Abstract
CCL17 is produced by conventional dendritic cells, signals through CCR4 on regulatory T (Treg) cells and drives atherosclerosis by suppressing Treg functions through yet undefined mechanisms. Here we show that conventional dendritic cells from CCL17-deficient mice display a pro-tolerogenic phenotype and transcriptome that is not phenocopied in mice lacking its cognate receptor CCR4. In the plasma of CCL17-deficient mice, CCL3 was the only decreased cytokine/chemokine. We found that CCL17 signaled through CCR8 as an alternate high-affinity receptor, which induced CCL3 expression and suppressed Treg functions in the absence of CCR4. Genetic ablation of CCL3 and CCR8 in CD4+ T cells reduced CCL3 secretion, boosted FoxP3+ Treg numbers and limited atherosclerosis. Conversely, CCL3 administration exacerbated atherosclerosis and restrained Treg differentiation. In symptomatic versus asymptomatic human carotid atheroma, CCL3 expression was increased, whereas FoxP3 expression was reduced. Together, we identified a non-canonical chemokine pathway whereby CCL17 interacts with CCR8 to yield a CCL3-dependent suppression of atheroprotective Treg cells.
Similar content being viewed by others
Main
Atherosclerosis is a lipid-driven chronic inflammatory disease of the arterial wall, underlying most cardiovascular diseases (CVDs)1. Dendritic cells (DCs) have been identified in healthy and inflamed arterial intima of mice and humans2,3,4 and advanced human plaques contain an increased number of DCs in clusters with T cells5. The chemokine CCL17 (TARC/thymus and activation-regulated chemokine) was found to be elevated in patients with CVD6 and atopic dermatitis, who are prone to develop CVD7,8. An intronic single-nucleotide polymorphism (rs223828) is associated with higher serum CCL17 and elevated CVD risk in humans9 and mouse studies revealed a proinflammatory role of CCL17 in atherosclerosis10 and colitis11. CCL17 is primarily expressed by a subset of antigen-presenting MHCII+ conventional DCs (cDCs), which migrates to draining lymph nodes (LNs) to prime naive T cells and plays a crucial role in the migration of various T cell subsets, including CD4+ T cells and Treg cells12,13,14. Treg cells use multiple effector mechanisms to modulate immune responses and to guard the balance between immune activation and tolerance toward self-antigens15. Naturally arising CD4+CD25+ Treg cells limit atherosclerosis development and have been implicated in the regression of established atherosclerotic lesions by licensing pro-resolving macrophage functions16,17. We found that CCL17 controls Treg homeostasis, restraining their expansion and thereby promoting atherosclerosis10. Accordingly, genetic deficiency or antibody blockade of CCL17 reduced atheroprogression by facilitating Treg expansion and survival in lymphoid organs, leading to Treg expansion10; however, the precise mechanisms, by which DC-derived CCL17 controls Treg homeostasis, remain to be elucidated, namely the relevant soluble mediators or the receptors involved.
To date, CCR4 is the only cognate signaling receptor for CCL17 identified and known to contribute to the recruitment and in vivo functions of Treg cells18. Yet, CCR4 deficiency did not phenocopy the effects on Treg cells and protection from atherosclerosis seen in CCL17-deficient mice10. Analogous findings were obtained in a model of atopic dermatitis, where the inflammatory burden was reduced in mice lacking CCL17 but not CCR4 (ref. 19). Consistently, deficiency in DC-derived CCL17 was protective against intestinal inflammation in a mouse model of colitis by creating a cytokine milieu that facilitated Treg expansion and, likewise, did not require CCR4 (ref. 11). In conjunction, these results suggest the existence of an alternative CCR4-independent receptor pathway triggered by CCL17. Here, we provide unequivocal evidence that CCL17 signals via CCR8 as an alternative high-affinity receptor expressed on T cell and DC subsets, harnessing the autocrine and paracrine release of CCL3 to mediate suppression of atheroprotective Treg cells via its receptor CCR1.
Results
CCR4 does not mediate the effects of CCL17 on atherogenesis
Mice with a targeted replacement of the Ccl17 gene by the enhanced green fluorescent protein gene (eGFP; Apoe−/−Ccl17e/e mice) displayed a reduced atherosclerotic lesion size in the aortic root, thoraco-abdominal aorta and aortic arch but unchanged lesional macrophage, smooth muscle cell (SMC) and collagen content after 12 weeks of western-type diet (WD), when compared to Apoe−/− littermate controls (Extended Data Fig. 1a–g). This was related to a higher frequency of CD25+Foxp3+CD3+CD4+ Treg cells in para-aortic LNs, spleen, axillary and inguinal LNs of Apoe−/−Ccl17e/e mice, compared to controls (Extended Data Fig. 1h–l). In contrast, mice lacking the canonical CCL17 receptor CCR4 did not phenocopy the effects observed in CCL17-deficient mice. Neither hematopoietic Ccr4 deficiency10 nor systemic deletion in Apoe−/−Ccr4−/− mice altered atherosclerotic lesion size, composition or Treg frequencies after 12 weeks of WD, compared to controls, and the decrease in apoptotic Treg cells in LNs of CCL17-deficient mice10 was not found in LNs of CCR4-deficient mice (Fig. 1a–f and Extended Data Fig. 2a–g). Lipid levels, circulating leukocyte and thrombocyte counts did not differ between Apoe−/−Ccl17e/e, Apoe−/−Ccr4−/− and respective control mice (Supplementary Tables 1 and 2) and thus did not explain the observed phenotype. Using recombinant CCL17, CCL22 and the CCR4 inhibitor C021 in Transwell assays, we confirmed CCR4-dependent chemotaxis of CD4+ T cells isolated from Apoe−/− mice and from human peripheral blood mononuclear cells (PBMCs) in vitro, as C021 abrogated CCL22-induced migration and inhibited CCL17-induced migration (Extended Data Fig. 2h,i). Together, these data indicate that an alternative signaling pathway is involved in mediating the effects of CCL17, a notion reinforced by investigating Treg homeostasis. Co-culturing CD4+CD62L+ T cells from Apoe−/− and Apoe−/−Ccr4−/− mice with ex vivo isolated cDCs revealed an increased differentiation of CD4+CD25+Foxp3+ Treg cells in the presence of Apoe−/−Ccl17e/e cDCs but not Apoe−/−Ccr4−/− or Apoe−/− cDCs (Fig. 1g). To explore whether CCL17-deficient cDCs differentially express putative mediators of this effect, we sorted CD45+CD11c+CD3−CD19− cells from LNs of Apoe−/−Ccl17wt/e or Apoe−/−Ccl17e/e mice and performed single-cell RNA sequencing (scRNA-seq) (GSE200862). Our analysis identified seven different DC clusters, with CCL17-expressing eGFP+ cells almost exclusively located among CCR7+ cDCs (Fig. 1h and Extended Data Fig. 3a–g). In CCL17-deficient samples, CCR7+ cDCs were enriched and had a higher tolerogenic score compared to other DC clusters (Fig. 1i). A tolerogenic profile was defined by high expression of Aldh1a2, Cd83 and Cd274 in CCR7+ cDCs (Fig. 1j and Extended Data Fig. 3d–g) and analysis of a gene set defining immunogenic and tolerogenic cell properties (Supplementary Table 3). While the tolerogenic score was similar in hetero- and homozygous samples, the number of CCR7+ tolerogenic cDCs was higher in CCL17-deficient mice (Fig. 1j,k), showing that more cDCs acquire a tolerogenic phenotype in the absence of CCL17. Accordingly, flow cytometry of cDCs in aortic LNs of Apoe−/− and Apoe−/−Ccl17e/e mice uncovered a higher percentage of CD83+, CCR7+ and CD83+CCR7+ cDCs (Extended Data Fig. 3h–j). Deletion of CD83 in cDCs confers a proinflammatory DC phenotype driving antigen-dependent T cell proliferation, type 17 helper T (TH17) cell commitment but impairing Treg suppressive capacity and resolving mechanisms20. Gene set variation analysis (GSVA) further revealed an enrichment of proinflammatory pathways in CCL17-competent cDCs (Extended Data Fig. 3k–m), in line with an atheroprotective role of CCR7 found in Apoe−/− mice21.
a, Experimental scheme of WD feeding. b, Representative sections and quantification of lesion area after Oil-Red-O staining (ORO) for lipid deposits in the aortic root of Apoe−/− (n = 16) or Apoe−/−Ccr4−/− (n = 20) mice. Scale bar, 500 µm. c, Quantification of lesion area after ORO in the thoraco-abdominal aorta of Apoe−/− (n = 5) or Apoe−/−Ccr4−/− (n = 19) mice. d, Atherosclerotic lesion size quantified by hematoxylin and eosin (H&E) staining in the aortic arches of Apoe−/− (n = 16) or Apoe−/−Ccr4−/− (n = 19) mice. e,f, Representative dot plots and flow cytometric quantification of CD45+CD3+CD4+CD25+FoxP3+ Treg cells in para-aortic LNs (e) and spleen (f) of Apoe−/− (e, n = 16; f, n = 16) or Apoe−/−Ccr4−/− (e, n = 19; f, n = 20) mice. g, Co-culture of CD45+CD11c+MHCII+ DCs sorted from Apoe−/−, Apoe−/−Ccl17e/e or Apoe−/−Ccr4−/− mice with splenic CD4+CD62L+ T cells isolated from Apoe−/− or Apoe−/−Ccr4−/− mice. After 72 h, the abundance of CD45+CD3+CD4+CD25+FoxP3+ Treg cells was determined by flow cytometry (individual number of experiments per bar from left to right n = 7, 5, 4, 6 and 5). Besides the cells, no other factors were added to the culture. h–j, CD45+CD3−CD19−CD11c+ cells were sorted from pooled LNs of Apoe−/−Ccl17wt/e or Apoe−/−Ccl17e/e mice on chow diet. Uniform Manifold Approximation and Projection (UMAP) projection of 4,731 single cells colored by inferred cell types consisting of seven distinct DC clusters (h). Depicted are eGFP+ cell counts in CCR7+ DCs or other (CCR7−) DCs (percentage of total numbers) and proportions of four distinct CCR7+ DC clusters among all single cells (CD45+CD3−CD19−CD11c+) (bottom) (i). j,k, Tolerogenic score calculated for four distinct CCR7+ DC clusters and other (CCR7−) DC clusters based on the top 20 genes differentially expressed between the groups (top) as (1 + mean (top 20 upregulated tolerogenic genes))/(1 + mean(top 20 upregulated immunogenic genes)) (j). k, Table depicts cell counts of four CCR7+ DC and other DC clusters. LNs for scRNA-seq were obtained from Apoe−/−Ccl17wt/e (n = 8) and Apoe−/−Ccl17e/e (n = 6). Data represent mean ± s.e.m. (a–k). Two-sided P values as indicated and as analyzed by unpaired Student’s t-test (b,e,f), Mann–Whitney U-test (c,d) or Kruskal–Wallis H with Dunn’s post hoc test (g,j). Differences in proportion in DC clusters between Apoe−/−Ccl17wt/e and Apoe−/−Ccl17e/e were computed by Pearson’s chi-squared test (k).
CCL17 induces CCL3 release independent of CCR4 expression
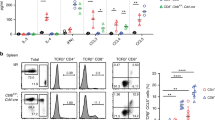
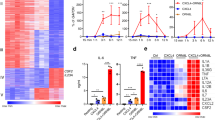
Because CCL17-deficient mice displayed increased numbers of tolerogenic cDCs, we performed an unbiased screen for differentially regulated inflammatory mediators. A multiplex bead array for cytokines and chemokines identified only CCL3 to be substantially reduced in plasma of Apoe−/−Ccl17e/e mice versus Apoe−/− controls after 12 weeks of WD (Fig. 2a,b and Extended Data Table 1), corresponding with decreased lesion size and increased Treg numbers (Extended Data Fig. 1). In contrast, CCL3 plasma levels in Apoe−/−Ccr4−/− mice were unaffected (Fig. 2a,c), consistent with unaltered atherosclerotic burden (Fig. 1a–f). Reconstituting Apoe−/−Ccl17e/e mice with CCL17-sufficient bone marrow (BM) restored CCL3 plasma levels to those seen in Apoe−/− controls (Fig. 2d,e). Given that CCL3 was diminished in systemic circulation of CCL17-deficient mice, we assessed which cell types release CCL3 in response to CCL17 by sorting CD11c+MHCII+ cDCs, CD3+ T cells and CD19+B220+ B cells from LNs, monocytes and neutrophils from spleen and BM for in vitro treatment with CCL17. ELISA or multiplex bead array identified cDCs and T cells as the main source of CCL3 upon CCL17 stimulation, whereas CCL3 release from monocytes, B cells and neutrophils was less marked or negligible (Fig. 2f,g and Extended Data Fig. 3n). CCL3 release was induced by CCL17 in distinct T cell subsets, namely splenic CD4+ T helper cells, CD4+CD62L+ naive/memory T cells and CD4+CD25+ Treg cells (Fig. 2h). Comparing sorted CCL17+ (eGFP−) with CCL17-deficient CD11c+MHCII+ cDCs (eGFP+), we found that baseline CCL3 secretion was lower in CCL17-deficient cDCs, whereas CCL17-stimulated CCL3 release was similar (Fig. 2i), implying that CCL17+ cDCs can release CCL17 in an autocrine or paracrine fashion to induce CCL3 secretion. This effect was independent of CCR4 in cDCs and CD4+ T cells (Fig. 2j,k), indicating that CCL17 mediates CCL3 secretion through an alternative receptor.
a, Experimental scheme of Apoe−/−, Apoe−/−Ccl17e/e and Apoe−/−Ccr4−/− mice fed a WD. b,c, CCL3 plasma concentrations in Apoe−/− (n = 17) and Apoe−/−Ccl17e/e (n = 18) mice (b) or Apoe−/− (n = 18) and Apoe−/−Ccr4−/− (n = 17) mice (c), as measured by ELISA. d, Experimental scheme of reconstituting irradiated Apoe−/− or Apoe−/−Ccl17e/e mice with Apoe−/− BM before WD feeding. e, CCL3 concentrations in plasma of Apoe−/−►Apoe−/− (n = 5) or Apoe−/−►Apoe−/−Ccl17e/e (n = 7) mice were measured by ELISA. f,g, Sorted CD11c+MHCII+ cDCs (control, n = 17 replicates over 11 independent experiments; CCL17, n = 23 replicates over 12 independent experiments) (f) or CD3+ T cells (control and CCL17, n = 17 replicates over six independent experiments) were cultured for 4 h with or without recombinant mouse CCL17 (g). CCL3 concentrations in the supernatant were measured by multiplex bead array. h, Isolated T cell subsets from LN suspensions of Apoe−/− mice were stimulated for 4 h in the presence or absence of CCL17. CCL3 concentrations in supernatants were measured by ELISA (n = 6 for each bar). i, Sorted CD11c+MHCII+eGFP− cDCs (CCL17-competent, control and CCL17, n = 10) or CD11c+MHCII+ eGFP+ cDCs (CCL17-deficient, control n = 12; CCL17 n = 10) from Apoe−/−Ccl17e/WT mice were cultured for 4 h in the presence or absence of CCL17. CCL3 concentrations in supernatants were measured by ELISA. j, Sorted CD11c+MHCII+ cDCs from LNs of Apoe−/− mice were cultured for 4 h in the presence or absence of CCL17 with or without the CCR4 inhibitor C021, and CCL3 concentrations in supernatants of isolated cDCs were measured by multiplex bead array (n = 20 replicates across six independent experiments for all conditions). k, Isolated T cell subsets from LN suspensions of Apoe−/− mice were stimulated with or without CCL17 in the presence of absence of the CCR4 inhibitor C021 for 4 h and CCL3 concentrations in supernatants were measured by ELISA (control, n = 39 replicates; C021, n = 38 replicates; CCL17, 42 replicates; CCL17 + C021, n = 40 replicates; all distributed over nine independent experiments). Data represent mean ± s.e.m. (a–k). Two-sided P values as indicated versus control/Apoe−/− and as analyzed by unpaired Student’s t-test (b,c,e), nested t-test (f,g), two-way analysis of variance (ANOVA) (h) or nested ANOVA (j,k) with Holm–Šídák’s post hoc test and Kruskal–Wallis H with Dunn’s post hoc test (i).
CCL17 binds and activates CCR8 as a non-canonical receptor
Searching for alternative receptors, we revisited the notion that CCR8 could be a receptor for CCL17 (ref. 22) (findings subsequently contested23), as CCR8 has also been implicated in controlling the migration and function of Treg cells24,25. To probe for binding of CCL17 to CCR8, we used surface plasmon resonance (SPR) to record the concentration-dependent binding of CCR8-carrying liposomes to biotinylated CCL17 immobilized on a BIAcore C1 sensor chip, using CCR4-carrying, mock protein-carrying or pure liposomes as positive or negative controls (Fig. 3a–d). CCR8-bearing liposomes displayed saturable binding with a Kd (koff/kon) of 1.1 ± 0.4 nM, indicating a high-affinity interaction between CCL17 and CCR8 (Fig. 3b,c). While CCR4-bearing liposomes showed even stronger binding, irrelevant protein-bearing or empty liposomes did not support binding to CCL17 (Fig. 3a). A CCL5-chip used as a negative control did not support binding of CCR8-bearing liposomes (Fig. 3d). We next used a proximity-ligation assay in CCR4- or CCR8-transfected Jurkat cells or in adherent cDCs from LNs of Apoe−/− mice treated with CCL17 or the cognate CCR8 ligand CCL1 and subsequently with non-blocking antibodies to CCL17 or CCL1 and to CCR4, CCR8 or CCR5 to yield signals reporting receptor interactions of CCL17 or CCL1. Proximity-ligation signals and their quantification revealed an interaction of CCL17 with both CCR4 and CCR8, whereas CCL1 interacted with CCR8 but not with CCR4 (Fig. 3e,f and Extended Data Fig. 4a,b). We further performed receptor binding competition studies using fluorescently labeled CCL1 and CCL17 in CCR8-expressing HEK293 transfectants (Extended Data Fig. 4c). Inhibition of CCL1AF647 binding to CCR8-transfectants with increasing concentrations of unlabeled CCL17 yielded a half-maximum inhibitory concentration (IC50) of 9.4 ± 4.4 nM. Inhibition of CCL17AF647 binding to CCR8-transfectants with increasing concentrations of unlabeled CCL1 yielded an IC50 of 0.58 ± 0.24 nM, and with unlabeled CCL18 yielded an IC50 of 4.1 ± 1.7 nM, similar to unlabeled CCL17 (IC50 4.9 ± 2.4 nM) (Fig. 3g–j). Using primary CD4+ T cells from thymus or LNs of tamoxifen-inducible CCR8-competent UniCreErt2−Ccr8flox/flox (Ccr8WTApoe−/−) or CCR8-deficient UniCreErt2+Ccr8flox/flox (Ccr8KOApoe−/−) Apoe−/− mice, we found CCR8 internalization upon CCL17 treatment in CCR8-expressing but not CCR8-deficient T cells (Fig. 4a,b). The expression of CCR4 was unaltered in CCR8-deficient T cells and CCR4 internalization by CCL17 was preserved, while CCR8 internalization by CCL1 and CCL17 was dose-dependent in these cells (Extended Data Fig. 4d–f). In addition, human CD4+ T cells exhibited both CCR4 and CCR8 internalization upon CCL17 treatment, whereas CCL1 only internalized CCR8 and CCL22 only internalized CCR4 (Fig. 4c,d). These experiments indicate that CCL17 binds to CCR8.
a–d, SPR to detect CCL17–CCR8 interactions. RU, response unit. Human biotinylated CCL17 was immobilized on C1 sensor chips and sensorgrams of human CCR8-, CCR4-, mock protein-carrying or pure liposomes perfused at 0.5 µg ml−1 were recorded (a). Binding kinetics of CCR8-carrying liposomes perfused at indicated concentrations on immobilized CCL17 were determined after fitting (red) of curves (blue) with a 1:1 interaction model (b). Concentration-independent dissociation was low (koff 4.2 ± 0.8 × 10−4 s−1), indicating high stability. Calculating the koff/kon ratio using the theoretical molecular weight of CCR8 (40.2 kD) yielded an apparent Kd of 1.1 ± 0.4 nM. Shown is one representative of five experiments. Saturation binding of experiments from b was fitted with one-site-specific binding, yielding a Kd of 11.7 ± 2.1 nM (c). Data represent mean ± s.d. (n = 5 independent experiments). Sensorgrams of CCR8-carrying (red) or pure liposomes (black) perfused at 0.5 µg ml−1 on biotinylated human CCL5 immobilized as a negative control (d). e,f, Interactions of mouse CCL17 or CCL1 with CCR4 and CCR8 were analyzed in stably transfected Jurkat cells using Duolink proximity-ligation assays. Signals generated by close proximity of non-blocking antibodies bound to ligands or receptors on CCR8- (e, all n = 9 replicates over four independent experiments) or CCR4-transfectants (f, anti-CCL1, anti-CCL17 + anti-CCR4, n = 19 replicates each over six independent experiments; anti-CCL17, n = 22 replicates; anti-CCR4, n = 18 replicates; anti-CCL1 + anti-CCR4, n = 22 replicates over seven independent experiments) were quantified by flow cytometry. For anti-CCL17 and anti-CCL1 incubation, recombinant CCL17 (100 ng ml−1) or CCL1 (50 ng ml−1) was added, respectively. g–j, HEK293 cells stably transfected with human CCR8 were incubated with CCL1 or CCL17 labeled with Alexa-Fluor 647 (AF467) at the C terminus (10 nM). Background binding to mock-transfected HEK293 cells was subtracted; data were normalized to binding without unlabeled chemokine (control) and subjected to nonlinear fitting. Data represent mean ± s.d. (n = 3 independent experiments). Inhibition of CCL1AF647 binding to CCR8-transfectants with increasing concentrations of unlabeled CCL17; IC50 9.4 ± 4.4 nM (g). Inhibition of CCL17AF647 binding to CCR8-transfectants with increasing concentrations of unlabeled CCL1 (h, IC50 0.58 ± 0.24 nM), CCL17 (i, IC50 4.9 ± 2.4 nM) or CCL18 (j, IC50 4.1 ± 1.7 nM). Data represent mean ± s.e.m. (e,f). Two-sided P values versus respective controls were analyzed by nested ANOVA.
a,b, Representative histogram and quantification of CCR8 surface availability and internalization upon stimulation with recombinant mouse CCL17 (100 ng ml−1) on CD4+ T cells isolated from thymus (a) and LNs (b) of UniCreErt2−Ccr8flox/floxApoe−/− (Ccr8WTApoe−/−, a,b n = 6 for all conditions) or CCR8-deficient UniCreErt2+Ccr8flox/floxApoe−/− (Ccr8KOApoe−/− a, n = 5 for both, b, n = 6 for both) mice. FMO, fluorescence minus one. c,d, Representative histogram and quantification of CCR8 (c) and CCR4 (d) surface expression and internalization upon stimulation with recombinant human CCL17 (100 ng ml−1), CCL1 (50 ng ml−1) or CCL22 (50 ng ml−1) on CD4+ T cells isolated from human PBMCs (number of replicates in parentheses over number of independent experiments per bar from left to right for c, n = (59) 12, (28) 6, (29) 6 and (23) 6 and d, n = (56) 12, (27) 6, (23) 6 and (27) 6). e,f, Glosensor-HEK293 cells transfected with either CCR4 (e) or CCR8 (f). Cells were stimulated with recombinant human CCL17, CCL1 and CCL20 (all 100 ng ml−1) or PBS as vehicle control after 25 min of equilibration. Shown is one of four representative experiments. g, Transwell migration of CD4+ T cells isolated from Ccr8WTApoe−/− or Ccr8KOApoe−/− mice toward recombinant murine chemokines. Migrated cells were quantified by flow cytometry; chemotactic index induced by the chemokines CCL17 (100 ng ml−1), CCL1 (50 ng ml−1) and CCL22 (50 ng ml−1) was calculated as the ratio of chemokine-stimulated to unstimulated migration (number of independent experiments per bar from left to right, n = 10, 10, 6, 7, 5 and 5). h, Transwell migration of CD4+ T cells isolated from Ccr8WTApoe−/− or Ccr8KOApoe−/− mice toward recombinant murine CCL17 displayed in a checkerboard heat map format. Columns indicate CCL17 concentrations (ng ml−1) in the upper chamber; rows indicate CCL17 concentrations (ng ml−1) in the lower chamber. While blue represents enhanced, gray indicates reduced migration toward CCL17 in the bottom chamber. Each box represents a mean value of three independent experiments. Data represent mean ± s.e.m. (a–d,g). Two-sided P values as indicated versus Ccr8WTApoe−/− (a,b,g) or control (c,d) and as analyzed by two-way (a,b,g) or univariate (c,d) ANOVA with Holm–Šídák’s post hoc test.
To test whether CCL17 can elicit Gi-protein-coupled signaling via CCR8, we determined downstream cAMP levels in Glosensor-HEK293 cells transfected with CCR4 or CCR8 (Extended Data Fig. 4c,g) and stimulated with recombinant human CCL17, CCL1 or CCL20 (Fig. 4e,f). Dose–response curves revealed a near maximal CCR8 activation by CCL1 at a concentration of ≈1.2 nM, corresponding to the IC50 for receptor binding competition, whereas CCL17 and CCL18 were less effective (Extended Data Fig. 4h). Previous studies reported CCL17 binding to CCR8 but lack of subsequent calcium signaling22. In contrast, our results revealed that CCL17 induced Gi-mediated signaling both in CCR4- and CCR8-transfectants. CCL1 induced Gi-signaling in CCR8- but not CCR4-transfectants, whereas CCL20, as a negative control, had no effect (Fig. 4e,f). We next performed Transwell migration assays with CD4+ T cells from Ccr8WTApoe−/− or Ccr8KOApoe−/− mice. CCL17-induced migration of CD4+ T cells lacking CCR8 was lower than that of wild-type cells, and more markedly reduced than by a CCR8 antibody in human T cells, whereas migration toward CCL1 was abolished and that toward CCL22 was unaffected (Fig. 4g and Extended Data Fig. 4i). The migration of Apoe−/− CD4+ T cells induced by CCL1, CCL17 and CCL22 followed a bell-shaped dose–response curve mediated by CCR8 and CCR4, respectively, as shown by receptor blockade (Extended Data Fig. 4j–l) and was chemotactic, as evident by a checkerboard analysis applying CCR8-competent and -deficient CD4+ T cells with CCL17 in the upper chamber (Fig. 4h). Together, our data demonstrate that CCL17 activates CCR8 to induce Gi-signaling and functional responses.
CCL17–CCR8–CCL3 axis interferes with Treg differentiation
Having established the interaction of CCL17 with CCR8, we assessed which cell types express CCR8. Screening the Human Protein Atlas (https://www.proteinatlas.org; ref. 26) we found CCR8 mostly expressed on T cell subsets with an enrichment in Treg cells (Extended Data Fig. 5a,b). Accordingly, scRNA-seq of aortic LNs from CCL17-competent and CCL17-deficient mice revealed a prominent expression of CCR8 in CD4+ T cells, mainly in Treg cells and follicular T helper cells, and detectable CCR8 expression in a cDC subset (Extended Data Fig. 5c,d). We next assessed whether the CCL17–CCR8 pathway directly mediates CCL3 secretion by culturing CD4+ T cells from Apoe−/− mice with or without CCL17 and an antibody to CCR8 for 4 h. ELISA revealed an increase in CCL3 release induced by CCL17 alone but not when CCR8 was blocked (Extended Data Fig. 5e). Likewise, CD4+CD62L+ T cells sorted from Ccr8WTApoe−/− or Ccr8KOApoe−/− mice were co-cultured with CCR8-competent cDCs with or without CCL17 addition for 3 d to quantify CCL3 release and Treg differentiation (Fig. 5a). CCL3 secretion was induced by CCL17 in CCR8-competent but not CCR8-deficient CD4+ T cell/DC co-cultures (Fig. 5b). Correspondingly, the number of Treg cells in co-cultures of cDCs with CCR8-deficient CD4+ T cells was higher than in CCR8-bearing controls, even after addition of CCL17 (Fig. 5c). This indicates that CCL17-induced signaling via CCR8 on CD4+ T cells and a subsequent autocrine CCL3 release are important for restraining Treg differentiation, whereas cDC-derived paracrine production of CCL3 seems rather redundant.
a, Schematic of co-culture experiment, where isolated splenic CD4+CD62L+ T cells from Ccr8WTApoe−/− or Ccr8KOApoe−/− mice were combined with sorted CD11c+MHCII+ cDCs from LN of CCR8WTApoe−/− mice and cultured for 3 d in the absence or presence of recombinant murine CCL17 (100 ng ml−1). b, CCL3 concentrations were measured in cell supernatants by ELISA (number of independent experiments per bar from left to right, n = 7, 9, 6, 9). c, CD4+CD25+Foxp3+ Treg cells were quantified using flow cytometry analysis (number of independent experiments per bar from left to right: n = 10, 12, 10 and 13). d, Scheme of co-culture experiment, where isolated splenic CD4+CD62L+ T cells from Ccr8WTApoe−/− or Ccr8KOApoe−/−mice were combined with sorted CD11c+MHCII+ cDCs from LN of Apoe−/− or Apoe−/−Ccl17e/e mice and cultured for 3 d. e, CCL3 concentrations in the supernatant were determined by ELISA (number of independent experiments per bar from left to right, n = 6, 5, 4 and 6). f, CD4+CD25+Foxp3+ Treg cells were quantified by flow cytometry (number of independent experiments per bar from left to right, n = 8, 13, 4 and 6). g, Scheme of co-culture experiment, where splenic CD4+CD62L+ T cells isolated from Apoe−/− or Apoe−/−Ccl3−/− mice were combined with sorted CD11c+MHCII+ cDCs from LN of Apoe−/− mice and cultured for 3 d. h, CCL3 concentrations in the supernatant were determined by ELISA (number of independent experiments per bar from left to right, n = 5, 6, 5 and 6). i, CD4+CD25+Foxp3+ Treg cells were quantified by flow cytometry (number of independent experiments per bar from left to right, n = 6, 5, 6 and 5). Data represent mean ± s.e.m. (a–i). Two-sided P values as indicated versus respective controls and as analyzed by two-way ANOVA (b,c,e,f,h,i) with Holm–Šídák’s post hoc test.
Next, CD4+CD62L+ T cells sorted from Ccr8WTApoe−/− or Ccr8KOApoe−/− mice were co-cultured with CCL17-competent or CCL17-deficient cDCs for 3 d (Fig. 5d). Combining CCR8-competent naive T cells with CCL17-deficient compared to CCL17-competent cDCs reduced CCL3 levels, whereas co-cultures with CCR8-deficient naive T cells showed lower CCL3 levels with either CCL17-competent or CCL17-deficient cDCs (Fig. 5e). This was accompanied by inverse changes in Treg numbers, which were elevated in the presence of CCL17-deficient cDCs or using CCR8-deficient CD4+ T cells independent of CCL17 (Fig. 5f). To verify the requirement for T cell-derived CCL3, we co-cultured CD4+CD62L+ T cells from CCL3-competent or CCL3-deficient mice with CCR8-competent cDCs in the absence and presence of CCL17 for 3 d (Fig. 5g). CCL3 release was induced by CCL17 compared to baseline in CCL3-competent T cells and abolished in co-cultures with CCL3-deficient CD4+ T cells, where background CCL3 secretion from cDC was less sensitive to CCL17 (Fig. 5h). The number of Treg cells in co-cultures of cDCs with CCL3-deficient CD4+ T cells was increased, as compared to controls, and remained higher and not diminished upon addition of CCL17 (Fig. 5i). Together, our data demonstrate that CCL17 interaction with CCR8, particularly on CD4+ T cells, is critical in mediating CCL3 secretion and restraining Treg differentiation.
Atheroprotection by blockade or CD4-specific deletion of CCR8
To test whether CCR8 inhibition affects in vivo Treg numbers and atherosclerosis, we injected a blocking antibody to CCR8 or isotype control three-times weekly into Apoe−/− mice receiving WD for 4 weeks (Fig. 6a). The atherosclerotic lesion size in the aortic roots and arches was reduced (Fig. 6b,c), whereas lesional macrophage, SMC and collagen content was unaltered in anti-CCR8-treated mice (Extended Data Fig. 5f–i). Accordingly, CCL3 expression was reduced, whereas FoxP3+CD25+ Treg numbers were elevated in para-aortic LNs and spleens of anti-CCR8-treated mice (Fig. 6d–f) and lipid levels, circulating leukocyte and thrombocyte counts remained unaltered (Supplementary Table 4). Atherosclerotic lesion size in aortic root, thoracic-abdominal aorta and aortic arch was decreased in Ccr8KOApoe−/− versus Ccr8WTApoe−/− mice fed a WD for 12 weeks (Extended Data Fig. 6a–d). Because our in vitro data indicated a role of CCR8 on CD4+ T cells in controlling Treg differentiation, we generated CD4CreCcr8flox/floxApoe−/− mice (CD4Ccr8KOApoe−/−) lacking CCR8 in CD4+ T cells (Fig. 6g). Compared to CD4Cre−Ccr8flox/floxApoe−/− (CD4Ccr8WTApoe−/−) controls, we found a reduced atherosclerotic lesion burden in the aortic arch and thoraco-abdominal aorta after 12 weeks of WD (Fig. 6h,i). Lesional content of macrophages, SMCs and collagen was unaltered (Extended Data Fig. 6e–j), CCL3 expression in LNs was reduced, whereas Treg numbers were increased in para-aortic LNs and spleens but not in blood and thymus of CD4Ccr8KOApoe−/− mice (Fig. 6k,l and Supplementary Table 5). As Helios is a marker distinguishing thymic-derived from peripherally induced Foxp3+ Treg cells27, we quantified CD4+FoxP3+Helios+ Treg cells in aortic LNs, blood and thymus of CD4Ccr8KOApoe−/−and CD4Ccr8WTApoe−/− mice but did not observe any differences (Supplementary Table 5). Yet, flow cytometry analysis of aortic cell suspensions revealed increased CD4+FoxP3+ Treg numbers in the aorta of CD4Ccr8KOApoe−/− versus control mice after 12 weeks of WD (Fig. 6m), whereas the suppression capacity of CCR8-deficient Treg cells was reduced compared to CCR8-competent Treg cells (Fig. 6n,o). These data indicate that the CCL17–CCR8-CCL3–CCR1 axis affects Treg numbers directly at sites of inflammation (Fig. 6m) and in aortic LNs (Fig. 6k) rather than in blood and thymus. Lipid levels, circulating leukocyte and platelet counts, other T cell subsets in blood, spleen, aortic and axillary LNs did not differ in CD4Ccr8KOApoe−/− versus control mice (Supplementary Table 6), except a reduction of naive CD44−CD62L+ T cells in aortic LNs, reciprocating increased Treg frequencies (Supplementary Table 7 and Fig. 6k). This corroborates a crucial role for CCR8 on CD4+ T cells in conferring atherogenic effects of CCL17 via CCL3 release and Treg suppression.
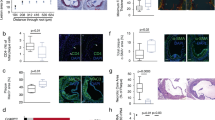
a, Experimental scheme of Apoe−/− mice fed a WD and injected three-times weekly with a blocking antibody to CCR8 or isotype control. b, Representative images and quantification of atherosclerotic lesion size after H&E staining in aortic arches of Apoe−/− mice (isotype, n = 9; anti-CCR8, n = 8). Scale bar, 200 µm. c, Lesion area after ORO in aortic roots of Apoe−/− mice (isotype, n = 10; anti-CCR8, n = 9). d, CCL3 mRNA expression levels in LNs of Apoe−/− mice. 18S rRNA was used as a housekeeping gene and changes in expression are given as fold change calculated with the 2−∆∆Ct method (isotype, n = 10; anti-CCR8, n = 8). e,f, Flow cytometry analysis of CD45+CD3+CD4+CD25+FoxP3+ Treg cells in para-aortic LNs (e) and spleens (f) of Apoe−/− mice (isotype, n = 10; anti-CCR8, n = 9). g, Experimental scheme of CD4Cre−Ccr8flox/floxApoe−/− (CD4Ccr8WTApoe−/−) and CD4Cre+Ccr8flox/floxApoe−/− (CD4Ccr8KOApoe−/−) mice fed a WD. h, Representative images and quantification of lesion area after H&E staining in the aortic arches of CD4Ccr8WTApoe−/− (n = 19) or CD4Ccr8KOApoe−/− (n = 22) mice. Scale bar, 200 µm. i, Atherosclerotic lesion size after ORO in aortas of CD4Ccr8WTApoe−/− (n = 20) or CD4Ccr8KOApoe−/− (n = 22) mice. j, CCL3 mRNA expression levels in LNs of CD4Ccr8WTApoe−/− (n = 19) or CD4Ccr8KOApoe−/− (n = 16) mice. 18S rRNA was used as a housekeeping gene and changes in expression are given as fold change calculated with the 2−∆∆Ct method. k,l, Flow cytometry analysis of CD45+CD3+CD4+CD25+FoxP3+ Treg cells in para-aortic LNs (n = 18 or 23) (k) and spleen (n = 20 or 23) (l) of CD4Ccr8WTApoe−/− (k, n = 18; l, n = 20) or CD4Ccr8KOApoe−/− (k,l, n = 23 each) mice. m, Flow cytometry analysis of CD45+CD4+FoxP3+ Treg cells in aortic cell suspensions from CD4Ccr8WTApoe−/− (n = 13) or CD4Ccr8KOApoe−/− (n = 7) mice fed a WD. n, Experimental scheme of a Treg suppression assay conducted with Apoe−/− CD4 effector T cells or Treg cells from CD4Ccr8WTApoe−/− or CD4Ccr8KOApoe−/− mice. o, Capacity of CD4Ccr8WTApoe−/− or CD4Ccr8KOApoe−/− Treg cells to suppress CD4+ effector T cell proliferation at varying Treg:Teff ratios (n = 2 replicates for each dilution over six independent experiments). Data represent mean ± s.e.m. (a–o). Two-sided P values as indicated and as analyzed by Mann–Whitney U-test (b,i,j,m), unpaired Student’s t-test (c,d–f,h,k,l) or a generalized linear model and Wald test for main model effects (o).
CCL17-driven CCL3 release limits Treg differentiation via CCR1
To explore which CCL3 receptor (CCR1 or CCR5) expressed on CD4+ T cells (Extended Data Fig. 6k,l) mediates the effects of CCL17-induced CCL3 release, we cultured CD4+CD62L+ T cells from spleens of Apoe−/−, Apoe−/−Ccr1−/− or Apoe−/−Ccr5−/− mice under Treg-polarizing conditions with or without CCL3 (Fig. 7a). Flow cytometry analysis revealed a decrease in CD4+CD25+Foxp3+ Treg frequencies among CD4+ T cells, when comparing CCL3-treated Apoe−/− or Apoe−/−Ccr5−/− T cell cultures to controls (TGFβ only), whereas this did not occur in Apoe−/−Ccr1−/− T cells (Fig. 7b), indicating that CCL3 restrains CD4+CD25+Foxp3+ Treg differentiation via CCR1. Evaluating the frequency of CD4+FoxP3+Tbet+ cells (as a subset with pro-atherogenic functions28) in LNs of CCL17- and CCL3-deficient mice, we observed a reduction of these cells in absence of CCL17 or CCL3 (Extended Data Fig. 6m). To confirm a role of the CCL3–CCR1 axis in mediating effects of CCL17, we sorted eGFP+ cDCs from Apoe−/−Ccl17wt/e (CCL17-competent) or Apoe−/−Ccl17e/e (CCL17-deficient) mice for co-culture with naive CD4+CD62L+ T cells from Apoe−/−, Apoe−/−Ccr1−/− or Apoe−/−Ccr5−/− mice (Fig. 7c). Apoe−/−Ccl17wt/e DCs reduce CD4+CD25+ Foxp3+ Treg frequencies in co-culture with Apoe−/− or Apoe−/−Ccr5−/− but not Apoe−/−Ccr1−/− T cells, establishing the importance of a CCL17-instructed CCL3–CCR1 axis in restraining Treg differentiation (Fig. 7d). As compared to controls, lesion size in the aortic root, arch and thoraco-abdominal aorta was reduced, lesional SMC content was increased, macrophage and collagen content were unaltered in Apoe−/−Ccr1−/− mice (Fig. 7e–h and Extended Data Fig. 7a–d). While CD3+CD4+CD25+Foxp3+ Treg cells were elevated in para-aortic, axillary and inguinal LNs and spleen of Apoe−/−Ccr1−/− mice, CCL3 plasma levels (Fig. 7i,j and Extended Data Fig. 7e–g), lipid levels and blood cell counts were unaltered (Supplementary Table 8). Our data are in line with reduced lesion size in Apoe−/−Ccr1−/− mice after 4 weeks of WD29.
a, Experimental scheme wherein CD4+CD62L+ T cells isolated from spleens of Apoe−/−, Apoe−/−Ccr1−/− or Apoe−/−Ccr5−/− mice were cultured for 3 d under Treg-polarizing conditions (TGFβ at 100 ng ml−1) in the presence or absence of recombinant mouse CCL3 (100 ng ml−1). b, Quantification of CD45+CD4+CD25+FoxP3+ Treg cells (number of independent experiments per bar from left to right, n = 7, 7, 7, 7, 7, 7, 6, 6 and 6) using flow cytometry. c, Scheme of co-culture experiment wherein CD4+CD62L+ T cells isolated from spleens of Apoe−/−, Apoe−/−Ccr1−/− or Apoe−/−Ccr5−/− mice were combined with sorted CD45+CD11c+MHCII+eGFP+ cDCs from LNs of Apoe−/−Ccl17wt/e or Apoe−/−Ccl17e/e mice and cultured for 3 d. d, Quantification of CD45+CD4+CD25+FoxP3+ Treg cells (number of independent experiments per bar from left to right, n = 5, 5, 5, 4, 5 and 3) using flow cytometry. e, Experimental scheme of Apoe−/− or Apoe−/−Ccr1−/− mice fed a WD for 12 weeks (e–j). f, Representative images and quantification of lesion area after ORO staining for lipid deposits in the aortic root of Apoe−/− (n = 10) or Apoe−/−Ccr1−/− (n = 9) mice. Scale bar, 500 µm. g, Quantification of lesion area after ORO staining for lipid deposits in the thoraco-abdominal aorta of Apoe−/− (n = 10) or Apoe−/−Ccr1−/− (n = 8) mice. h, Representative images and quantification of atherosclerotic lesion size in aortic arches of Apoe−/− (n = 11) or Apoe−/−Ccr1−/− (n = 8) mice after H&E staining. i,j, Flow cytometric quantification of CD45+CD3+CD4+CD25+FoxP3+ Treg cells in para-aortic LNs (i) and spleen (j) of Apoe−/− (i,j, n = 13 each) or Apoe−/−Ccr1−/− (i, n = 8; j, n = 9) mice. Data represent mean ± s.e.m. (a–j). Two-sided P values as indicated and as analyzed by a mixed-effects regression model followed by pairwise contrasts for fixed factors corrected by a Holm–Šídák’s procedure (b), unpaired Student’s t-test (d,f,i) or Mann–Whitney U-test (g,h,j).
CCL3 confers atherosclerosis and reduced Treg numbers in vivo
Under steady-state conditions, we found an increase in CD4+CD25+Foxp3+ Treg cells among CD4+ T cells in para-aortic, axillary and inguinal LNs and spleen of Ccl3−/− mice compared to wild-type controls (Extended Data Fig. 7h–k). A pro-atherogenic role of hematopoietic CCL3 was evidenced by protection in Ldlr−/− mice bearing CCL3-deficient BM cells30. Similar to CCL17-deficient mice, Apoe−/−Ccl3−/− mice displayed a marked reduction in lesion size in the aortic root, arch and thoraco-abdominal aorta (Fig. 8a–d), elevated FoxP3+ cells in aortic root lesions (Fig. 8e) and an increase in Treg numbers in aortic, axillary or inguinal LNs and spleen, when compared to controls (Fig. 8f,g and Extended Data Fig. 7l,m). The suppression capacity of Apoe−/−Ccl3−/− Treg cells in a 3-d assay with CD4+ effector T cells from Apoe−/− mice was enhanced (Fig. 8h,i). The lesional content of macrophages, SMCs and collagen (Extended Data Fig. 7n–p), body weight, lipid levels and blood cell counts (Supplementary Table 9) were unaltered in Apoe−/−Ccl3−/− versus control mice.
a, Experimental scheme. b, Representative images and quantification of lesion area after ORO staining in aortic roots of Apoe−/− (n = 9) or Apoe−/−Ccl3−/− (n = 12) mice. Scale bar, 500 µm. c, Quantification of lesion area after ORO in the thoraco-abdominal aorta of Apoe−/− (n = 10) or Apoe−/−Ccl3−/− (n = 12) mice. d, Representative images (scale bar, 200 µm) and quantification of lesion area after H&E staining of aortic arches from Apoe−/− or Apoe−/−Ccl3−/− mice (n = 8 each). e, Representative image and immunohistochemical quantification of lesional FoxP3+ cell numbers in aortic roots of Apoe−/− (n = 9) or Apoe−/−Ccl3−/− (n = 12) mice. f,g, Flow cytometry analysis of CD45+CD3+CD4+CD25+FoxP3+ Treg cells in para-aortic LNs (f) and spleen (g) of Apoe−/− (f, n = 8; g, n = 10) or Apoe−/−Ccl3−/− (f, n = 8; g, n = 12) mice. h, Experimental scheme of Treg suppression assays with Apoe−/− CD4+ effector T cells. i, Capacity of Apoe−/− or Apoe−/−Ccl3−/− Treg cells to suppress CD4+ T cell proliferation at varying Treg:Teff ratios (n = 6 each). j, Experimental scheme of Apoe−/− or Apoe−/−Ccl3−/− mice injected twice with isotype control or anti-CD25 antibody for Treg depletion. k, Lesion quantification in aortic roots of isotype control-treated Apoe−/− (n = 22), Apoe−/−Ccl3−/− (n = 7) or anti-CD25-treated Apoe−/−Ccl3−/− mice (n = 9). l, Flow cytometry analysis of CD45+CD3+CD4+CD25+FoxP3+ Treg cells in para-aortic LNs of isotype control-treated Apoe−/− (n = 21), Apoe−/−Ccl3−/− (n = 7) or anti-CD25-treated Apoe−/−Ccl3−/− mice (n = 9). m, Experimental scheme of Apoe−/− or Apoe−/−Ccl17e/e mice injected three-times weekly with PBS or recombinant mouse CCL3 (20 µg i.p. injection). i.p., intraperitoneal. n, Representative images and quantification of lesion area after ORO in aortic roots of PBS-treated Apoe−/− or Apoe−/−Ccl17e/e (n = 12 each) or CCL3-treated Apoe−/−Ccl17e/e (n = 8) mice. Scale bar, 500 µm. o,p, Flow cytometry analysis of CD45+CD3+CD4+CD25+FoxP3+ Treg cells in para-aortic LNs (o) and spleen (p) of PBS-treated Apoe−/− (o, n = 10; p, n = 13) or Apoe−/−Ccl17e/e (o, n = 15; p, n = 14) or CCL3-treated Apoe−/−Ccl17e/e mice (o, n = 9; p, n = 10). Data represent mean ± s.e.m. (a–p). Two-sided P values were analyzed by unpaired Student’s t-test or Mann–Whitney U-test (b–g), a generalized linear model and Wald test for main model effects (i) or a generalized linear model with mixed effects and post hoc pairwise contrasts within fixed factors corrected by a step-down Holm–Šídák’s procedure (k–p).
To establish that the atheroprotective effects of CCL3 deficiency were mediated by Treg cells, we depleted CD25+ cells in Apoe−/−Ccl3−/− mice using an antibody to CD25. As compared to isotype control, treatment of Apoe−/−Ccl3−/− mice with anti-CD25 antibody increased lesion size in the aortic root and decreased Treg frequencies in aortic LNs, restoring the levels to those observed in isotype control-treated Apoe−/− mice (Fig. 8j–l). Body weight, lipid levels and blood cell counts were unaltered in these mice (Supplementary Table 10). We next tested whether supplementing Apoe−/−Ccl17e/e mice with CCL3 would reinstate the phenotype of CCL17-competent mice (Fig. 8m). Injection of CCL3 (three-times weekly) into Apoe−/−Ccl17e/e mice during 4 weeks of WD increased aortic root lesion size and reduced axillary and splenic Treg numbers to levels seen in Apoe−/− controls (Fig. 8n–p). Body weight, lipid levels and blood cell counts were unaltered (Supplementary Table 11). Our data demonstrate the importance of CCL3 in restraining Treg cells and promoting atherosclerosis.
Proof-of-principle analysis in human gene expression datasets revealed higher CCL3 levels in atherosclerotic carotid artery segments with advanced (thick or thin fibrous cap atheroma) versus early lesions (intimal thickening or xanthoma) (Extended Data Fig. 8a; GSE28829) or in carotid atheroma specimens (stage IV) containing plaque core and shoulders versus remote, macroscopically intact tissue (stages I/II) (Extended Data Fig. 8b; GSE43292). We found increased CCL3 expression in human coronary arteries with atherosclerotic lesions from symptomatic compared to asymptomatic patients with coronary artery disease (Extended Data Fig. 8c; GSE11138). Likewise, CCL3 transcript levels were higher in carotid atherectomy specimens from symptomatic patients (characteristics in Extended Data Table 2) with ipsilateral neurological events, for example, transient ischemic attacks (n = 16), than in those from asymptomatic patients (n = 13) (Extended Data Fig. 8d). This was mirrored by lower FoxP3 expression indicative of reduced Treg abundance (Extended Data Fig. 8e). These data confirm increased CCL3 levels in progressing human lesions and imply a role in suppressing Treg differentiation in humans.
Discussion
Our quest to disambiguate the mechanisms underlying effects of CCL17 in an atherogenic context uncovered that aortic LNs of CCL17-deficient mice contain more tolerogenic cDCs, which license atheroprotective Treg maintenance. In turn, mice lacking the canonical CCL17 receptor CCR4 failed to phenocopy the effects of CCL17 deficiency. Instead, we could identify CCR8 as a functional high-affinity CCL17 receptor expressed by cDCs, CD4+ T cells and Treg cells. Further analysis established that CCL17–CCR8 interactions on CD4+ T cells facilitate CCL3 release, thereby suppressing Treg differentiation. Accordingly, interference with CCR8 by antibody blockade or CD4+ T cell-specific deletion blunted CCL3 levels and atherosclerotic lesion formation. Likewise, CCL3 deficiency attenuated lesion development and increased Treg numbers, whereas CCL3 applied in CCL17-deficient mice worsened atherosclerosis and hindered Treg differentiation, an effect that was dependent on CCR1. We found increased CCL3 expression and reduced FoxP3 levels in human plaques versus healthy arteries and in symptomatic versus asymptomatic plaques.
CCR7 is a key receptor guiding cDCs into T cell rich regions of lymphatic organs, enabling them to stimulate or suppress T cell immunity31. CCR7 has also been implicated in mediating egress of antigen-presenting cells from atherosclerotic lesions32. We provide evidence that the CCR7-expressing DCs cluster in aortic LNs harbors both CCL17+ and CCL17-deficient cDC populations. In LNs from CCL17-deficient mice, the number of CCR7-expressing DCs with a tolerogenic gene expression profile was twofold higher than in controls. Hence, an increased number of tolerogenic cDCs together with locally decreased CCL3 levels might explain higher Treg frequencies in lymphoid organs of CCL17-deficient mice. This is consistent with the hypothesis that CCL17+ DCs regulate the homeostatic mechanisms of T cells, including Treg differentiation in lymphoid tissues, and are thereby able to affect the development of atherosclerosis10. The involvement of Treg cells in limiting chronic inflammation and immune responses in mouse models of atherosclerosis17,33 and in alleviating atherosclerosis-related diseases in humans34,35,36 is well documented.
It is notable that a deficiency of CCR4, conventionally considered as the sole CCL17 receptor, failed to recapitulate any experimental features associated with CCL17 deficiency in our models. These findings mirrored related reports in experimental models of atopic dermatitis19 and colitis11, where reduced inflammation was observed in CCL17-deficient but not CCR4-deficient mice. A similar discrepancy was evident in models of allograft tolerance, where CCR4-deficient mice fail to develop tolerance due to diminished Treg recruitment, whereas CCL17-deficient mice show prolonged allograft survival13,37. Hence, we revisited the previously proposed but later contested concept22,25,38 that CCR8 may serve as an alternate CCL17 receptor and unequivocally establish that CCR8 indeed acts as a functional high-affinity receptor for CCL17. CCR8 is mainly expressed on CD4+ T cells and specifically on Treg cells39,40 but also present on monocytes, natural killer (NK) cells, group 2 innate lymphoid cells and DCs, depending on disease context and tissue location41,42,43. While the role of CCR8 in cancer has received great attention44,45, reports on its contributions to chronic inflammation remain scarce. CCR8 has been implicated in airway inflammation46 and in promoting pathogenic functions of interleukin (IL)-5+ TH2 cells during atopic dermatitis47. The role of CCR8 in atherosclerosis has been addressed in a study showing that genetic deletion of CCL1 in Apoe−/− mice reduced Treg recruitment to inflamed arteries and increased lesion formation48. In Ldlr−/− mice reconstituted with BM cells expressing FoxP3-driven red fluorescent protein, treatment with a CCR8-blocking antibody increased lesion size48, contrasting our findings likely due to different experimental setups. Whereas we used Apoe−/− mice for CCR8 blocking studies, the Ldlr−/− mice were subjected to BM transplantation48 and fed a cholesterol-rich diet for 1 week, an unusually short time span for evaluating the pathogenic role of adaptive immune cells in atherosclerosis. Yet, CCR8-expressing Treg cells interacting with CCL1 have been identified as key drivers of suppressive immunity in models of autoimmune encephalomyelitis24. We cannot exclude that CCL1–CCR8 interactions driving Treg recruitment contribute to atheroprotective effects in CCL17-deficient mice, nor that differences in receptor affinity or local availability of its ligands or biased signaling may shape anti- versus proinflammatory immune responses mediated by CCR8. In fact, both CCR8 ligands may be involved and differential levels in a given pathology may determine functional outcomes.
Expression of CCR8 was initially identified on human monocytes and lymphocytes49. Mouse pre-B cell transfectants (4DE4) expressing CCR8 dose-dependently migrated and exhibited specific calcium transients in response to CCL1 but not other chemokines tested (albeit not including CCL17)49. Subsequently, CCL17 was suggested to act as a functional CCR8 ligand, evidenced by a dose-dependent migration of Jurkat CCR8-transfectants toward CCL17 (ref. 22). This was supported by a study revealing CCR8 expression and dose-dependent migration of human IL-2-activated NK cells (IANK) in response to CCL17 (ref. 36). While CCL1 induced a CCR8-dependent calcium flux in IANK cells, partially inhibiting CCL17-induced calcium flux, CCL17 fully desensitized the calcium response to CCL1. This discrepancy was explained by the expression of CCR4 on IANK cells, which cannot be desensitized by CCL1 (ref. 41). Accordingly, other groups were unable to show migration, calcium flux or receptor internalization in CCR8-transfected 4DE4 cells in response to CCL17 (refs. 38,50). This may be related to the fact that 4DE4 transfectants are a suboptimal model for signaling studies, whereas primary human IANK41 cells such as CD4+ T cells used herein represent more physiological CCR8-bearing cell types. Still, calcium flux induced by CCL17 in IANK cells was predominantly mediated by CCR4 (ref. 41). Given these inconsistencies, we applied assays beyond migration and receptor internalization, both of which documented CCL17 activity for CCR8, to confirm a functional high-affinity CCL17 interaction with CCR8. Proximity ligation in DCs or Jurkat CCR8-transfectants, SPR and binding competition revealed binding of CCL17 to CCR8 with apparent affinities ranging from 1.1 nM (Kd SPR) to 9.4 nM (IC50 CCL1 competition), thus equivalent to that for CCL18 (Kd 1.9 nM) but lower than that found for CCL1 by us (IC50 0.58 nM) and others (Ki/IC50 0.11–0.22 nM)50,51. Determining cAMP levels in CCR4- or CCR8-transfected HEK293 cells confirmed that CCL17 induced Gi-signaling via both receptors. This extends findings that CCR8 mediates chemotactic migration toward CCL17, unequivocally establishing that CCR8 as a signaling high-affinity receptor for CCL17. Our data can be reconciled with a report that CCL17 induced chemotaxis of Jurkat CCR8-transfectants, albeit without eliciting calcium mobilization22. Findings disputing the assignment of CCL17 as a CCR8 ligand may have been due to insufficient bioactivity, as no positive controls were provided22. A role of CCR8 in mediating the restraint of Treg homeostasis may thus serve to complement or counter-balance functions of CCR4 in Treg recruitment during inflammation and cancer52,53. Preliminary evidence that CCR4 and CCR8 can engage in a heterodimeric interaction may further imply alternative mechanisms of modulation that will be subject of future studies. For instance, CCL17 inhibited Treg recruitment through biased activation of CCR4, activating Gq-signaling but inhibiting CCL22-stimulated β-arrestin signaling to explain the abundance of Treg cells in injured myocardium of CCL17-deficient mice54, an effect possibly attributable to CCL17 activity mediated by CCR8.
It is tempting to speculate that only chronic inflammatory conditions, as present in atherosclerosis, atopic dermatitis19 or colitis11, foster the development of CCL17-expressing cDCs, which then trigger Treg restraint by inducing CCL3 release through CCR8 in lymphoid organs. Notably, our data show that it is primarily CCR8 on CD4+ T cells, which orchestrates the restraint of Treg cells by upregulating CCL3 in response to CCL17, as evident by decreased lesion size and increased Treg numbers in Apoe−/− mice lacking CCR8 on CD4+ T cells. Because CCR8 is prominently expressed in Treg cells, it is conceivable that at sites of inflammation or in T cell rich areas of LNs CCL17 directs Treg trafficking but also prevents Treg differentiation through induction of CCL3. This mechanism would explain why isolated CD25+CD4+ T cells secreted CCL3 in response to CCL17. Under chronic inflammatory conditions like atherosclerosis, however, CCL17+ cDCs are continuously present and skewing CD4+ T cell responses toward a proinflammatory type. This concept is corroborated by mechanistic studies in psoriasis as a chronic inflammatory autoimmune disease. The transcription factor Grainyhead-like 3 is crucial for maintaining barrier integrity of the skin, whereas its knockdown upregulates CCL17 in keratinocytes, driving their proliferation and an inflammatory T cell infiltration pattern resembling psoriasis55. Moreover, elevated CCL3 inversely correlates with FoxP3 levels in Treg cells of psoriatic patients and CCL3 interferes with FoxP3 stability by promoting ubiquitination-dependent degradation56. Psoriatic disease may thus be prompted by CCL17-induced CCL3 expression to impair FoxP3 stability and reduce Treg numbers. It will be intriguing to dissect whether CCL3 induction by CCL17 is restricted to cell types expressing CCR8, whether additional cell types are licensed by CCR8 expression to enact this mechanism of Treg control and which specific signaling pathways couple CCR8 to CCL3 release.
Previous evidence on the role of CCL3 in atherosclerosis, despite not pinpointing the cellular sources of CCL3, lends support to our findings. In Ldlr−/− mice reconstituted with Ccl3−/− BM, aortic lesion formation and inflammatory neutrophil adhesion was reduced; however, an involvement of T cell subsets was not examined30. Atorvastatin inhibits the 5-lipoxgenase pathway in Apoe−/− mice, thereby downregulating CCL3 expression and attenuating lesion development, to implicate CCL3 as a therapeutic target in atherosclerosis57. Mice lacking CCL3 are also protected from aortic inflammation and aneurysm formation58. Here we show that genetic deletion of CCL3 in Apoe−/− mice reduced lesion size and increased Treg numbers and depletion studies indicated that atheroprotection was mediated by Treg cells. Probing CCL3 receptors, CCR5 deficiency conferred a protective phenotype in different mouse models of atherosclerosis59, whereas results on CCR1 deficiency were more ambiguous29,59. Our data demonstrate that Treg restraint by CCL3 is afforded by CCR1 and that CCR1 deficiency in Apoe−/− mice decreased lesion development and enhanced Treg numbers. Nevertheless, findings may be reconciled depending on the mouse model and disease phenotype, as CCR1 engages multiple other chemokine ligands. Thus, cell subsets interacting in the vicinity and the local tissue environment may determine the availability of CCR1 ligands and how CCL3 shapes the immune response at relevant interfaces.
In synopsis, our data establish that CCL17 binds to CCR8 as its second functional high-affinity receptor besides CCR4, and introduce CCL17 to the unique ligand spectrum of CCR8, including its major ligand CCL1, CCL8 (ref. 47), a chemokine responsible for pathogenic circuits in atopic dermatitis and the widely expressed inflammatory chemokine CCL18 (ref. 50). The functional relevance in primary cells (Treg cells), unfolds another facet within the remarkable versatility of the chemokine-receptor family60. Our data show that CCL17 signaling via CCR8 on CD4+ T cells triggers CCL3 secretion, which engages CCR1 and suppresses Treg differentiation to drive atherogenic effects of CCL17 (Extended Data Fig. 8f). We propose that the specific instruction of CD4+ T cells by CCL17+ cDCs dictating a CCL3-dependent restraint of Treg cells may constitute a broadly relevant mechanism in chronic inflammatory disease and identify the sequential CCL17–CCR8–CCL3–CCR1 pathway as a target for multilayered therapeutic intervention.
Methods
Mice
All experiments were approved by local authorities and complied with German animal protection law (Regierung von Oberbayern; ROB-55.2-2532.Vet_02-14-189, ROB-55.2-2532.Vet_02-18-96 and ROB-55.2-2532.Vet_02-20-26). Every effort was made to minimize suffering. Ccr4−/− mice61 were kindly provided by K. Pfeffer (Heinrich-Heine-Universität) and Ccl17e/e (GFP reporter knock-in) mice13 were kindly provided by I. Förster (Universität Bonn). Ccl3−/− mice were purchased from the Jackson Laboratories. Ccr1−/− mice and Ccr5−/− mice were kindly provided by P.M. Murphy and W.A. Kuziel, respectively, and have been previously characterized59,62. Ccr4−/−, Ccr1−/−, Ccr5−/−, Ccl17e/e and Ccl3−/− mice were crossed with Apoe−/− mice purchased from the Jackson Laboratories. Ccr8flox/flox mice were generated at Ozgene, backcrossed into a C57BL/6 background and crossed with C57BL/6 Apoe−/− mice in house. Apoe−/−UniCreErt2 (ubiquitous inducible Cre expression), described previously63, and CD4Cre (purchased from Jackson laboratory) bred to Apoe−/− were crossed in house with Apoe−/−Ccr8flox/flox mice to generate whole body or T cell-specific Ccr8 knockout mice, respectively. To induce Ccr8 deletion in UniCreErt2 mice the mice received an i.p. injection with tamoxifen (50 mg kg−1 body weight, from Sigma-Aldrich and dissolved in Miglyol, Caelo) for five consecutive days. All strains were backcrossed for at least ten generations to the C57BL/6 background. All mice were housed under specific-pathogen-free conditions in 12-h light–dark cycles at 21 °C and 50% humidity with ad libitum food and water. Depending on the type of study, mice were either fed a normal chow diet (steady state) or WD (for atherosclerosis studies) containing 21% fat and 0.15–0.2% cholesterol (Altromin 132010, Sniff TD88137) starting at 8–10 weeks of age for 4 or 12 weeks before being killed. For the rescue experiment using CCL3 injections, mice were injected 3× weekly with 20 µg recombinant mouse CCL3 or PBS control by i.p. injection. For depletion of CD25+ cells, including Treg cells, Apoe−/− or Apoe−/−Ccl3−/− mice were fed a WD for 4 weeks and injected twice (every second week) with isotype control or anti-CD25 antibody (each 250 µg antibody per i.p. injection). For the experiment using anti-CCR8 blocking antibody, mice were injected 3× weekly with 5 µg anti-CCR8 antibody or isotype control by i.p. injection. For scRNA-seq, male Ccl17wt/eApoe−/− and Ccl17e/eApoe−/− mice were fed a normal chow diet or 6 weeks of WD. All experimental mice were sex- and age-matched.
Histology and immunofluorescence
Atherosclerotic lesion size was assessed by analyzing cryosections of the aortic root by staining for lipid depositions with ORO. In brief, hearts with the aortic root were embedded in Tissue-Tek O.C.T. compound (Sakura) for cryosectioning. ORO+ atherosclerotic lesions were quantified in 4-µm transverse sections and averages were calculated from three sections. The thoraco-abdominal aorta was fixed with 4% paraformaldehyde and opened longitudinally, mounted on glass slides and stained enface with ORO. Aortic arches with the main branch points (brachiocephalic artery, left subclavian artery and left common carotid artery) were fixed with 4% paraformaldehyde and embedded in paraffin. Lesion size was quantified after H&E staining of 4-µm transverse sections and averages were calculated from 3–4 sections. For analysis of the cellular composition or inflammation of atherosclerotic lesions, aortic root sections were stained with antibodies to Mac2 (Cedarline), smooth muscle actin (Dako) or FoxP3 (Abcam). Nuclei were counter-stained by 4′,6-diamidino-2-phenylindol (DAPI). After incubation with a secondary FITC- or Cy3-conjugated antibody (Life Technologies) for 30 min at room temperature, sections were embedded with VectaShield Hard Set Mounting Medium (Vector Laboratories) and analyzed using a Leica DM4000B LED fluorescence microscope and charge-coupled device camera. For FoxP3 staining, an Avidin/Biotin Blocking kit, VECTASTAIN ABC-AP and Vector Red Substrate kit were applied (all from Vector Laboratories). Blinded image analysis was performed using Diskus, Leica Qwin Imaging (Leica) or ImageJ software. For each mouse and staining, two to three root sections were analyzed and the average was taken.
Laboratory blood parameters and flow cytometry
Whole blood from the mice was collected in EDTA-buffered tubes. Thrombocyte counts were determined using a Celltac Automated Hematology Analyzer (Nihon Kohden). Afterwards, samples were subjected to red-blood-cell lysis for further analysis using flow cytometry. Spleen and LNs were mechanically crushed and passed through a 30-μm cell strainer (Cell-Trics, Partec) using Hank’s medium (HBSS + 0.3 mmol l−1 EDTA + 0.1% BSA; Gibco by Life Technologies) to obtain single-cell suspensions. Leukocyte subsets were analyzed using the following surface marker combinations: neutrophils (CD45+CD11b+CD115-Gr1high), classical (CD45+CD11b+CD115+GR1high) and non-classical (CD45+CD11b+CD115+GR1low) monocytes, B cells (CD45+B220+), T cells (CD45+CD3+). Treg cells were classified as CD45+CD3+CD4+CD25+FoxP3+ (the gating strategy used to identify Treg cells throughout the manuscript is depicted in Extended Data Fig. 1j) and its subpopulation as CD45+CD3+CD4+FoxP3+Tbet+. Foxp3 transcription factor was stained using the Foxp3/Transcription Factor Staining Buffer Set (eBioscience). Annexin-V+ cells were analyzed using the Dead Cell Apoptosis kit (Thermo Fisher Scientific). Cell populations and marker expression were analyzed using a FACSCanto II, FACSDiva software v.8.0 (BD Biosciences) and the FlowJo analysis program v.10 (Tree Star).
All aortas, including the aortic root, aortic arch and thoracic portions were subjected to a house-made enzymatic digestion and post-digestion protocol64. Single-cell suspensions were obtained by mashing aortas through a 70-μm cell strainer. Live/dead staining was performed with Zombie Violet Fixable Viability kit (BioLegend, 423113) followed by surface staining with antibodies: anti-CD45-APC-Cy7 (BioLegend, clone 30-F11, 1:300 dilution), anti-CD11b-PerCP-Cy5.5 (BioLegend, clone M1/70, 1:300 dilution), anti-CD3e-FITC (BioLegend, clone 145-2C11, 1:300 dilution) and anti-CD4-PE-Cy7 (BioLegend, clone RM4-5; 1:500 dilution) including unconjugated anti-CD16/32 (BioLegend, clone 93, 1:500 dilution). Intracellular staining for FoxP3 was performed using anti-FoxP3-PE antibody (eBiosciences, clone FJK-16s, 1:50 dilution) and FoxP3/Transcription Factor Staining Buffer Set (eBiosciences, 00-5523-00) according to the manufacturer’s protocol. Data were acquired using flow cytometry (BD FACSCanto II, BD Biosciences) and analyzed using FlowJo v.10 (FlowJo).
Plasma lipid levels
Cholesterol and triglyceride levels were analyzed using mouse EDTA-buffered plasma and quantified using enzymatic assays (c.f.a.s. Cobas, Roche Diagnostics) according to the manufacturer’s protocol.
Fluorescence-activated cell sorting and tolerogenic DC analysis
For the isolation of DCs, LNs were mechanically crushed and passed through a 30-μm cell strainer (Cell-Trics, Partec) using Hank’s medium (HBSS + 0.3 mmol l−1 EDTA + 0.1% BSA; Gibco by Life Technologies) to obtain single-cell suspensions. cDCs were isolated from this suspension by fluorescence-activated cell sorting (BD FACSAria), by gating for CD45+CD11c+MHCII+ cells. Furthermore, eGFP+Ccl17wt/e and eGFP+Ccl17e/e DCs were isolated by gating for the endogenous eGFP signal in the FITC channel (pre-gating, CD45+CD11c+MHCII+). Flow cytometric analysis of tolerogenic DCs in aortic LNs was performed by pre-gating for CD45+CD11c+ MHCII+ followed by evaluation of CD83, CCR7, IDO and CD274 on pre-gated cDCs20,65,66,67.
For the isolation of T and B cells, spleens were mechanically crushed and passed through a 30-μm cell strainer (Cell-Trics, Partec) using Hank’s medium (HBSS + 0.3 mmol l−1 EDTA + 0.1% BSA; Gibco by Life Technologies) to obtain single-cell suspensions. Cell subsets were isolated by fluorescence-activated cell sorting (BD FACSAria), by gating for CD45+CD3+ cells (T cells) or CD45+CD19+ cells (B cells). After sorting, all cells were cultured in 96-well round-bottom plates (1 × 105 cells per well) (Corning Costar by Sigma-Aldrich/Merck) in RPMI-1640 medium supplemented with 10% (v/v) fetal calf serum, 2 mM l-glutamine and 1% penicillin/streptomycin (all Gibco by Life Technologies), unless stated otherwise and with/without specific stimuli as indicated for the individual experiments.
Immunomagnetic cell isolation
For the isolation of monocytes and neutrophils, BM cells were collected by flushing femurs with Hank’s medium (HBSS + 0.3 mmol l−1 EDTA + 0.1% BSA) (Gibco by Life Technologies). Monocytes are isolated using the mouse Monocyte Isolation kit and an LS separation column (all Miltenyi Biotec), according to the manufacturer’s protocol. Neutrophils were isolated using the mouse Neutrophil Isolation kit and an LS separation column (all Miltenyi Biotec), according to the manufacturer’s protocol. After isolation, all cells were cultured in 96-well round-bottom plates (1 × 105 cells per well) (Corning Costar by Sigma-Aldrich/Merck) in RPMI-1640 medium supplemented with 10% (v/v) fetal calf serum, 2 mM l-glutamine and 1% penicillin/streptomycin (All Gibco by Life Technologies) unless stated otherwise, with/without specific stimuli as indicated for the individual experiments.
For the isolation of CD4+, CD4+CD62L+ and CD4+CD25+ T cells, spleens were mechanically crushed and passed through a 30-μm cell strainer (Cell-Trics, Partec) using Hank’s medium (HBSS + 0.3 mmol l−1 EDTA + 0.1% BSA; Gibco by Life Technologies) to obtain single-cell suspensions. CD4+ T cells were subsequently isolated using the mouse CD4+ T cell Isolation kit and an LS separation column (all Miltenyi Biotec), according to the manufacturer’s protocol. CD4+CD62L+ T cells were subsequently isolated using the mouse CD4+CD62L+ T cell Isolation kit II and an LS separation column (all Miltenyi Biotec), according to the manufacturer’s protocol. CD4+CD25+ T cells were subsequently isolated using the mouse CD4+CD25+ T cell Isolation kit and an LS separation column (all Miltenyi Biotec), according to the manufacturer’s protocol. After isolation, cells were cultured in 96-well round-bottom plates (1 × 105 cells per well) (Corning Costar by Sigma-Aldrich/Merck) in RPMI-1640 medium supplemented with 10% (v/v) fetal calf serum, 2 mM l-glutamine and 1% penicillin/streptomycin (all Gibco by Life Technologies), unless stated otherwise and with/without specific stimuli as indicated for the individual experiments.
For the isolation of human CD4+ T cells, 18 ml whole blood was collected from healthy volunteers and mixed with 2 ml citrate to avoid blood coagulation (approved by the local ethics committee, LMU Munich, no. 18–283). Whole blood was then diluted with same volume of T cell isolation buffer (phosphate-buffered saline + 2 mmol l−1 EDTA + 0.1% BSA; Gibco by Life Technologies) and gently layered over twofold volume of Biocoll solution (1.077 g ml−1; Bio&SELL), followed by centrifugation for 25 min at 600g without brake. The top layer of plasma was removed and mononuclear cells in the middle layer were carefully collected and transferred to a new tube. The mononuclear cells were washed with T cell isolation buffer twice and centrifuged for 10 min at 300g. The supernatants were discarded and cell pellets were resuspended with T cell isolation buffer to reach a final density of 1 × 108 cells per ml. Human CD4+ T cells were isolated from this cell suspension with Dynabeads Untouched Human CD4 T Cells kit (Invitrogen) according to the manufacturer’s protocol.
Transmigration assay
Mouse and human CD4+ T cells were isolated according to the manufacturer’s protocols as detailed above. Transmigration assays were performed using HTS Transwell 96-well plates (3.0-μm pore size with polycarbonate membrane; Corning). Murine or human recombinant CCL17 (BioLegend) was added to the bottom chambers at a concentration of 100 ng ml−1 or as indicated in RPMI-1640 medium containing 0.5% BSA. Murine or human recombinant CCL1 or CCL22 (Peprotech) was added to the bottom chambers at a concentration of 50 ng ml−1 or as indicated in RPMI-1640 medium containing 0.5% BSA. Mouse CD4+ T cells from Apoe−/−, Ccr8WTApoe−/− or Ccr8KOApoe−/− mice or human CD4+ T cells (1 × 105) were added to the top chamber in the presence or absence of CCR4 receptor antagonist C 021 dihydrochloride (Tocris) at a concentration of 0.5 µM; or human CD4+ T cells (1 × 105) were pretreated with or without anti-CCR8 antibody (R&D Systems) and added to the top chamber and allowed to migrate for 3 h. The number of cells migrated was analyzed by flow cytometry (FACSCanto II, BD Biosciences) and FlowJo v.10 software (Tree Star). The chemotactic index was calculated as the ratio of chemokine-stimulated to unstimulated migration.
In another quantitative transmigration assay (checkerboard), HTS Transwell 96-well plates (3.0-μm pore size with polycarbonate membrane; Corning) were also used. Isolated CD4+ T cells (1 × 105) were added to the upper chamber of each well in a total volume of 80 μl of RPMI-1640 medium containing 0.5% BSA. Murine recombinant CCL17 (BioLegend) was used at concentrations of 1 µg ml−1, 100 ng ml−1, 10 ng ml, 1 ng ml−1 or 0 ng ml−1 in RPMI-1640 medium containing 0.5% BSA in the lower, upper or both lower and upper chambers of the Transwell to generate ‘checkerboard’ analysis matrix of positive, negative and the absent gradients of CCL17, respectively. Cells were collected from the lower chamber 3 h later and counted. The number of cells migrated was analyzed by flow cytometry (FACSCanto II, BD Biosciences) and FlowJo v.10 software (Tree Star). The chemotactic index was calculated as the ratio of migrated cell counts of each well to unstimulated migration without murine recombinant CCL17 in both lower and upper chamber.
T effector polarization assay
Splenic CD4+CD62L+ T cells were obtained by immunomagnetic cell isolation as described previously. CD4+CD62L+ T cells (1 × 105) were cultured in 96-well tissue round-bottom culture plates in the presence of anti-CD3e (pre-coated overnight, 5 µg ml−1), anti-CD28 (1 µg ml−1) and supplemented with TGFβ (5 ng ml−1) in the presence or absence of CCL17 (100 ng ml−1) for 3 d. Treg cells were classified as CD45+CD3+CD4+CD25+Foxp3+. The number of Treg cells was analyzed by flow cytometry (FACSCanto II, BD Biosciences) and FlowJo v.10 software (Tree Star).
Treg suppression assay
Splenic CD4+CD25− T cells and CD4+CD25+ Treg cells from Apoe−/−, Apoe−/−Ccl3−/− or Ccr8KOApoe−/− were isolated using CD4+CD25+ Regulatory T Cell Isolation kit (Miltenyi Biotec, 130-091-041) according to manufacturer’s instructions. Isolated CD4+CD25− T cells were labeled with 5 µM Cell Proliferation Dye eFluor 670 dye (eBioscience, 65-0840-90) according to the manufacturer´s instructions and co-cultured with different concentrations of CD4+CD25+ Treg cells and with Dynabeads Mouse T-Activator CD3/CD28 (Thermo Fisher Scientific, 11456D) for 72 h at 37 °C with 5% CO2. Proliferation of CD4+CD25− T cells was assessed by flow cytometry (BD FACSCanto II, BD Biosciences). Data were analyzed using FlowJo v.10 (Tree Star).
Co-culture experiments
DCs were isolated from cell suspension from LNs by fluorescence-activated cell sorting (BD FACSAria), by gating for CD45+CD11c+MHCII+ cells as previously mentioned. Sorted DCs were subsequently co-cultured for 3 d in 96-well tissue flat-bottom culture plates with splenic naive CD4+CD62L+ T cells obtained by immunomagnetic cell isolation as described previously in a DC:T cell ratio of 1:2 (in general 2.5 × 104 to 5 × 104 cells), with/without specific stimuli as indicated for the individual experiments. The percentage of Treg cells (CD45+CD3+CD4+CD25+FoxP3+ cells) relative to CD4+ T cells was analyzed by flow cytometry (FACSCanto II, BD Biosciences) and FlowJo v.10 software (Tree Star). Supernatants were collected for further ELISA or multiplex bead array analysis.
Cell culture experiments for CCL3 release
To measure CCL3 release of CD4+, CD4+CD62L+, CD4+CD25+, CD45+CD11c+MHCII+, eGFP+Ccl17wt/e DCs, eGFP+Ccl17e/e DCs CD45+CD3+ cells (T cells) or CD45+CD19+ B cells and monocytes as well as neutrophils cells were seeded at 1 × 105 into a 96-well round-bottom plate and incubated in the absence or presence of CCL17 (100 ng ml−1) in combination with C021 (5 µM) or anti-CCR8 antibody (2 µg ml−1) for 4 h. Thereafter, supernatants were collected and measured by CCL3 ELISA as detailed below.
Multiplex bead array
Cell culture supernatants and mouse plasma were analyzed for various cytokines using the ‘Cytokine & Chemokine 26-Plex Mouse ProcartaPlex Panel 1’ (Thermo Fisher Scientific, eBioscience), sample preparation and analysis were performed according to the manufacturer’s protocol. The kit allows the simultaneous detection and quantification of soluble murine IFNγ; IL-12p70; IL-13; IL-1β; IL-2; IL-4; IL-5; IL-6; TNFα; granulocyte–macrophage colony-stimulating factor; IL-10; IL-17A; IL-18; IL-22; IL-23; IL-27; IL-9; GROα (CXCL1); IP-10 (CXCL10); MCP-1 (CCL2); MCP-3 (CCL7); MIP-1α (CCL3); MIP-1β (CCL4); MIP-2 (CXCL2); RANTES (CCL5); eotaxin (CCL11). The bead-based assay followed the principles of a sandwich immunoassay. Fluorescent magnetic beads were coupled with antibodies specific to the analytes to be detected. Beads were differentiated by their sizes and distinct spectral signature (color-coded) by flow cytometry using Luminex xMAP, data were collected with xPONENT software (Thermo Fisher Scientific, v.4.2) and analyzed with ProcartaPlex Analyst software (Thermo Fisher Scientific, v.1.0).
ELISA
CCL3 plasma (EDTA-plasma of full blood) levels or CCL3 levels in cell culture supernatants were quantified by ELISA using a commercially available kit (CCL3 Mouse Uncoated ELISA kit with plates from Thermo Fisher Scientific or Mouse CCL3/MIP-1 alpha Quantikine ELISA kit by R&D System) following the manufacturer’s protocol. The final measurement of absorbance was carried out using a plate reader (Tecan) set to 450 nm with a correction factor of 550 nm.
Cyclic AMP signaling
Levels of cyclic adenosine monophosphate (cAMP) were measured in confluent Flp-In system and Flp-In TREx-293 (HEK293) cells (Invitrogen). HEK293 cells were transfected using plasmids harboring sequences of CCR4 and CCR8 (Missouri S&T cDNA Resource Center; www.cdna.org). The sequence of the luciferase-cAMP binding site fusion protein from the pGloSensor-20F-vector (Promega) was amplified and ligated into a bicistronic pIRESneo vector (Clontech) to obtain the reporter gene plasmid. HEK293 cells were transfected with the reporter gene vector using Eco-Transfect (OZBioscience), stable clones were selected as host cell lines for expressing receptor constructs using the Flp-In system68 and reselected with G418 and hygromycin B. After incubation with luciferin-EF (2.5 mM, Promega) at room temperature for 2 h, cells were stimulated with CCL17, CCL1 or CCL20 (100 ng ml−1 each) or left unstimulated (PBS control) and luminescence indicating the reduction of cAMP was recorded over time.
Proximity-ligation assay
Proximity ligation was carried out using the Duolink In Situ Red kit goat/rabbit (Sigma-Aldrich) on PFA-fixated mouse DCs cultured on collagen-coated coverslips that were pre-incubated with recombinant mouse CCL1 (Peprotech) and CCL17 (BioLegend) using primary polyclonal antibodies to mouse CCL17 (R&D systems), mouse CCL1 (Acris), mouse CCR4 (Thermo Scientific), mouse CCR5 (Santa Cruz Biotechnology) and mouse CCR8 (Abcam) according to the manufacturer’s instructions. Imaging was performed using fluorescence microscopy (Leica DM4000) after which deconvolution algorithms for wide-field microscopy were applied to improve overall image quality (Huygens Professional 16.10; SVI). The number of Duolink-detected interactions was determined in the processed images using the Leica LAS 4.2 analyses software. To more accurately resolve the interactions detected with Duolink, representative DC samples of each condition were also visualized with a Leica SP8 3X microscope using a combination of 3D confocal microscopy (DAPI) and 3D STED nanoscopy (Duolink Red). Image processing and deconvolution of the resultant 3D datasets was performed using the Leica LAS X and Huygens professional software packages.
Proximity ligation was also carried out using the Duolink In Situ Probe anti-goat or anti-rabbit or anti-mouse kit (Sigma-Aldrich) on PFA-fixated CCR4-transfected or CCR8-transfected Jurkat cells (ATCC), which were pre-incubated with or without recombinant human CCL1 (Peprotech) and CCL17 (BioLegend) using primary polyclonal antibodies against human CCL17 (Thermo Fisher Scientific), human CCL1 (R&D Systems), human CCR4 (Thermo Fisher Scientific or BioLegend) and human CCR8 (Thermo Fisher Scientific). Cells were then treated with ligase and polymerase according to the manufacturer’s instructions of Duolink flowPLA Detection Far Red kit (Sigma-Aldrich). The fluorescent signal was analyzed by flow cytometry (FACSCanto II, BD Biosciences) and FlowJo v.10 software (Tree Star).
Expression, purification and labeling of CCL17
The gene encoding native CCL17 was inserted into a pET-32a(+) vector between Kpn I and Xho I restriction sites. The expression of recombinant CCL17 in One Shot BL21(DE3)Star E. coli in LB medium was induced by 0.1 mM IPTG when the OD600 reached 0.8–1.0. Inclusion bodies were isolated and resuspended in binding buffer (50 mM Tris, 500 mM NaCl, 4 M Gnd-HCl, 40 mM imidazole, 10 mM β-mercaptoethanol, pH 7.4). The extract was loaded on a HisTrap HP column (Cytiva Europe) equilibrated with equilibration buffer (50 mM Tris, 500 mM NaCl and 6 M Gnd-HCl, pH 7.4). After washing with 2% of elution buffer (50 mM Tris, 500 mM NaCl and 2 M imidazole, pH 7.4), the protein was eluted using a gradient of 2–50% elution buffer, followed by dialysis against acetic acid and lyophilization. The lyophilizate was resuspended in 10 mM Tris (pH 8.0), 3 U EKMax protease (Thermo Fisher Scientific) was added and the solution was incubated for 16 h at 37 °C to remove the tag. The cleaved product was further purified using a 3-ml RESOURCE RPC column with an acetonitrile + 0.1% TFA gradient. After lyophilization, the protein was refolded in 50 mM Tris, 10 mM cysteine and 0.5 mM cystine (pH 8.0) for 24 h at 4 °C under gentle stirring, purified using a HiTrap Heparin HP column and HPLC. The correct mass and folding were verified by mass spectrometry and NMR, respectively.
CCL17 was labeled using 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) and Alexa-Fluor 647 Cadaverine (Thermo Fisher Scientific). Briefly, CCL17 was incubated in presence of tenfold molar excess of EDC and dye in 10 mM MES (pH 6.0). After 10 min the labeled product was purified using Zeba Spin Desalting Columns (Life Technologies) and stored at 4 °C.
Surface plasmon resonance
SPR was performed on a BIAcore X100 instrument (Cytiva Europe) using neutravidin-modified C1 sensor chips69 coated with biotinylated human recombinant CCL17 or CCL5 to 1,300 resonance units. Sensograms were obtained by injecting different concentrations of CCR8-carrying proteoliposomes or with CCR4-carrying liposomes (positive control), mock protein-carrying or pure liposomes (negative controls) in running buffer (62.5–2,000 ng ml−1 in HBS-EP + buffer). Analytes were perfused over the chip for 270 s (at 20 µl min−1) followed by a final dissociation phase of 180 s. The sensor chip was regenerated with two pulses of 60 s of NAS (30% acetonitrile, 100 mM NaOH and 0.1% SDS). Responses from analyte injections were overlaid with the fit of 1:1 interaction model (Langmuir) determined using BIACORE X100 evaluation 2.0 software.
Competitive chemokine receptor-ligand binding
HEK293 cells stably transfected with human CCR8 (HA-tagged to monitor expression) and mock HEK293 cells (105 each) were incubated in binding buffer (HBSS supplemented with 20 mM HEPES and 0.2% BSA) with increasing concentrations of unlabeled human recombinant CCL1, CCL17 or CCL18. After 20 min at 4 °C, recombinant human CCL17 or synthetic human CCL1 (ALMAC) labeled with Alexa-Fluor 647 at the C terminus was added at a final concentration of 10 nM and further incubated for 30 min. After washing with binding buffer and fixation in 2% PFA/PBS, fluorescence intensity was measured by flow cytometry (FACSCanto II) and analyzed using FlowJo v.10 software (Ashland). Background binding to HEK293 mock-transfectants was subtracted and data were normalized to binding without unlabeled chemokine (control) and subjected to nonlinear fitting. Data represent mean ± s.d. from three independent experiments performed in duplicate or triplicate.
CCR8 and CCR4 internalization assay
For isolation of CD4+ T cells from Ccr8WTApoe−/− or Ccr8KOApoe−/− mice, thymus or LNs were mechanically crushed and passed through a 30-μm cell strainer (Cell-Trics, Partec) using Hank’s medium (HBSS + 0.3 mmol l−1 EDTA + 0.1% BSA; Gibco by Life Technologies) to obtain single-cell suspensions. Subsequently, CD4+ T cells were isolated using the mouse CD4+ T cell Isolation kit (Miltenyi Biotec). Cells (1 × 105) were incubated with recombinant murine CCL17 (100 ng ml−1), CCL1 (50 ng ml−1) or CCL22 (50 ng ml−1) at 37 °C for 1 h and surface stained with antibodies against CD4, CCR8 and CCR4 at 4 °C for 30 min. CCR8 fluorescence intensity of CD4+ T cells was analyzed by flow cytometry (FACSCanto II, BD Biosciences) and FlowJo v.10 software (Tree Star). Human CD4+ T cells were isolated from PBMCs with Dynabeads Untouched Human CD4 T Cells kit (Invitrogen) according to the manufacturer’s protocol as previously mentioned. Cells (1 × 105) were incubated with recombinant human CCL17 (100 ng ml−1), CCL1 (50 ng ml−1) or CCL22 (50 ng ml−1, all Peprotech) at 37 °C for 20 min or 40 min and surface stained with antibodies against CD4, CCR8 and CCR4 at 4 °C for 30 min. Cells were then washed with PBS. CCR8 and CCR4 fluorescence intensity of CD4+ T cells was analyzed by flow cytometry (FACSCanto II, BD Biosciences) and FlowJo v.10 software (Tree Star).
RNA purification and real-time PCR
Total mRNA was isolated from frozen mouse axillary LNs with Trizol (Invitrogen). Isolated RNA was subsequently transcribed into cDNA using an iScript cDNA Synthesis kit (Bio-Rad) according to the manufacturer’s instructions. Real-time PCR was then performed with QuantiNova Probe PCR kit (QIAGEN) in QuantStudio 6 Real-Time PCR system (Thermo Fisher). The threshold cycle (Ct) values of the target genes were normalized to that of the housekeeping gene (endogenous control) encoding 18S ribosomal RNA (rRNA). All data were analyzed by adopting 2-ΔΔCt method. Relative mRNA expression is shown, with the average from control samples set as 1. The TaqMan gene expression assays used in this study were Mm00441259_g1 (CCL3) and Mm03928990_g1 (Rn18S) (all Life Technologies).
Quantification of FoxP3 mRNA and CCL3 copy numbers in human plaque specimens was performed as described70,71,72 and correlated with the clinical phenotype, either defined by asymptomatic stable atherosclerosis or by symptomatic disease, as apparent by cerebral ischemic events, for example transient ischemic attacks or stroke. The following primers/probes were applied: FoxP3_h_fwd GCCCGGATGTGAGAAGGTCTT, FoxP3_h_rev GCCCTGCCCTTCTCATCCAG, FoxP3_h_Probe 5′FAM-CTTCCTCAAGCACTGCCAGGCGGAC-3′TAM; hCCL3-fwd: CTGCACCATGGCTCTCTGC; hCCL3-rev: CTGAAGCAGCAGGCGGTC, hCCL3-Probe:CTCTGCATCACTTGCTGCTGACACGC. The use of human material was approved by the local ethics committee (LMU Munich, no. 18–296). The study procedure was in accordance with the Helsinki Declaration and all participants provided their written informed consent.
Microarray data acquisition and data processing of published datasets
For the CCR8 mRNA expression in various human tissues and in different blood cell types, datasets were downloaded from the Human Protein Atlas (https://www.proteinatlas.org).
For the CCL3, CCR8 and FOXP3 mRNA expression in human plaques or T cells from patients with familial hypercholesterolemia, four microarray datasets (GSE43292 (ref. 73), GSE28829 (ref. 74), GSE11138 (ref. 75)) were downloaded from the Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo). The GSE43292 dataset contained 32 human atheroma plaques or 32 paired distant macroscopically intact tissue. The GSE28829 dataset consisted of 16 advanced atherosclerotic plaques and 13 early lesions. The GSE11138 dataset consisted of eight symptomatic and six asymptomatic patients with carotid and coronary plaque. Data preprocessing included transforming gene probes into gene symbols, data consolidation and normalization. Probes without gene symbols were deleted. Probes with maximal expression were retained for further analysis if the probes contained more than one probe.
Cell suspension preparation for scRNA-seq
For the scRNA-seq experiment of pooled lymph nodes (including mesenteric, para-aortic, inguinal, axillary, brachial and mandibular LNs), eight Ccl17wt/eApoe−/− and six Ccl17e/eApoe−/− male mice (C57BL/6J background) were fed a normal chow diet. Perfusion was performed with 5 ml PBS through left ventricular puncture until the liver yielded a pale color. Pooled LNs from either Ccl17wt/eApoe−/− or Ccl17e/eApoe−/− mice were mechanically crushed and passed through a 30-μm cell strainer (Cell-Trics, Partec) using Hank’s medium (HBSS + 0.04% BSA; Gibco by Life Technologies) to obtain single-cell suspensions, followed by B cell depletion using CD19 MicroBeads (Miltenyi Biotec) according to the manufacturer’s instructions. B cell-depleted fractions were stained with Fixable Viability Dyes eFluor 660, anti-CD45, anti-CD3, anti-CD19 and anti-CD11c (All from eBioscience) at 4 °C for 20 min. After washing in PBS for 5 min, cells were resuspended in Hank’s medium (HBSS + 0.04% BSA; Gibco by Life Technologies) and then isolated by fluorescence-activated cell sorting (BD FACSAria), by gating for live CD45+CD11c+CD3−CD19− cells. The cell suspension with viability >80% was ready for subsequent single-cell capture and library preparation.
For the scRNA-seq experiment of para-aortic LNs, seven Ccl17wt/eApoe−/− and ten Ccl17e/eApoe−/− male mice (C57BL/6J background) were fed a 6-week WD. Para-aortic LNs of same genotype were pooled and strained as previously mentioned. Cells were resuspended in Hank’s medium (HBSS + 0.04% BSA; Gibco by Life Technologies), stained and isolated by fluorescence-activated cell sorting (BD FACSAria), by gating for live CD45+CD19−MHCII+ cells. The cell suspension with viability > 80% was ready for subsequent single-cell capture and library preparation.
Single-cell RNA sequencing
Cell suspensions were loaded into a 10x Genomics Chromium Next GEM Chips and encapsulated with Single Cell 3ʹ v.3.1 barcoded gel beads using the 10x Genomics Chromium controller, according to the manufacturer’s instructions. Single-cell libraries were then constructed according to the manufacturer’s instructions. Libraries from individual samples were sequenced on an Illumina NovaSeq 6000 platform. The sequencing depth was set to around 50,000 reads per cell for pooled LNs from various positions and around 65,000 reads per cell for para-aortic LNs.
Analysis of scRNA-seq data
Fastq files of sorted CD45+CD11c+CD3−CD19− cells from LNs of Ccl17wt/eApoe−/− and Ccl17e/eApoe−/− mice on a chow diet were aligned to the customized reference genome (eGFP was added to the mm10 reference) individually using CellRanger Software (v.3.0.0, 10x Genomics). Individual datasets were then aggregated using the CellRanger aggr command without subsampling normalization. The aggregated dataset was then analyzed using the R package Seurat (v.3.1.4)76,77. The dataset was trimmed of cells expressing <200 or >5,000 genes for exclusion of non-cell or cell aggregates. Cells containing >10% mitochondrial genes were presumed to be of poor quality and were also discarded. A ‘logNormalize’ method was employed to normalize the gene expression for each cell by the total expression, the resulting expression values were then multiplied by 10,000 and log-transformed. The most highly variable genes in the dataset were discovered with FindVariableFeatures function and used in principal-component analysis (PCA), followed by a linear transformation (‘scaling’) following the standard pipeline. PCA was used for dimensionality reduction and UMAP was then used for two-dimensional visualization of the clusters. Visualization of gene expression with feature plot was generated with Seurat function FeaturePlot. Marker genes of each cluster were found by FindAllMarkers function.
Similar alignment was performed in sorted CD45+CD19−MHCII+ cells from para-aortic LNs of Ccl17wt/eApoe−/− and Ccl17e/eApoe−/− mice on a WD for 6 weeks individually using CellRanger Software (v.3.0.0., 10x Genomics). Individual datasets were then aggregated using the CellRanger aggr command without subsampling normalization. Cells expressing <200 or >4,000 genes were filtered out for the exclusion of non-cell or cell aggregates. Cells containing >5% mitochondrial genes were also discarded. Similar normalization, scaling, PCA, clustering and UMAP analysis were then performed.
Tolerogenic score
XCR1+ tolerogenic DCs undergo a continuous homeostatic maturation that is essential for central tolerance and that occurs irrespective of IFN-I according to previous study78. The 82 specifically upregulated genes during thymic and peripheral homeostatic XCR1+ DC maturation in this study were listed as tolerogenic genes. Interferon-stimulated genes were among the few discriminators of immunogenic and tolerogenic XCR1+ DCs. The 31 interferon-stimulated genes from this study are listed as immunogenic genes (Supplementary Table 3). The tolerogenic score was then calculated using the top 20 genes that distinguished tolerogenic DCs and immunogenic DCs as follows: tolerogenic score = (1 + mean (top 20 upregulated tolerogenic genes))/(1 + mean (top 20 upregulated immunogenic genes)).
Gene set variation analysis enrichment score
To generate a GSVA enrichment score of each eGFP-expressing CCL17-deficient cell from tolerogenic DCs of Apoe−/− Ccl17e/e mice and each CCL17-expressing cell from tolerogenic DCs of Apoe−/−Ccl17wt/e mice fed on chow diet, the ontology gene sets v.7.1 were provided in MSigDB (https://www.gsea-msigdb.org/gsea/msigdb). The analysis was implemented using R package gsva.
Statistical analysis
Statistical analyses were performed with GraphPad Prism v.10 (GraphPad Software) and IBM SPSS Statistics v.29.0 (IBM). Data distribution and homogeneity of variance were tested by the Shapiro–Wilk and Levene’s tests, respectively. Data violating the assumption of Gaussian distribution were analyzed by Mann–Whitney U-test (two-group comparisons) or Kruskal–Wallis H test with Dunn’s post hoc test. For normally distributed data, unpaired Student’s t-test with Welch correction when appropriate (two-group comparisons) or univariate ANOVA with Holm–Šídák’s post hoc test (three or more groups) was performed. In analyses involving two or more factors, factorial (two-way) ANOVA with Holm–Šídák’s post hoc test for pairwise comparisons was applied. Data from multiple biological replicates over independent experiments were analyzed with a nested approach by fitting mixed-effect regression based on generalized linear models with nested data (within the independent experiment) as random factors and robust estimation of the covariance matrix to avoid the influence of violation of model assumptions (nested ANOVA). Pairwise contrasts within fixed factors were corrected by step-down Holm–Šídák’s procedure. Main model effects were tested by Wald chi-squared test. For models involving dependency of measurements, the assumption of sphericity was verified with Mauchly’s W test, and the Greenhouse–Geisser correction was applied in case of violation. Differences were considered significant for a two-tailed P value < 0.05. Data were reported as mean ± s.e.m., unless otherwise stated. For mouse experiments, a priori calculation of power based on a two-sample t-test design and previous data or pilot experiments was performed with the software Java Applets for Power and Sample Size (available at http://www.stat.uiowa.edu/~rlenth/Power) and aimed at 80% statistical power for detecting biological relevant changes (30%) with a two-tailed α-value of 5%. For in vitro experiments, group sample sizes were determined empirically based on pilot experiments.
Materials
A detailed overview of used materials is provided in the Major Resources Table (Supplementary Table 12).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data associated with this study are presented within the paper and associated files. The scRNA-seq data of Ccl17wt/eApoe−/− and Ccl17e/eApoe−/− mice fed on a chow diet or a WD for 6 weeks have been deposited in Gene Expression Omnibus and are available under accession code GSE200862. The following publicly available datasets for human atherosclerosis were included: GSE28829 (advanced atherosclerotic plaques or early lesions), GSE43292 (human atheroma or paired distant macroscopically intact tissue) and GSE11138 (symptomatic or asymptomatic patients with carotid and coronary plaques). Schematic panels in the figures were created using www.biorender.com. Source data are provided with this paper.
References
Lutgens, E. et al. Immunotherapy for cardiovascular disease. Eur. Heart J. 40, 3937–3946 (2019).
Jongstra-Bilen, J. et al. Low-grade chronic inflammation in regions of the normal mouse arterial intima predisposed to atherosclerosis. J. Exp. Med. 203, 2073–2083 (2006).
Millonig, G. et al. Network of vascular-associated dendritic cells in intima of healthy young individuals. Arter. Thromb. Vasc. Biol. 21, 503–508 (2001).
Choi, J. H. et al. Identification of antigen-presenting dendritic cells in mouse aorta and cardiac valves. J. Exp. Med. 206, 497–505 (2009).
Gil-Pulido, J. & Zernecke, A. Antigen-presenting dendritic cells in atherosclerosis. Eur. J. Pharmacol. 816, 25–31 (2017).
Ye, Y. et al. Serum chemokine CCL17/thymus activation and regulated chemokine is correlated with coronary artery diseases. Atherosclerosis 238, 365–369 (2015).
Brunner, P. M. et al. The atopic dermatitis blood signature is characterized by increases in inflammatory and cardiovascular risk proteins. Sci. Rep. 7, 8707 (2017).
He, H. et al. Increased cardiovascular and atherosclerosis markers in blood of older patients with atopic dermatitis. Ann. Allergy Asthma Immunol. 124, 70–78 (2020).
Ye, Y. et al. Association between a CCL17 genetic variant and risk of coronary artery disease in a Chinese Han population. Circulation 82, 224–231 (2017).
Weber, C. et al. CCL17-expressing dendritic cells drive atherosclerosis by restraining regulatory T cell homeostasis in mice. J. Clin. Investig. 121, 2898–2910 (2011).
Heiseke, A. F. et al. CCL17 promotes intestinal inflammation in mice and counteracts regulatory T cell-mediated protection from colitis. Gastroenterology 142, 335–345 (2012).
Steinman, R. M. Decisions about dendritic cells: past, present, and future. Annu. Review. Immunol. 30, 1–22 (2012).
Alferink, J. et al. Compartmentalized production of CCL17 in vivo: strong inducibility in peripheral dendritic cells contrasts selective absence from the spleen. J. Exp. Med. 197, 585–599 (2003).
Saigusa, R., Winkels, H. & Ley, K. T cell subsets and functions in atherosclerosis. Nat. Rev. Cardiol. 17, 387–401 (2020).
Arce-Sillas, A. et al. Regulatory T cells: molecular actions on effector cells in immune regulation. J. Immunol. Res. 2016, 1720827 (2016).
Sharma, M. et al. Regulatory T cells license macrophage pro-resolving functions during atherosclerosis regression. Circ. Res. 127, 335–353 (2020).
Ait-Oufella, H. et al. Natural regulatory T cells control the development of atherosclerosis in mice. Nat. Med. 12, 178–180 (2006).
Imai, T. et al. The T cell-directed CC chemokine TARC is a highly specific biological ligand for CC chemokine receptor 4. J. Biol. Chem. 272, 15036–15042 (1997).
Stutte, S. et al. Requirement of CCL17 for CCR7- and CXCR4-dependent migration of cutaneous dendritic cells. PNAS 107, 8736–8741 (2010).
Wild, A. B. et al. CD83 orchestrates immunity toward self and non-self in dendritic cells. JCI Insight 4, e126246 (2019).
Wan, W., Lionakis, M. S., Liu, Q., Roffe, E. & Murphy, P. M. Genetic deletion of chemokine receptor Ccr7 exacerbates atherogenesis in ApoE-deficient mice. Cardiovasc.Res. 97, 580–588 (2013).
Bernardini, G. et al. Identification of the CC chemokines TARC and macrophage inflammatory protein-1 β as novel functional ligands for the CCR8 receptor. Eur. J. Immunol. 28, 582–588 (1998).
Garlisi, C. G. et al. The assignment of chemokine-chemokine receptor pairs: TARC and MIP-1 β are not ligands for human CC-chemokine receptor 8. Eur. J. Immunol. 29, 3210–3215 (1999).
Barsheshet, Y. et al. CCR8+FOXp3+ Treg cells as master drivers of immune regulation. PNAS 114, 6086–6091 (2017).
Iellem, A. et al. Unique chemotactic response profile and specific expression of chemokine receptors CCR4 and CCR8 by CD4+CD25+ regulatory T cells. J. Exp. Med. 194, 847–853 (2001).
Uhlen, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Thornton, A. M. et al. Expression of Helios, an Ikaros transcription factor family member, differentiates thymic-derived from peripherally induced Foxp3+ T regulatory cells. J. Immunol. 184, 3433–3441 (2010).
Li, J. et al. CCR5+T-bet+FoxP3+ effector CD4 T cells drive atherosclerosis. Circ. Res. 118, 1540–1552 (2016).
Soehnlein, O. et al. Distinct functions of chemokine receptor axes in the atherogenic mobilization and recruitment of classical monocytes. EMBO Mol. Med. 5, 471–481 (2013).
de Jager, S. C. et al. Leukocyte-specific CCL3 deficiency inhibits atherosclerotic lesion development by affecting neutrophil accumulation. Arter. Thromb. Vasc. Biol. 33, e75–e83 (2013).
Schneider, M. A., Meingassner, J. G., Lipp, M., Moore, H. D. & Rot, A. CCR7 is required for the in vivo function of CD4+ CD25+ regulatory T cells. J. Exp. Med. 204, 735–745 (2007).
Worbs, T., Hammerschmidt, S. I. & Forster, R. Dendritic cell migration in health and disease. Nat. Rev. Immunol. 17, 30–48 (2017).
Klingenberg, R. et al. Depletion of FOXP3+ regulatory T cells promotes hypercholesterolemia and atherosclerosis. J. Clin. Investig. 123, 1323–1334 (2013).
Mor, A., Luboshits, G., Planer, D., Keren, G. & George, J. Altered status of CD4+CD25+ regulatory T cells in patients with acute coronary syndromes. Eur. Heart J. 27, 2530–2537 (2006).
Zhang, W. C. et al. Impaired thymic export and increased apoptosis account for regulatory T cell defects in patients with non-ST segment elevation acute coronary syndrome. J. Biol. Chem. 287, 34157–34166 (2012).
Albany, C. J., Trevelin, S. C., Giganti, G., Lombardi, G. & Scotta, C. Getting to the heart of the matter: the role of regulatory T-cells (Tregs) in cardiovascular disease (CVD) and atherosclerosis. Front. Immunol. 10, 2795 (2019).
Lee, I. et al. Recruitment of Foxp3+ T regulatory cells mediating allograft tolerance depends on the CCR4 chemokine receptor. J. Exp. Med. 201, 1037–1044 (2005).
Fox, J. M. et al. Structure/function relationships of CCR8 agonists and antagonists. Amino-terminal extension of CCL1 by a single amino acid generates a partial agonist. J. Biol. Chem. 281, 36652–36661 (2006).
Freeman, C. M. et al. CCR8 is expressed by antigen-elicited, IL-10-producing CD4+CD25+ T cells, which regulate Th2-mediated granuloma formation in mice. J. Immunol. 174, 1962–1970 (2005).
Sebastiani, S. et al. Chemokine receptor expression and function in CD4+ T lymphocytes with regulatory activity. J. Immunol. 166, 996–1002 (2001).
Inngjerdingen, M., Damaj, B. & Maghazachi, A. A. Human NK cells express CC chemokine receptors 4 and 8 and respond to thymus and activation-regulated chemokine, macrophage-derived chemokine, and I-309. J. Immunol. 164, 4048–4054 (2000).
Puttur, F. et al. Pulmonary environmental cues drive group 2 innate lymphoid cell dynamics in mice and humans. Sci. Immunol. 4, eaav7638 (2019).
Sokol, C. L., Camire, R. B., Jones, M. C. & Luster, A. D. The chemokine receptor CCR8 promotes the migration of dendritic cells into the lymph node parenchyma to initiate the allergic immune response. Immunity 49, 449–463 (2018).
De Simone, M. et al. Transcriptional landscape of human tissue lymphocytes unveils uniqueness of tumor-infiltrating T regulatory cells. Immunity 45, 1135–1147 (2016).
Eruslanov, E. et al. Expansion of CCR8+ inflammatory myeloid cells in cancer patients with urothelial and renal carcinomas. Clinical Cancer Res. 19, 1670–1680 (2013).
Mitsuyama, E., Kunori, Y., Kamimura, T. & Kaminuma, O. Functional chemokine receptors in allergic diseases: is CCR8 a novel therapeutic target? Mini Rev. Med. Chem. 6, 463–466 (2006).
Islam, S. A. et al. Mouse CCL8, a CCR8 agonist, promotes atopic dermatitis by recruiting IL-5+ T(H)2 cells. Nat. Immunol. 12, 167–177 (2011).
Vila-Caballer, M. et al. Disruption of the CCL1–CCR8 axis inhibits vascular Treg recruitment and function and promotes atherosclerosis in mice. J. Mol. Cell. Cardiol. 132, 154–163 (2019).
Tiffany, H. L. et al. Identification of CCR8: a human monocyte and thymus receptor for the CC chemokine I-309. J. Exp. Med. 186, 165–170 (1997).
Islam, S. A., Ling, M. F., Leung, J., Shreffler, W. G. & Luster, A. D. Identification of human CCR8 as a CCL18 receptor. J. Exp. Med. 210, 1889–1898 (2013).
Liu, L. et al. Biological characterization of ligands targeting the human CC chemokine receptor 8 (CCR8) reveals the biased signaling properties of small molecule agonists. Biochem. Pharmacol. 188, 114565 (2021).
Haas, J. et al. Specific recruitment of regulatory T cells into the CSF in lymphomatous and carcinomatous meningitis. Blood 111, 761–766 (2008).
Oo, Y. H. et al. Distinct roles for CCR4 and CXCR3 in the recruitment and positioning of regulatory T cells in the inflamed human liver. J. Immunol. 184, 2886–2898 (2010).
Feng, G. et al. CCL17 aggravates myocardial injury by suppressing recruitment of regulatory T cells. Circulation 145, 765–782 (2022).
Goldie, S. J. et al. Loss of GRHL3 leads to TARC/CCL17-mediated keratinocyte proliferation in the epidermis. Cell Death Dis. 9, 1072 (2018).
Chen, L., Wu, J., Pier, E., Zhao, Y. & Shen, Z. mTORC2-PKBα/Akt1 serine 473 phosphorylation axis is essential for regulation of FOXP3 stability by chemokine CCL3 in psoriasis. J. Investig. Dermatol. 133, 418–428 (2013).
Yang, L.-X. et al. Atorvastatin inhibits the 5-lipoxygenase pathway and expression of CCL3 to alleviate atherosclerotic lesions in atherosclerotic ApoE knockout mice. J. Cardiovasc. Pharmacol. 62, 205–211 (2013).
Ishida, Y. et al. Prevention of CaCl2-induced aortic inflammation and subsequent aneurysm formation by the CCL3–CCR5 axis. Nat. Commun. 11, 5994 (2020).
Braunersreuther, V. et al. Ccr5 but not Ccr1 deficiency reduces development of diet-induced atherosclerosis in mice. Arterioscler. Thromb. Vasc. Biol. 27, 373–379 (2007).
Bachelerie, F. et al. International union of basic and clinical pharmacology. [corrected]. LXXXIX. Update on the extended family of chemokine receptors and introducing a new nomenclature for atypical chemokine receptors. Pharmacol. Rev. 66, 1–79 (2014).
Huser, N. et al. CCR4-deficient mice show prolonged graft survival in a chronic cardiac transplant rejection model. Eur. J. Immunol. 35, 128–138 (2005).
Zernecke, A. et al. Deficiency in CCR5 but not CCR1 protects against neointima formation in atherosclerosis-prone mice: involvement of IL-10. Blood 107, 4240–4243 (2006).
Doring, Y. et al. CXCL12 derived from endothelial cells promotes atherosclerosis to drive coronary artery disease. Circulation 139, 1338–1340 (2019).
Park, I. et al. C-type lectin receptor CLEC4A2 promotes tissue adaptation of macrophages and protects against atherosclerosis. Nat. Commun. 13, 215 (2022).
Domogalla, M. P., Rostan, P. V., Raker, V. K. & Steinbrink, K. Tolerance through education: how tolerogenic dendritic cells shape immunity. Front. Immunol. 8, 1764 (2017).
Vendelova, E. et al. Tolerogenic transcriptional signatures of steady-state and pathogen-induced dendritic cells. Front. Immunol. 9, 333 (2018).
Kryczanowsky, F., Raker, V., Graulich, E., Domogalla, M. P. & Steinbrink, K. IL-10-modulated human dendritic cells for clinical use: identification of a stable and migratory subset with improved tolerogenic activity. J. Immunol. 197, 3607–3617 (2016).
Feierler, J. et al. Helix 8 plays a crucial role in bradykinin B(2) receptor trafficking and signaling. J. Biol. Chem. 286, 43282–43293 (2011).
von Hundelshausen, P. et al. Heterophilic interactions of platelet factor 4 and RANTES promote monocyte arrest on endothelium. Blood 105, 924–930 (2005).
Holdt, L. M. et al. ANRIL expression is associated with atherosclerosis risk at chromosome 9p21. Arterioscler. Thromb. Vasc. Biol. 30, 620–627 (2010).
Holdt, L. M. et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat. Commun. 7, 12429 (2016).
El Housni, H., Heimann, P., Parma, J. & Vassart, G. Single-nucleotide polymorphism genotyping by melting analysis of dual-labeled probes: examples using factor V Leiden and prothrombin 20210A mutations. Clin. Chem. 49, 1669–1672 (2003).
Ayari, H. & Bricca, G. Identification of two genes potentially associated in iron-heme homeostasis in human carotid plaque using microarray analysis. J. Biosci. 38, 311–315 (2013).
Doring, Y. et al. Auto-antigenic protein-DNA complexes stimulate plasmacytoid dendritic cells to promote atherosclerosis. Circulation 125, 1673–1683 (2012).
Cagnin, S. et al. Reconstruction and functional analysis of altered molecular pathways in human atherosclerotic arteries. BMC Genomics 10, 13 (2009).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Ardouin, L. et al. Broad and largely concordant molecular changes characterize tolerogenic and immunogenic dendritic cell maturation in thymus and periphery. Immunity 45, 305–318 (2016).
Acknowledgements
We are indebted to A. Rot (Queen Mary University) for thoughtful discussions. We thank L. Saroyan, O. Schengel and staff at the animal facility (all LMU Munich) for excellent technical assistance. We thank S. Michaelides (LMU Munich) for generously providing Jurkat CCR4- and CCR8-transfectants, K. Pfeffer (Heinrich-Heine-Universität) for kindly providing Ccr4−/− mice and I. Förster (Universität Bonn) for generously providing Ccl17e/e (GFP reporter knock-in) mice. This work was supported by Deutsche Forschungsgemeinschaft (DFG) SFB1123-A1/A10 to Y.D. and C.W., SFB1123-A2 to P.v.H., SFB1123-B1 to D.T. and L.H., SFB1123-Z1 to R.T.A.M., SFB1123-A5 to D.A., SFB1123-B5 to D.S. and 390857198-EXC 2145 to C.W.; by the European Research Council (AdG °692511 to C.W.); by the German Ministry of Education and Research (BMBF) and the German Center for Cardiovascular Research (DZHK) to Y.D., C.W., D.S. and E.P.C.v.d.V. (81X2600254, 81Z0600202 and 81Z0600203); by grants from the Interdisciplinary Center for Clinical Research within the faculty of Medicine at the RWTH Aachen University and NWO-ZonMw Veni (91619053) to E.P.C.v.d.V.; by the DFG YI 133/3-5 and Friedrich-Baur-Stiftung 39/20 to C.Y.; by the DFG HA 1083/15-4, HA 1083/15-5 and ERA-CVD (PLAQUEFIGHT) 01KL1808 to A.J.R.H. and ERA-CVD (AtheroInside) 01KL1908 to R.T.A.M. The position of L.P. was kindly supported by DZHK (81X2600271). C.W. is van der Laar-Professor of Atherosclerosis.
Author information
Authors and Affiliations
Contributions
Y.D. conceived and supervised the study, designed experiments, provided funding performed experiments, analyzed data and wrote the manuscript. E.P.C.v.d.V. supervised the study, designed experiments, performed experiments, analyzed data and contributed to writing the manuscript. Y.Y. and C.N. designed experiments, performed experiments and analyzed the data. Y.J. performed and analyzed mouse experiments and data. X.B., J.L. and Y.L. performed and analyzed plasmon resonance and receptor binding assays. M.K., S.B., L.J.F.P., S.G., L.P. and R.T.A.M. contributed to the conduction of mouse experiments and data analysis. M.H. and K.B. performed cell culture and cell-sorting experiments; C.Y., X.Z. and A.J.R.H. contributed to scRNA-seq experiments and data analysis. A.F. helped to perform and analyze cAMP signaling assays and provided critical reagents. D.T. and L.H. provided human plaque material and analysis. C.M. and I.P. provided critical reagents, contributed to scRNA-seq experiments and data analysis and provided intellectual input. M.K., K.N. and D.A. contributed to Treg suppression and T cell migration studies. D.S. provided critical revision of the manuscript, analyzed data and calculated statistics. P.v.H. contributed to receptor binding experiments, analyzed data, provided supervision and intellectual input, and contributed to writing the manuscript. C.W. conceived and supervised the study, designed experiments, provided funding and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Christoph Binder, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effects of CCL17 deficiency on atherosclerosis and Treg numbers.
(a) Experimental scheme of Apoe−/− or Apoe−/−Ccl17e/e mice fed a Western-diet (WD) for 12 weeks; (b) Representative images and quantification of lesion area of Apoe−/− (n = 16) or Apoe−/−Ccl17e/e (n = 14) mice measured after Oil-Red-O staining (ORO) for lipid deposits in the aortic root. Scale bar = 500 µm; (c) Quantification of lesion area measured after ORO for lipid deposits in the thoraco-abdominal of Apoe−/− (n = 13) or Apoe−/−Ccl17e/e (n = 14); (d) Atherosclerotic lesion size in aortic arches of Apoe−/− (n = 15) or Apoe−/−Ccl17e/e (n = 12), as quantified after H&E staining; (e-g) Quantification of the percentage of lesional Mac2+ macrophages (e), smooth muscle α-actin (SMA)+ smooth muscle cells (SMCs) (f) and collagen content (g) of Apoe−/− (n = 11) or Apoe−/−Ccl17e/e (n = 10) mice fed a WD for 12 weeks; (h-i) Representative dot plots and flow cytometric quantification of CD45+CD3+CD4+ CD25+FoxP3+ Tregs in para-aortic lymph nodes (LNs) (h) and spleen (i) of Apoe−/− (h, n = 8; i, n = 11) or Apoe−/−Ccl17e/e (h, n = 8; i, n = 10) mice; (j) Gating strategy for CD45+CD3+ CD4+CD25+FoxP3+ Tregs in lymphatic organs; (k-l) flow cytometric quantification of CD45+CD3+CD4+CD25+FoxP3+ Tregs in axillary (k) and inguinal LNs (l) of Apoe−/− (k, n = 11; l, n = 10) or Apoe−/−Ccl17e/e (k, n = 10; l, n = 9) mice; (a-l) Data represent mean ± SEM. Two-sided P values as indicated and analyzed by unpaired Student’s t-test.
Extended Data Fig. 2 Effects of CCR4 deficiency on atherosclerosis and CCL17-induced migration via CCR4.
(a) Experimental scheme of Apoe−/− or Apoe−/−Ccr4−/− mice fed a Western-diet (WD) for 12 weeks; (b, c) Flow cytometric quantification of CD45+CD3+CD4+CD25+FoxP3+ Tregs, axillary (b) and inguinal LNs (c) of Apoe−/− (b, n = 13; c, n = 16) or Apoe−/−Ccr4−/− (b, n = 18; c, n = 20) mice after 12 weeks of WD; (d-f) Quantification of the percentage of lesional Mac2+ macrophages (d), SMA+ SMCs (e) and collagen content (f) in Apoe−/− (d, e n = 20; f, n = 19) or Apoe−/−Ccr4−/− (d, n = 19; e, n = 20; f = 16) mice fed a WD for 12 weeks; (g) Analysis of Annexin-V (AnnV) expression on Tregs (CD45+CD3+CD4+CD25+FoxP3+) from isolated LNs (para-aortic, axillary and inguinal combined) of Apoe−/− (n = 8) or Apoe−/−Ccl17e/e (n = 6); (h) Transwell migration assay with CD4+ T cells (isolated from Apoe−/− mice) towards recombinant mouse CCL17 (100 ng/ml) or CCL22 (50 ng/ml) in the presence or absence of the CCR4 inhibitor C021 dihydrochloride (0.5 µM), number of replicates in parentheses over number of independent experiments per bar from left to right: n = (40)8, (40)8, (27)6, (27)6, (25)5, (25)5; (i) Transwell migration assay with CD4+ T cells (isolated from human blood PBMCs) towards recombinant human CCL17 (100 ng/ml) or CCL22 (50 ng/ml), in the presence or absence of the CCR4 inhibitor C021 dihydrochloride (0.5 µM), number of replicates in parentheses over number of independent experiments per bar from left to right: n = (37)7, (33)7, (31)5, (28)5, (16)4, (16)4; (h,i) All migrated cells were quantified by flow cytometry, chemotactic index calculated as the ratio of chemokine-stimulated and unstimulated migration is depicted; (a-i) Data represent mean ± SEM. Two-sided P values as indicated and analyzed by unpaired Student’s t-test, Mann-Whitney U (b-h), or nested ANOVA with Holm-Šídák’s post hoc test (h, i).
Extended Data Fig. 3 Identification of tolerogenic DCs in the LNs of Apoe−/−Ccl17wt/e or Apoe−/−Ccl17e/e mice.
(a) UMAP projection of 4,731 single cells, colored by inferred cell types, in sorted cells (viable CD45+CD3−CD11c+) from LNs of Apoe−/−Ccl17wt/e or Apoe−/−Ccl17e/e mice; (b) UMAP visualization overlaid with the expression of eGFP+ Apoe−/−Ccl17wt/e (left panels) or Apoe−/−Ccl17e/e (right panel); (c) Heat map of the top 20 marker genes from each cluster and cell type assignment of each cluster; (d-f) UMAP visualization overlaid with the expression of Aldh1a2 (d), Cd83 (e) and Cd274 (f) in 7 distinct DC clusters of sorted cells (viable CD45+CD3−CD11c+) from LNs of Apoe−/−Ccl17wt/e (left panel) or Apoe−/−Ccl17e/e (right panel) mice; (g) UMAP projection of single cells, colored by inferred cell types including tolerogenic DCs and other DCs, in sorted cells (viable CD45+CD3−CD11c+) from LNs of Apoe−/−Ccl17wt/e or Apoe−/−Ccl17e/e mice; (h-j) Representative dot plots and flow cytometric quantification of CD83+ (h), CCR7+ (i) and CD83+CCR7+ (j) tolerogenic DCs among CD45+CD11c+ MHCII+ cDCs in aortic LNs of Apoe−/− or Apoe−/−Ccl17e/e mice (h-j; each bar n = 7); (k-m) GSVA score was calculated in GO term CCR chemokine receptor binding (k), myeloid leukocyte migration (l) and positive regulation of acute inflammatory response (m) in n = 172 eGFP-expressing CCL17-deficient cells from tolerogenic DCs of Apoe−/−Ccl17e/e mice or n = 92 CCL17-expressing cells from tolerogenic DCs of Apoe−/−Ccl17wt/e mice fed on a chow diet; (n) Sorted CD19+B220+ B cells from lymph nodes (LNs), isolated CD115+ monocytes and Ly6G+ neutrophils from spleen and bone marrow were cultured for 4 hours in the presence or absence of recombinant mouse CCL17 (100 ng/ml). CCL3 concentrations in the supernatant were measured by multiplex bead array (each condition n = 3); (a-n) Data represent mean ± SEM. Two-sided P values as indicated and analyzed compared to control by Mann-Whitney U-test (h-j), unpaired Student’s t-test (k-m), or two-way ANOVA with Holm-Šídák’s post hoc test (n).
Extended Data Fig. 4 Analysis of CCR8 ligand binding, internalization and migration capacities.
(a, b) Interactions between mouse CCL17 or CCL1 and CCR4, CCR5 or CCR8 were assessed on the surface of adherent cDCs isolated from LNs of Apoe−/− mice using Duolink proximity-ligation assay after incubation with recombinant mouse CCL17, CCL1 (100 ng/ml) or PBS (control) and respective antibodies to CCR4, CCR5 and CCR8, as indicated. (a) Shown are representative images recorded with a Leica SP8 confocal microscope for anti-CCR8 and anti-CCL17 after PBS and CCL17 treatment (scale bar = 10 µm); (b) Signals generated by interactions between ligands and receptors on the cDC surface were quantified and normalized to untreated controls (dotted line); number of independent experiments per bar from left to right: n = 5, 4, 4, 4, 3, 5; (c) Representative histograms displaying CCR8 expression in HEK293 CCR8-transfectants and controls; (d,e) CCR4 internalization in CCR8-deficient or CCR8-competent CD4+ T cells from thymus (d, number of independent experiments per bar from left to right: n = 3, 4, 3, 4) and LNs (e, number of independent experiments per bar from left to right: n = 3, 4, 3, 4) stimulated with CCL17 (100 ng/ml) or vehicle and analyzed by flow cytometry; (f) Dose-response curve to compare CCR8 internalization by CCL1 and CCL17 at indicated concentrations in primary CCR8-expressing human T cells; (g) Representative histograms displaying CCR4 expression in HEK293 CCR4-transfectants and controls; (h) Area under the curve (AUC) for Rluc in Glosensor assays monitoring cAMP levels in HEK293 CCR8-transfectants to obtain dose-response curves for CCL1, CCL17, CCL18 or irrelevant CCL20 using indicated concentrations (CCL1, CCL18, CCL20, n = 6 independent experiments; CCL17, n = 3 independent experiments); (i) Transwell migration assay with human CD4+ T cells towards recombinant human CCL17 (100 ng/ml) or CCL1 (50 ng/ml) in the presence or absence of a blocking antibody to CCR8 (2 µg/ml). Migrated cells were quantified by flow cytometry; number of replicates in parentheses over number of independent experiments per bar from left to right: n = (40)8, (16)5, (46)8, (21)5, (18)5, (18)5; (j-l) Transwell migration assays depicted as dose-response curves of Apoe−/−CD4+ T cell migration induced by CCL1 (j), CCL17 (k) or CCL22 (l) at indicated concentrations in the presence or absence of anti-CCR8 antibody (2 µg/ml) or C021 (0.5 µM) (j-l, n = 4 independent experiments; CCL17+anti-CCR8, n = 3 independent experiments). (a-l) Data represent mean ± SEM. Two-sided P values as indicated and analyzed by Mann-Whitney U-test or unpaired Student’s t-test (b), two-way ANOVA (d,e) or nested ANOVA (i) with Holm-Šídák’s post hoc test (d,e), generalized linear model (f).
Extended Data Fig. 5 Analysis of CCR8 transcript expression across tissues and cell types and consequences of CCR8 blocking.
(a) CCR8 mRNA expression in different tissues of consensus datasets from The Human Protein Atlas. Tissues of the same system (nervous system, endocrine system, digestion system, etc.) are depicted in the same color; (b) CCR8 mRNA expression in different blood cell types of consensus datasets from The Human Protein Atlas; (c) Ccr8 mRNA expression in different T cell populations of the para-aortic LNs from Apoe−/−Ccl17e/wt and Apoe−/−Ccl17e/e mice fed a WD for 6 weeks; (d) Flow cytometric quantification of CCR8 expression on DCs from LN of the indicated mouse strains (n = 5 per strain); (e) CCL3 titers in supernatants of CD4+ T cells from Apoe−/− mice stimulated with CCL17 or vehicle for 4 hours in the presence or absence of an antibody to CCR8 (number of replicates in parentheses over number of independent experiments per bar from left to right: n = 8, 8, 7, 7); (f) Experimental scheme of Apoe−/− mice injected with isotype control or CCR8-blocking antibody during WD feeding for 4 weeks. (g-i) Quantification of the percentage of lesional Mac2+ macrophages (g, isotype, n = 9; anti-CCR8, n = 5), SMA+ SMCs (h, isotype, n = 9; anti-CCR8, n = 6) and collagen content (i, isotype, n = 10; anti-CCR8, n = 5) of Apoe−/− mice receiving an isotype or CCR8 blocking antibody during WD feeding for 4 weeks. (a-i) Data represent mean ± SEM. Two-sided P values as indicated and analyzed by Mann-Whitney U-test or unpaired Student’s t-test (c, g-i) or two-way ANOVA with Holm-Šídák’s post hoc test (e).
Extended Data Fig. 6 Effect of systemic and T cell-specific CCR8 deficiency on atherosclerosis and analysis of CCR1 and CCR5 expression in different T cell subsets.
(a) Experimental scheme of CD4Cre-Ccr8flox/floxApoe−/− (CD4Ccr8WTApoe−/−) and CD4Cre+Ccr8flox/flox Apoe−/− (CD4Ccr8KOApoe−/−) mice fed a WD for 12 weeks; (b) Representative images and quantification of lesion area measured after HE-staining in the aortic root of CD4Ccr8WT Apoe−/− (n = 16) or CD4Ccr8KO Apoe−/− (n = 9) mice. Scale bar = 500 µm; (c) Quantification of lesion area measured after Oil-Red-O staining for lipid deposits in the thoraco-abdominal aorta of CD4Ccr8WT Apoe−/− (n = 17) or CD4Ccr8KO Apoe−/− (n = 8) mice. (d) Atherosclerotic lesion size in the aortic arch, as quantified after H&E staining in CD4Ccr8WT Apoe−/− (n = 14) or CD4Ccr8KO Apoe−/− (n = 8) mice. (e) Experimental scheme of CD4Ccr8WT Apoe−/− or CD4Ccr8KO Apoe−/−mice fed a WD for 12 weeks; (f-j) Representative images and quantification of the percentage of lesional Mac2+ macrophages (f,g) or SMA+ SMCs (h,i) and collagen content (j) in aortic arch sections of CD4Ccr8WT Apoe−/− (g,i,j all n = 20) or CD4Ccr8KO Apoe−/− (g,i,j all n = 23) mice fed a WD for 12 weeks. Scale bar = 250 µm; (k,l) Ccr1 and Ccr5 mRNA expression in different T cell populations of the para-aortic LNs from Apoe−/−Ccl17e/wt and Apoe−/−Ccl17e/e mice fed a WD for 6 weeks; (m) Flow cytometric quantification of CD45+CD3+CD4+FoxP3+Tbet+ cells in aortic, axillary and mesenteric LNs of Apoe−/−, Apoe−/−Ccl3−/− or Apoe−/−Ccl17e/e mice (all n = 8). (a-m). Data represent mean ± SEM. Two-sided P values as indicated and analyzed by by Mann-Whitney U test or unpaired Student’s t-test (b-j), multiple Mann-Whitney U test with false discovery rate (FDR) (k,l), one-way ANOVA with Holm-Šídák’s post hoc test (aortic and axillary LNs) or Kruskal-Wallis H with Dunn’s post hoc test (mesenteric LNs) (m).
Extended Data Fig. 7 Effect of CCR1 and CCL3 on lesional characteristics and Treg numbers.
(a) Representative images of staining for Mac2+ macrophages, SMA+ SMCs and DAPI (nuclei) in aortic root sections of Apoe−/− or Apoe−/−Ccr1−/− mice after 12 weeks of WD. Scale bar = 250 µm; (b-d) Quantification of the percentage of lesional Mac2+ macrophages (b), SMA+ SMCs (c) and collagen content (d) in Apoe−/− (b, n = 8; c, n = 7; d, n = 7) or Apoe−/−Ccr1−/− (b-d, n = 5 each) mice after 12 weeks of WD; (e-f) Quantification of Tregs (CD45+CD3+CD4+CD25+ FoxP3+) in axillary (e) and inguinal LNs (f) of Apoe−/− (e, n = 7; f, n = 8) Apoe−/−Ccr1−/− (e, n = 5; f, n = 4) mice after 12 weeks of WD; (g) Plasma concentrations of CCL3 in Apoe−/− (n = 7) or Apoe−/−Ccr1−/− (n = 5) mice after 12 weeks of WD, as measured by ELISA; (h-k) Quantification of Tregs (CD45+CD3+CD4+CD25+FoxP3+) in aortic LN (h), spleen (i), axillary (j) and inguinal LNs (k) of C57Bl6 (h, n = 9; i, n = 6; j, n = 10; k, n = 9) or Ccl3−/− (h-k, n = 10 each) mice; (l,m) Quantification of Tregs (CD45+CD3+CD4+ CD25+FoxP3+) in axillary (l) and inguinal LNs (m) of Apoe−/− (l, n = 9; m, n = 10) or Apoe−/−Ccl3−/− (l,m, n = 13 each) mice, after 12 weeks of WD; (n) Representative images of staining for Mac2+ macrophages, SMA+ SMCs and DAPI (nuclei) in aortic root sections of Apoe−/− or Apoe−/−Ccl3−/− mice after 12 weeks of WD. Scale bar = 250 µm. (n-p) Quantification of the percentage of lesional Mac2+ macrophages (n), SMA+ SMCs (o) and collagen content (p) in aortic root sections of Apoe−/− (n, n = 9; o, n = 9; p, n = 10) or Apoe−/−Ccl3−/− (n, n = 8; o, n = 13; p, n = 13) mice after 12 weeks of WD. (a-p). Data represent mean ± SEM. Two-sided P values as indicated and compared to Apoe−/− or Bl6 control mice analyzed by unpaired Student’s t-test or Mann Whitney U test (b-p).
Extended Data Fig. 8 CCL3 and FOXP3 mRNA expression in human plaques and synopsis of the proposed pathway.
(a) mRNA expression of CCL3 in advanced atherosclerotic plaques (n = 16) or early (n = 13) lesions derived from GSE28829 dataset; (b) mRNA expression of CCL3 in human atheroma plaque (atheroma) or paired distant macroscopically intact tissue (adjacent) derived from GSE43292 dataset (n = 32 each); (c) CCL3 mRNA expression in symptomatic (n = 8) or asymptomatic (n = 6) patients with carotid and coronary plaques derived from GSE11138 dataset; (d,e) Quantification of CCL3 (d) and FOXP3 (e) mRNA copy numbers normalized to housekeeping mRNA (105 GAPDH or β-actin mRNA copies, respectively) in atherosclerotic lesions of carotid atherectomy specimens from symptomatic (d, e n = 13) or asymptomatic (d, n = 16; e, n = 15) patients using real-time PCR; (a-e) Data represent mean ± SEM. Two-sided P values as indicated versus corresponding controls, as analyzed by unpaired Student’s t-test (a-c), or Mann Whitney U test (d, e); (f) Pathway synopsis (I.) Sterile inflammation triggers the activation of a subset of cDCs, which respond by releasing CCL17. (II.) In turn, CCL17 binds to CCR8 on cDCs (autocrine) and on CD4+ T cells (paracrine) to stimulate an upregulation of CCL3 expression and release. (III.) Subsequently, CCL3 interacts with CCR1 on naïve T cells, thereby blocking the differentiation and expansion of Tregs.
Supplementary information
Supplementary Information
Supplementary Tables 1–11.
Supplementary Table 12
Twelve major resources table.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Table 1
Statistical source data.
Source Data Extended Data Table 2
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Döring, Y., van der Vorst, E.P.C., Yan, Y. et al. Identification of a non-canonical chemokine-receptor pathway suppressing regulatory T cells to drive atherosclerosis. Nat Cardiovasc Res 3, 221–242 (2024). https://doi.org/10.1038/s44161-023-00413-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-023-00413-9