identity maintained by nucleosomes assembled with histone H3 containing the CENP-A targeting domain. Mol Cell 25:309–322
Bloom KS, Amaya E, Carbon J, Clarke L, Hill A, Yeh E (1984) Chromatin conformation of yeast centromeres. J Cell Biol 99:1559–1568
Blower MD, Sullivan BA, Karpen GH (2002) Conserved organization of centromeric chromatin in flies and humans. Dev Cell 2:319–330
Brenner S, Pepper D, Berns MW, Tan E, Brinkley BR (1981) Kinetochore structure, duplication, and distribution in mammalian cells: analysis by human autoantibodies from scleroderma patients. J Cell Biol 91:95–102
Camahort R, Li B, Florens L, Swanson SK, Washburn MP, Gerton JL (2007) Scm3 is essential to recruit the histone h3 variant cse4 to centromeres and to maintain a functional kinetochore. Mol Cell 26:853–865
Camahort R, Shivaraju M, Mattingly M, Li B, Nakanishi S, Zhu D, Shilatifard A, Workman JL, Gerton JL (2009) Cse4 is part of an octameric nucleosome in budding yeast. Mol Cell 35:794–805
Cardinale S, Bergmann JH, Kelly D, Nakano M, Valdivia MM, Kimura H, Masumoto H, Larionov V, Earnshaw WC (2009) Hierarchical inactivation of a synthetic human kinetochore by a chromatin modifier. Mol Biol Cell 20:4194–4204
Carroll CW, Silva MC, Godek KM, Jansen LE, Straight AF (2009) Centromere assembly requires the direct recognition of CENP-A nucleosomes by CENP-N. Nat Cell Biol 11:896–902
Carroll CW, Milks KJ, Straight AF (2010) Dual recognition of CENP-A nucleosomes is required for centromere assembly. J Cell Biol 189:1143–1155
Castillo AG, Mellone BG, Partridge JF, Richardson W, Hamilton GL, Allshire RC, Pidoux AL (2007) Plasticity of fission yeast CENP-A chromatin driven by relative levels of histone H3 and H4. PLoS Genet 3:e121. doi:10.1371/journal.pgen.0030121
Cheeseman IM, Desai A (2008) Molecular architecture of the kinetochore–microtubule interface. Nat Rev Mol Cell Biol 9:33–46
Cheeseman IM, Niessen S, Anderson S, Hyndman F, Yates JR 3rd, Oegema K, Desai A (2004) A conserved protein network controls assembly of the outer kinetochore and its ability to sustain tension. Genes Dev 18:2255–2268
Cheeseman IM, Chappie JS, Wilson-Kubalek EM, Desai A (2006) The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell 127:983–997
Cheeseman IM, Hori T, Fukagawa T, Desai A (2008) KNL1 and the CENP-H/I/K complex coordinately direct kinetochore assembly in vertebrates. Mol Biol Cell 19:587–594
Chen Y, Baker RE, Keith KC, Harris K, Stoler A, Fitzgerald-Hayes M (2000) The N-terminus of the centromere H3-like protein Cse4p performs an essential function distinct from that of the histone fold domain. Mol Cell Biol 20:7037–7048
Choo KH (2001) Domain organization at the centromere and neocentromere. Dev Cell 1:165–177
Collins KA, Furuyama S, Biggins S (2004) Proteolysis contributes to the exclusive centromere localization of the yeast Cse4/CENP-A histone H3 variant. Curr Biol 14:1968–1972
Conde e Silva N, Black BE, Sivolob A, Filipski J, Cleveland DW, Prunell A (2007) CENP-A-containing nucleosomes: easier disassembly versus exclusive centromeric localization. J Mol Biol 370:555–573
Cooke CA, Bernat RL, Earnshaw WC (1990) CENP-B: A major human centromere protein located beneath the kinetochore. J Cell Biol 110:1475–1488
Cooke CA, Schaar B, Yen TJ, Earnshaw WC (1997) Localization of CENP-E in the fibrous corona and outer plate of mammalian kinetochores from prometaphase through anaphase. Chromosoma 106:446–455
Dalal Y, Wang H, Lindsay S, Henikoff S (2007) Tetrameric structure of centromeric nucleosomes in interphase Drosophila cells. PLoS Biol 5:e218. doi:10.1371/journal.pbio.0050218
DeLuca JG, Dong Y, Hergert P, Strauss J, Hickey JM, Salmon ED, McEwen BF (2005) Hec1 and nuf2 are core components of the kinetochore outer plate essential for organizing microtubule attachment sites. Mol Biol Cell 16:519–531
DeLuca JG, Gall WE, Ciferri C, Cimini D, Musacchio A, Salmon ED (2006) Kinetochore microtubule dynamics and attachment stability are regulated by Hec1. Cell 127:969–982
Desai A, Rybina S, Müller-Reichert T, Shevchenko A, Shevchenko A, Hyman A, Oegema K (2003) KNL-1 directs assembly of the microtubule-binding interface of the kinetochore in C. elegans. Genes Dev 17:2421–2435
du Sart D, Cancilla MR, Earle E, Mao J, Saffery R, Tainton KM, Kalitsis P, Martyn J, Barry AE, Choo A (1997) A functional neo-centromere formed through activation of a latent human centromere and consisting of non-alpha-satellite DNA. Nat Genet 16:144–153
Dunleavy EM, Pidoux AL, Monet M, Bonilla C, Richardson W, Hamilton GL, Ekwall K, McLaughlin PJ, Allshire RC (2007) A NASP (N1/N2)-related protein, Sim3, binds CENP-A and is required for its deposition at fission yeast centromeres. Mol Cell 28:1029–1044
Dunleavy EM, Roche D, Tagami H, Lacoste N, Ray-Gallet D, Nakamura Y, Daigo Y, Nakatani Y, Almouzni-Pettinotti G (2009) HJURP is a cell-cycle-dependent maintenance and deposition factor of CENP-A at centromeres. Cell 137:485–497
Earnshaw WC, Migeon BR (1985) Three related centromere proteins are absent from the inactive centromere of a stable isodicentric chromosome. Chromosoma 92:290–296
Earnshaw WC, Rothfield N (1985) Identification of a family of human centromere proteins using autoimmune sera from patients with scleroderma. Chromosoma 91:313–321
Earnshaw WC, Machlin PS, Bordwell BJ, Rothfield NF, Cleveland DW (1987) Analysis of anticentromere autoantibodies using cloned autoantigen CENP-B. Proc Natl Acad Sci USA 84:4979–4983
Erhardt S, Mellone BG, Betts CM, Zhang W, Karpen GH, Straight AF (2008) Genome-wide analysis reveals a cell cycle-dependent mechanism controlling centromere propagation. J Cell Biol 183:805–818
Folco HD, Pidoux AL, Urano T, Allshire RC (2008) Heterochromatin and RNAi are required to establish CENP-A chromatin at centromeres. Science 319:94–97
Foltz DR, Jansen LET, Black BE, Bailey AO, YateIII JR, Cleveland DW (2006) The human CENP-A centromeric nucleosome-associaed complex. Nat Cell Biol 8:458–469
Foltz DR, Jansen LE, Bailey AO, Yates JR 3rd, Bassett EA, Wood S, Black BE, Cleveland DW (2009) Centromere-specific assembly of CENP-A nucleosomes is mediated by HJURP. Cell 137:472–484
Fujita Y, Hayashi T, Kiyomitsu T, Toyoda Y, Kokubu A, Obuse C, Yanagida M (2007) Priming of centromere for CENP-A recruitment by human hMis18alpha, hMis18beta, and M18BP1. Dev Cell 12:17–30
Fukagawa T, Brown WR (1997) Efficient conditional mutation of the vertebrate CENP-C gene. Hum Mol Genet 6:2301–2308
Fukagawa T, Pendon C, Morris J, Brown W (1999) CENP-C is necessary but not sufficient to induce formation of a functional centromere. EMBO J 18:4196–4209
Fukagawa T, Mikami Y, Nishihashi A, Regnier V, Haraguchi T, Hiraoka Y, Sugata N, Todokoro K, Brown W, Ikemura T (2001) CENP-H, a constitutive centromere component, is required for centromere targeting of CENP-C in vertebrate cells. EMBO J 20:4603–4617
Fukagawa T, Nogami M, Yoshikawa M, Ikeno M, Okazaki T, Takami Y, Nakayama T, Oshimura M (2004) Dicer is essential for formation of the heterochromatin structure in vertebrate cells. Nat Cell Biol 6:784–791
Furuyama T, Henikoff S (2009) Centromeric nucleosomes induce positive DNA supercoils. Cell 138:104–113
Furuyama T, Dalal Y, Henikoff S (2006) Chaperone-mediated assembly of centromeric chromatin in vitro. Proc Natl Acad Sci USA 103:6172–6177
Gascoigne KE, Takeuchi K, Suzuki A, Hori T, Fukagawa T, Cheeseman IM (2011) Induced ectopic kinetochore assembly bypasses the requirement for CENP-A nucleosomes. Cell 145:410–422
Hayashi T, Fujita Y, Iwasaki O, Adachi Y, Takahashi K, Yanagida M (2004) Mis16 and Mis18 are required for CENP-A loading and histone deacetylation at centromeres. Cell 118:715–729
Heintz N, Sive HL, Roeder RG (1983) Regulation of human histone gene expression: kinetics of accumulation and changes in the rate of synthesis and in the half-lives of individual histone mRNAs during the HeLa cell cycle. Mol Cell Biol 3:539–550
Hemmerich P, Weidtkamp-Peters HC, Schmiedeberg L, Erliandri I, Diekmann S (2008) Dynamics of inner kinetochore assembly and maintenance in living cells. J Cell Biol 180:1101–1114
Hereford LM, Osley MA, Ludwig TR 2nd, McLaughlin CS (1981) Cell-cycle regulation of yeast histone mRNA. Cell 24:367–375
Heun P, Erhardt S, Blower MD, Weiss S, Skora AD, Karpen GH (2006) Mislocalization of the Drosophila centromere-specific histone CID promotes formation of functional ectopic kinetochores. Dev Cell 10:303–315
Hewawasam G, Shivaraju M, Mattingly M, Venkatesh S, Martin-Brown S, Florens L, Workman JL, Gerton JL (2010) Psh1 is an E3 ubiquitin ligase that targets the centromeric histone variant Cse4. Mol Cell 40:444–454
Hori T, Haraguchi T, Hiraoka Y, Kimura H, Fukagawa T (2003) Dynamic behavior of Nuf2-Hec1 complex that localizes to the centrosome and centromere and is essential for mitotic progression in vertebrate cells. J Cell Sci 116:3347–3362
Hori T, Amano M, Suzuki A, Backer CB, Welburn JP, Dong Y, McEwen BF, Shang WH, Suzuki E, Okawa K, Cheeseman IM, Fukagawa T (2008a) CCAN makes multiple contacts with centromeric DNA to provide distinct pathways to the outer kinetochore. Cell 135:1039–1052
Hori T, Okada M, Maenaka K, Fukagawa T (2008b) CENP-O class proteins form a stable complex and are required for proper kinetochore function. Mol Biol Cell 19:843–854
Howell BJ, McEwen BF, Canman JC, Hoffman DB, Farrar EM, Rieder CL, Salmon ED (2001) Cytoplasmic dynein/dynactin drives kinetochore protein transport to the spindle poles and has a role in mitotic spindle checkpoint inactivation. J Cell Biol 155:1159–1172
Howman EV, Fowler KJ, Newson AJ, Redward S, MacDonald AC, Kalitsis P, Choo KHA (2000) Early disruption of centromeric chromatin organization in centromere protein A (CenpA) null mice. Proc Natl Acad Sci USA 97:1148–1153
Izuta H, Ikeno M, Suzuki N, Tomonaga T, Nozaki N, Obuse C, Kisu Y, Goshima N, Nomura F, Nomura N, Yoda K (2006) Comprehensive analysis of the ICEN (Interphase Centromere Complex) components enriched in the CENP-A chromatin of human cells. Genes Cells 11:673–684
Janke C, Ortiz J, Lechner J, Shevchenko A, Shevchenko A, Magiera MM, Schramm C, Schiebel E (2001) The budding yeast proteins Spc24p and Spc25p interact with Ndc80p and Nuf2p at the kinetochore and are important for kinetochore clustering and checkpoint control. EMBO J 20:777–791
Jansen LE, Black BE, Foltz DR, Cleveland DW (2007) Propagation of centromeric chromatin requires exit from mitosis. J Cell Biol 176:795–805
Johnston K, Joglekar A, Hori T, Suzuki A, Fukagawa T, Salmon ED (2010) Vertebrate kinetochore protein architecture: protein copy number. J Cell Biol 189:937–943
Jokelainen PT (1967) The ultrastructure and spatial organization of the metaphase kinetochore in mitotic rat cells. J Ultrastruct Res 19:19–44
Kalitsis P, Fowler KJ, Earle E, Hill J, Choo KH (1998) Targeted disruption of mouse centromere protein C gene leads to mitotic disarray and early embryo death. Proc Natl Acad Sci USA 95:1136–1141
Kang YH, Park JE, Yu LR, Soung NK, Yun SM, Bang JK, Seong YS, Yu H, Garfield S, Veenstra TD, Lee KS (2006) Self-regulated Plk1 recruitment to kinetochores by the Plk1-PBIP1 interaction is critical for proper chromosome segregation. Mol Cell 24:409–422
Kang YH, Park C, Kim TS, Soung NK, Bang JK, Kim BY, Park JE, Lee KS (2011) Mammalian polo-like kinase 1-dependent regulation of the PBIP1-CENP-Q complex at kinetochores. J Biol Chem 286:19744–19757
Kato T, Sato N, Hayama S, Yamabuki T, Ito T, Miyamoto M, Kondo S, Nakamura Y, Daigo Y (2007) Activation of Holliday junction recognizing protein involved in the chromosomal stability and immortality of cancer cells. Cancer Res 67:8544–8553
Kingwell B, Rattner JB (1987) Mammalian kinetochore/centromere composition: a 50 kDa antigen is present in the mammalian kinetochore/centromere. Chromosoma 95:403–407
Kiyomitsu T, Obuse C, Yanagida M (2007) Human Blinkin/AF15q14 is required for chromosome alignment and the mitotic checkpoint through direct interaction with Bub1 and BubR1. Dev Cell 13:663–676
Kline SL, Cheeseman IM, Hori T, Fukagawa T, Desai A (2006) The human Mis12 complex is required for kinetochore assembly and proper chromosome segregation. J Cell Biol 173:9–17
Koike A, Nishikawa H, Wu W, Okada Y, Venkitaraman AR, Ohta T (2010) Recruitment of phosphorylated NPM1 to sites of DNA damage through RNF8-dependent ubiquitin conjugates. Cancer Res 70:6746–6756
Kwon MS, Hori T, Okada M, Fukagawa T (2007) CENP-C is involved in chromosome segregation, mitotic checkpoint function, and kinetochore assembly. Mol Biol Cell 18:2155–2168
Lagana A, Dorn JF, De Rop V, Ladouceur AM, Maddox AS, Maddox PS (2010) A small GTPase molecular switch regulates epigenetic centromere maintenance by stabilizing newly incorporated CENP-A. Nat Cell Biol 12:1186–1193
Liao H, Winkfein RJ, Mack G, Rattner JB, Yen TJ (1995) CENP-F is a protein of the nuclear matrix that assembles onto kinetochores at late G2 and is rapidly degraded after mitosis. J Cell Biol 130:507–518
Liao WT, Wang X, Xu LH, Kong QL, Yu CP, Li MZ, Shi L, Zeng MS, Song LB (2009) Centromere protein H is a novel prognostic marker for human nonsmall cell lung cancer progression and overall patient survival. Cancer 115:1507–1517
Liu ST, Hittle JC, Jablonski SA, Campbell MS, Yoda K, Yen TJ (2003) Human CENP-I specifies localization of CENP-F, MAD1 and MAD2 to kinetochores and is essential for mitosis. Nat Cell Biol 5:341–345
Liu ST, Rattner JB, Jablonski SA, Yen TJ (2006) Mapping the assembly pathways that specify formation of the trilaminar kinetochore plates in human cells. J Cell Biol 175:41–53
Luger K, Mäder AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389:251–260
Maddox PS, Hyndman F, Monen J, Oegema K, Desai A (2007) Functional genomics identifies a Myb domain-containing protein family required for assembly of CENP-A chromatin. J Cell Biol 176:757–763
Maresca TJ, Salmon ED (2009) Intrakinetochore stretch is associated with changes in kinetochore phosphorylation and spindle assembly checkpoint activity. J Cell Biol 184:373–381
Marshall OJ, Chueh AC, Wong LH, Choo KH (2008) Neocentromeres: new insights into centromere structure, disease development, and karyotype evolution. Am J Hum Genet 82:261–282
Masumoto H, Masukata H, Muro Y, Nozaki N, Okazaki T (1989) A human centromere antigen (CENP-B) interacts with a short specific sequence in alphoid DNA, a human centromeric satellite. J Cell Biol 109:1963–1973
McCleland ML, Gardner RD, Kallio MJ, Daum JR, Gorbsky GJ, Burke DJ, Stukenberg PT (2003) The highly conserved Ndc80 complex is required for kinetochore assembly, chromosome congression, and spindle checkpoint activity. Genes Dev 17:101–114
McCleland ML, Kallio MJ, Barrett-Wilt GA, Kestner CA, Shabanowitz J, Hunt DF, Gorbsky GJ, Stukenberg PT (2004) The vertebrate Ndc80 complex contains Spc24 and Spc25 homologs, which are required to establish and maintain kinetochore-microtubule attachment. Curr Biol 14:131–137
McClelland SE, Borusu S, Amaro AC, Winter JR, Belwal M, McAinsh AD, Meraldi P (2007) The CENP-A NAC/CAD kinetochore complex controls chromosome congression and spindle bipolarity. EMBO J 26:5033–5047
Measday V, Hailey DW, Pot I, Givan SA, Hyland KM, Cagney G, Fields S, Davis TN, Hieter P (2002) Ctf3p, the Mis6 budding yeast homolog, interacts with Mcm22p and Mcm16p at the yeast outer kinetochore. Genes Dev 16:101–113
Mellone BG, Allshire RC (2003) Stretching it: putting the CEN(P-A) in centromere. Curr Opin Genet Dev 13:191–198
Mellone BG, Grive KJ, Shteyn V, Bowers SR, Oderberg I, Karpen GH (2011) Assembly of Drosophila Centromeric Chromatin Proteins during Mitosis. PLoS Genet 7:e1002068
Milks KJ, Moree B, Straight AF (2009) Dissection of CENP-C-directed centromere and kinetochore assembly. Mol Biol Cell 20:4246–4255
Minoshima Y, Hori T, Okada M, Kimura H, Haraguchi T, Hiraoka Y, Kawashima T, Kitamura T, Fukagawa T (2005) The constitutive centromere component CENP-50 is required for recovery from spindle damage. Mol Cell Biol 25:10315–10328
Mizuguchi G, Xiao H, Wisniewski J, Smith MM, Wu C (2007) Nonhistone Scm3 and histones CenH3-H4 assemble the core of centromere-specific nucleosomes. Cell 129:1153–1164
Moreno-Moreno O, Torras-Llort M, Azorín F (2006) Proteolysis restricts localization of CID, the centromere-specific histone H3 variant of Drosophila, to centromeres. Nucleic Acids Res 34:6247–6255
Morey L, Barnes K, Chen Y, Fitzgerald-Hayes M, Baker RE (2004) The histone fold domain of Cse4 is sufficient for CEN targeting and propagation of active centromeres in budding yeast. Eukaryot Cell 3:1533–1543
Moroi Y, Peebles C, Fritzler MJ, Steigerwald J, Tan EM (1980) Autoantibody to centromere (kinetochore) in scleroderma sera. Proc Natl Acad Sci USA 77:1627–1631
Nakano M, Cardinale S, Noskov VN, Gassmann R, Vagnarelli P, Kandels-Lewis S, Larionov V, Earnshaw WC, Masumoto H (2008) Inactivation of a human kinetochore by specific targeting of chromatin modifiers. Dev Cell 14:507–522
Nakaseko Y, Adachi Y, Funahashi S, Niwa O, Yanagida M (1986) Chromosome walking shows a highly homologous repetitive sequence present in all the centromere regions of fission yeast. EMBO J 5:1011–1012
Nakazawa N, Nakamura T, Kokubu A, Ebe M, Nagao K, Yanagida M (2008) Dissection of the essential steps for condensin accumulation at kinetochores and rDNAs during fission yeast mitosis. J Cell Biol 180:1115–1131
Narlikar GJ (2010) A proposal for kinetic proof reading by ISWI family chromatin remodeling motors. Curr Opin Chem Biol 14:660–665
Nekrasov VS, Smith MA, Peak-Chew S, Kilmartin JV (2003) Interactions between centromere complexes in Saccharomyces cerevisiae. Mol Biol Cell 14:4931–4946
Nishihashi A, Haraguchi T, Hiraoka Y, Ikemura T, Regnier V, Dodson H, Earnshaw WC, Fukagawa T (2002) CENP-I is essential for centromere function in vertebrate cells. Dev Cell 2:463–476
Obuse C, Iwasaki O, Kiyomitsu T, Goshima G, Toyoda Y, Yanagida M (2004a) A conserved Mis12 centromere complex is linked to heterochromatic HP1 and outer kinetochore protein Zwint-1. Nat Cell Biol 6:1135–1141
Obuse C, Yang H, Nozaki N, Goto S, Okazaki T, Yoda K (2004b) Proteomics analysis of the centromere complex from HeLa interphase cells: UV-damaged DNA binding protein-1 (DDB-1) is a component of the CEN-complex, while BMI-1 is transiently colocalized with the centromeric region in interphase. Genes Cells 9:105–120
Ohta S, Bukowski-Wills JC, Sanchez-Pulido L, Alves Fde L, Wood L, Chen ZA, Platani M, Fischer L, Hudson DF, Ponting CP, Fukagawa T, Earnshaw WC, Rappsilber J (2010) The protein composition of mitotic chromosomes determined using multiclassifier combinatorial proteomics. Cell 142:810–821
Okada M, Cheeseman IM, Hori T, Okawa K, McLeod IX, YateIII JR, Desai A, Fukagawa T (2006) The CENP-H–I complex is required for the efficient incorporation of newly syntehseized CENP-A. Nat Cell Biol 8:446–457
Okada M, Okawa K, Isobe T, Fukagawa T (2009) CENP-H-containing complex facilitates centromere deposition of CENP-A in cooperation with FACT and CHD1. Mol Biol Cell 20:3986–3995
Osakabe A, Tachiwana H, Matsunaga T, Shiga T, Nozawa RS, Obuse C, Kurumizaka H (2010) Nucleosome formation activity of human somatic nuclear autoantigenic sperm protein (sNASP). J Biol Chem 285:11913–11921
Palmer DK, O'Day K, Wener MH, Andrews BS, Margolis RL (1987) A 17-kD centromere protein (CENP-A) copurifies with nucleosome core particles and with histones. J Cell Biol 104:805–815
Palmer DK, O'Day K, Trong HL, Charbonneau H, Margolis RL (1991) Purification of the centromere-specific protein CENP-A and demonstration that it is a distinctive histone. Proc Natl Acad Sci USA 88:3734–3738
Pearson CG, Yeh E, Gardner M, Odde D, Salmon ED, Bloom K (2004) Stable kinetochore-microtubule attachment constrains centromere positioning in metaphase. Curr Biol 14:1962–1967
Perpelescu M, Nozaki N, Obuse C, Yang H, Yoda K (2009) Active establishment of centromeric CENP-A chromatin by RSF complex. J Cell Biol 185:397–407
Pidoux AL, Choi ES, Abbott JK, Liu X, Kagansky A, Castillo AG, Hamilton GL, Richardson W, Rappsilber J, He X, Allshire RC (2009) Fission yeast Scm3: A CENP-A receptor required for integrity of subkinetochore chromatin. Mol Cell 33:299–311
Pimpinelli S, Goday C (1989) Unusual kinetochores and chromatin diminution in Parascaris. Trends Genet 5:310–315
Pluta AF, Mackay AM, Ainsztein AM, Goldberg IG, Earnshaw WC (1996) The centromere: hub of chromosomal activities. Science 270:1591–1594
Przewloka MR, Venkei Z, Bolanos-Garcia VM, Debski J, Dadlez M, Glover DM (2011) CENP-C is a structural platform for kinetochore assembly. Curr Biol 21:399–405
Ranjitkar P, Press MO, Yi X, Baker R, MacCoss MJ, Biggins S (2010) An E3 ubiquitin ligase prevents ectopic localization of the centromeric histone H3 variant via the centromere targeting domain. Mol Cell 40:455–464
Rattner JB, RaoA FMJ, Valencia DW, Yen TJ (1993) CENP-F is a ca. 400 kDa kinetochore protein that exhibits a cell-cycle dependent localization. Cell Motil Cytoskeleton 26:214–226
Ravi M, Chan SW (2010) Haploid plants produced by centromere-mediated genome elimination. Nature 464:615–618
Regnier V, Vagnarelli P, Fukagawa T, Zerjal T, Burns E, Trouche D, Earnshaw W, Brown W (2005) CENP-A is required for accurate chromosome segregation and sustained kinetochore association of BubR1. Mol Cell Biol 25:3967–3981
Renou JP, Bierie B, Miyoshi K, Cui Y, Djiane J, Reichenstein M, Shani M, Henninghauzen L (2003) Identification of genes differentially expressed in mouse mammary epithelium transformed by an activated b-catenin. Oncogene 22:4594–4610
Ribeiro SA, Vagnarelli P, Dong Y, Hori T, McEwen BF, Fukagawa T, Flors C, Earnshaw WC (2010) A super-resolution map of the vertebrate kinetochore. Proc Natl Acad Sci USA 107:10484–10489
Rieder CL, Salmon ED (1998) The vertebrate cell kinetochore and its role during mitosis. Trends Cell Biol 8:310–318
Saffery R, Irvine DV, Griffiths B, Kalitsis P, Worderman L, Choo KH (2000) Human centromeres and neocentromeres show identical distribution patterns of >20 functionally imprtant kinetochore-associated proteins. Hum Mol Genet 9:175–185
Saffery R, Sumer H, Hassan S, Wong LH, Craig JM, Todokoro K, Anderson M, Stafford A, Choo KH (2003) Transcription within a functional human centromere. Mol Cell 12:509–516
Saitoh H, Tomkiel J, Cooke CA, Ratrie H III, Maurer M, Rothfield NF, Earnshaw WC (1992) CENP-C, an autoantigen in scleroderma, is a component of the human inner kinetochore plate. Cell 70:115–125
Saitoh S, Ishii K, Kobayashi Y, Takahashi K (2005) Spindle checkpoint signaling requires the mis6 kinetochore subcomplex, which interacts with mad2 and mitotic spindles. Mol Biol Cell 16:3666–3677
Sanchez-Pulido L, Pidoux AL, Ponting CP, Allshire RC (2009) Common ancestry of the CENP-A chaperones Scm3 and HJURP. Cell 137:1173–1174
Schuh M, Lehner CF, Heidmann S (2007) Incorporation of Drosophila CID/CENP-A and CENP-C into centromeres during early embryonic anaphase. Curr Biol 17:237–243
Screpanti E, De Antoni A, Alushin GM, Petrovic A, Melis T, Nogales E, Musacchio A (2011) Direct binding of Cenp-C to the Mis12 complex joins the inner and outer kinetochore. Curr Biol 21:391–398
Sekulic N, Bassett EA, Rogers DJ, Black BE (2010) The structure of (CENP-A-H4)(2) reveals physical features that mark centromeres. Nature 467:347–351
Shang WH, Hori T, Toyoda A, Kato J, Popendorf K, Sakakibara Y, Fujiyama A, Fukagawa T (2010) Chickens possess centromeres with both extended tandem repeats and short non-tandem-repetitive sequences. Genome Res 20:1219–1228
Shelby RD, Vafa O, Sullivan KF (1997) Assembly of CENP-A into centromeric chromatin requires a cooperative array of nucleosomal DNA contact sites. J Cell Biol 136:501–513
Shelby RD, Monier K, Sullivan KF (2000) Chromatin assembly at kinetochores is uncoupled from DNA replication. J Cell Biol 151:1113–1118
Shigeishi H, Higashikawa K, Ono S, Mizuta K, Ninomiya Y, Yoneda S, Taki M, Kamata N (2006) Increased expression of CENP-H gene in human oral squamous cell carcinomas harboring high-proliferative activity. Oncol Rep 16:1071–1075
Shiroiwa Y, Hayashi T, Fujita Y, Villar-Briones A, Ikai N, Takeda K, Ebe M, Yanagida M (2011) Mis17 is a regulatory module of the Mis6–Mal2–Sim4 centromere complex that is required for the recruitment of CenH3/CENP-A in fission yeast. PLoS One 6:e17761
Shuaib M, Ouararhni K, Dimitrov S, Hamiche A (2010) HJURP binds CENP-A via a highly conserved N-terminal domain and mediates its deposition at centromeres. Proc Natl Acad Sci USA 107:1349–1354
Singh TR, Saro D, Ali AM, Zheng XF, Du CH, Killen MW, Sachpatzidis A, Wahengbam K, Pierce AJ, Xiong Y, Sung P, Meetei AR (2010) MHF1-MHF2, a histone-fold-containing protein complex, participates in the Fanconi anemia pathway via FANCM. Mol Cell 37:879–886
Stoler S, Keith KC, Curnick KE, Fitzgerald-Hayes M (1995) A mutation in CSE4, an essential gene encoding a novel chromatin-associated protein in yeast, causes chromosome nondisjunction and cell cycle arrest at mitosis. Genes Dev 9:573–586
Stoler S, Rogers K, Weitze S, Morey L, Fitzgerald-Hayes M, Baker RE (2007) Scm3, an essential Saccharomyces cerevisiae centromere protein required for G2/M progression and Cse4 localization. Proc Natl Acad Sci USA 104:10571–10576
Sugata N, Munekata E, Todokoro K (1999) Characterization of a novel kinetochore protein, CENP-H. J Biol Chem 274:27343–27346
Sugata N, Li S, Earnshaw WC, Yen TJ, Yoda K, Masumoto H, Munekata E, Warburton PE, Todokoro K (2000) Human CENP-H multimers colocalize with CENP-A and CENP-C at active centromere-kinetochore complexes. Hum Mol Genet 9:2919–2926
Sugimoto K, Yata H, Muro Y, Himeno M (1994) Human centromere protein C (CENP-C) is a DNA-binding protein which possesses a novel DNA-binding motif. J Biochem 116:877–881
Sugimoto K, Kuriyama K, Shibata A, Himeno M (1997) Characterization of internal DNA-binding and C-terminal dimerization domains of human centromere/kinetochore autoantigen CENP-C in vitro: role of DNA-binding and self-associating activities in kinetochore organization. Chromosome Res 5:132–141
Sullivan BA, Karpen GH (2004) Centromeric chromatin exhibits a histone modification pattern that is distinct from both euchromatin and heterochromatin. Nat Struc Molec Biol 11:1076–1083
Sullivan KF, Hechenberger M, Masri K (1994) Human CENP-A contains a histone H3 related histone fold domain that is required for targeting to the centromere. J Cell Biol 127:581–592
Sullivan BA, Blower MD, Karpen GH (2001) Determining centromere identity: cyclical stories and forking paths. Nat Rev Genet 2:584–596
Suzuki A, Hori T, Nishino T, Usukura J, Miyagi A, Morikawa K, Fukagawa T (2011) Spindle microtubules generate tension-dependent changes in the distribution of inner kinetochore proteins. J Cell Biol 193:125–140
Tadeu AM, Ribeiro S, Johnston J, Goldberg I, Gerloff D, Earnshaw WC (2008) CENP-V is required for centromere organization, chromosome alignment and cytokinesis. EMBO J 27:2510–2522
Tagami H, Ray-Gallet D, Almouzni G, Nakatani Y (2004) Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 116:51–61
Takahashi K, Chen ES, Yanagida M (2000) Requirement of Mis6 centromere connector for localizing a CENP-A-like protein in fission yeast. Science 288:2215–2219
Takahashi K, Takayama Y, Masuda F, Kobayashi Y, Saitoh S (2005) Two distinct pathways responsible for the loading of CENP-A to centromeres in the fission yeast cell cycle. Philos Trans R Soc Lond B Biol Sci 360:595–606
Takayama Y, Sato H, Saitoh S, Ogiyama Y, Masuda F, Takahashi K (2008) Biphasic incorporation of centromeric histone CENP-A in fission yeast. Mol Biol Cell 19:682–690
Tomkiel J, Cooke CA, Saitoh H, Bernat RL, Earnshaw WC (1994) CENP-C is required for maintaining proper kinetochore size and for a timely transition to anaphase. J Cell Biol 125:531–545
Tomonaga T, Matsushita K, Yamaguchi S, Oohasi S, Shimada H, Ochiai T, Yoda K, Nomura F (2003) Overexpression and mistargeting of centromere protein-A in human primary colorectal cancer. Cancer Res 63:3511–3516
Tomonaga T, Matsushita K, Ishibashi M, Nezu M, Shimada H, Ochiai T, Yoda K, Nomura F (2005) Centromere protein H is up-regulated in primary human colorectal cancer and its overexpression induces aneuploidy. Cancer Res 65:4683–4689
Uchida KS, Takagaki K, Kumada K, Hirayama Y, Noda T, Hirota T (2009) Kinetochore stretching inactivates the spindle assembly checkpoint. J Cell Biol 184:383–390
Vafa O, Sullivan K (1997) Chromatin containing CENP-A and a-satellite DNA is a major component of the inner kinetochore plate. Curr Biol 7:897–900
Van Hooser AA, Ouspenski II, Gregson HC, Starr DA, Yen TJ, Goldberg ML, Yokomori K, Earnshaw WC, Sullivan KF, Brinkley BR (2001) Specification of kinetochore-forming chromatin by the histone H3 variant CENP-A. J Cell Sci 114:93529–93542
Ventura M, Weigl S, Carbone L, Cardone MF, Misceo D, Teti M, D'Addabbo P, Wandall A, Björck E, de Jong PJ, She X, Eichler EE, Archidiacono N, Rocchi M (2004) Recurrent sites for new centromere seeding. Genome Res 14:1696–1703
Vermaak D, Hayden HS, Henikoff S (2002) Centromere targeting element within the histone fold domain of Cid. Mol Cell Biol 22:7553–7561
Volpe TA, Kidner C, Hall IM, Teng G, Grewal SI, Martienssen RA (2002) Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297:1833–1837
Wade CM, Giulotto E, Sigurdsson S, Zoli M, Gnerre S, Imsland F, Lear TL, Adelson DL, Bailey E, Bellone RR, Blöcker H, Distl O, Edgar RC, Garber M, Leeb T, Mauceli E, MacLeod JN, Penedo MC, Raison JM et al (2009) Genome sequence, comparative analysis, and population genetics of the domestic horse. Science 326:865–867
Wan X, O'Quinn RP, Pierce HL, Joglekar AP, Gall WE, DeLuca JG, Carroll CW, Liu ST, Yen TJ, McEwen BF, Stukenberg PT, Desai A, Salmon ED (2009) Protein architecture of the human kinetochore microtubule attachment site. Cell 137:672–684
Warburton PE, Cooke CA, Bourassa S, Vafa O, Sullivan BA, Stetten G, Gimelli G, Warburton D, Tyler-Smith C, Sullivan KF, Poirier GG, Earnshaw WC (1997) Immunolocalization of CENP-A suggests a distinct nucleosome structure at the inner kinetochore plate of active centromere. Curr Biol 7:901–904
Wieland G, Orthaus S, Ohndorf S, Diekmann S, Hemmerich P (2004) Functional complementation of human centromere protein A (CENP-A) by Cse4p from Saccharomyces cerevisiae. Mol Cell Biol 24:6620–6630
Wiens GR, Sorger PK (1998) Centromeric chromatin and epigenetic effects in kinetochore assembly. Cell 93:313–316
Wigge PA, Kilmartin JV (2001) The Ndc80p complex from Saccharomyces cerevisiae contains conserved centromere components and has a function in chromosome segregation. J Cell Biol 152:349–360
Willard HF (1990) Centromeres of mammalian chromosomes. Trends Genet 6:410–416
Willard HF, Waye JS (1987) Hierarchical order in chromosome-specific human alpha satellite DNA. Trends Genet 3:192–198
Williams JS, Hayashi T, Yanagida M, Russell P (2009) Fission yeast Scm3 mediates stable assembly of Cnp1/CENP-A into centromeric chromatin. Mol Cell 33:287–298
Xu M, Long C, Chen X, Huang C, Chen S, Zhu B (2010) Partitioning of histone H3–H4 tetramers during DNA replication-dependent chromatin assembly. Science 328:94–98
Yamashita A, Ito M, Takamatsu N, Shiba T (2000) Characterization of Solt, a novel SoxLZ/Sox6 binding protein expressed in adult mouse testis. FEBS Lett 481:147–151
Yan Z, Delannoy M, Ling C, Daee D, Osman F, Muniandy PA, Shen X, Oostra AB, Du H, Steltenpool J, Lin T, Schuster B, Décaillet C, Stasiak A, Stasiak AZ, Stone S, Hoatlin ME, Schindler D, Woodcock CL, Joenje H, Sen R, de Winter JP, Li L, Seidman MM, Whitby MC, Myung K, Constantinou A, Wang W (2010) A histone-fold complex and FANCM form a conserved DNA-remodeling complex to maintain genome stability. Mol Cell 37:865–878
Yang CH, Tomkiel J, Saitoh H, Johnson DH, Earnshaw WC (1996) Identification of overlapping DNA-binding and centromere-targeting domains in the human kinetochore protein CENP-C. Mol Cell Biol 16:3576–3586
Yao X, Anderson KL, Cleveland DW (1997) The microtubule-dependent motor centromere-associated protein E (CENP-E) is an integral component of kinetochore corona fibers that link centromeres to spindle microtubules. J Cell Biol 139:435–447
Yen TJ, Compton DA, Wise D, Zinkowski RP, Brinkley BR, Earnshaw WC, Cleveland DW (1991) CENP-E, a novel human centromere-associated protein required for progression from metaphase to anaphase. EMBO J 10:1245–1254
Yoda K, Kitagawa K, Masumoto H, Muro Y, Okazaki T (1992) A human centromere protein, CENP-B, has a DNA binding domain containing four potential a-helices at the NH2 terminus, which is separable from dimerizing activity. J Cell Biol 119:1413–1427
Yoda K, Ando S, Morishita S, Houmura K, Hashimoto K, Takeyasu K, Okazaki T (2000) Human centromere protein A (CENP-A) can replace histone H3 in nucleosome reconstitution in vitro. Proc Natl Acad Sci USA 97:7266–7271
Zeitlin SG, Baker NM, Chapados BR, Soutoglou E, Wang JY, Berns MW, Cleveland DW (2009) Double-strand DNA breaks recruit the centromeric histone CENP-A. Proc Natl Acad Sci USA 106:1562–1567
Zhou Z, Feng H, Zhou BR, Ghirlando R, Hu K, Zwolak A, Miller Jenkins LM, Xiao H, Tjandra N, Wu C, Bai Y (2011) Structural basis for recognition of centromere histone variant CenH3 by the chaperone Scm3. Nature 472:234–237
Acknowledgments
The authors are very grateful to all members of the Fukagawa Lab for useful discussions and to Bill Earnshaw for useful comments. This work was supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan to M.P. and T.F. and the Cabinet Office, Government of Japan through its “Funding Program for the Next Generation World-Leading Researchers” to T.F.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by E. Nigg
Rights and permissions
About this article
Cite this article
Perpelescu, M., Fukagawa, T. The ABCs of CENPs. Chromosoma 120, 425–446 (2011). https://doi.org/10.1007/s00412-011-0330-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-011-0330-0