Abstract
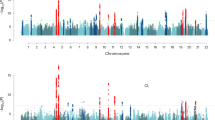
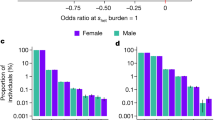
Age at first sexual intercourse and age at first birth have implications for health and evolutionary fitness. In this genome-wide association study (age at first sexual intercourse, N = 387,338; age at first birth, N = 542,901), we identify 371 single-nucleotide polymorphisms, 11 sex-specific, with a 5–6% polygenic score prediction. Heritability of age at first birth shifted from 9% [CI = 4–14%] for women born in 1940 to 22% [CI = 19–25%] for those born in 1965. Signals are driven by the genetics of reproductive biology and externalising behaviour, with key genes related to follicle stimulating hormone (FSHB), implantation (ESR1), infertility and spermatid differentiation. Our findings suggest that polycystic ovarian syndrome may lead to later age at first birth, linking with infertility. Late age at first birth is associated with parental longevity and reduced incidence of type 2 diabetes and cardiovascular disease. Higher childhood socioeconomic circumstances and those in the highest polygenic score decile (90%+) experience markedly later reproductive onset. Results are relevant for improving teenage and late-life health, understanding longevity and guiding experimentation into mechanisms of infertility.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Our policy is to make genome-wide summary statistics widely and publically available. Upon publication, summary statistics will be available on the GWAS catalogue website: https://www.ebi.ac.uk/gwas/downloads/summary-statistics.
The phenotype and genotype data for separate studies used in this GWAS are available upon application to each of the participating cohorts, who can be contacted directly to follow their different data access policies. Access to the UK Biobank is available through application with information available at: http://www.ukbiobank.ac.uk.
Code availability
No custom code was used, with all analyses and modelling using standard software as described in the Methods section and in detail in the Supplementary Information.
Change history
28 July 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41562-021-01179-5
References
Mercer, C. H. et al. Changes in sexual attitudes and lifestyles in Britain through the life course and over time: findings from the National Surveys of Sexual Attitudes and Lifestyles (Natsal). Lancet 382, 1781–1794 (2013).
Lara, L. A. S. & Abdo, C. H. N. Age at time of initial sexual intercourse and health of adolescent girls. J. Pediatr. Adolesc. Gynecol. 29, 417–423 (2016).
Polimanti, R. et al. The interplay between risky sexual behaviors and alcohol dependence: genome-wide association and neuroimaging support for LHPP as a risk gene. Neuropsychopharmacology 42, 598–605 (2017).
Karlsson Linnér, R. et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat. Genet. 51, 245–257 (2019).
Balbo, N., Billari, F. C. & Mills, M. Fertility in advanced societies: a review of research. Eur. J. Popul./Rev. Eur. Démographie 29, 1–38 (2013).
Mills, M. C. et al. Why do people postpone parenthood? Reasons and social policy incentives. Hum. Reprod. Update 17, 848–860 (2011).
Rahmioglu, N. et al. Genome-wide enrichment analysis between endometriosis and obesity-related traits reveals novel susceptibility loci. Hum. Mol. Genet. 24, 1185–1199 (2015).
Barban, N. et al. Genome-wide analysis identifies 12 loci influencing human reproductive behavior. Nat. Genet. 48, 1462–1472 (2016).
Martin, N. G., Eaves, L. J. & Eysenck, H. J. Genetical, environmental and personality factors influencing the age of first sexual intercourse in twins. J. Biosoc. Sci. 9, 91–97 (1977).
Day, F. R. et al. Physical and neurobehavioral determinants of reproductive onset and success. Nat. Genet. https://doi.org/10.1038/ng.3551 (2016).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Tropf, F. C. et al. Hidden heritability due to heterogeneity across seven populations. Nat. Hum. Behav. 1, 757–765 (2017).
Mills, M. C. Introducing Survival and Event History Analysis (Sage, 2011).
Singh, S., Darroch, J. E. & Frost, J. J. Socioeconomic disadvantage and adolescent women’s sexual and reproductive behavior: the case of five developed countries. Fam. Plann. Perspect. 33, 251 (2001).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Grotzinger, A. D. et al. Genomic structural equation modelling provides insights into the multivariate genetic architecture of complex traits. Nat. Hum. Behav. 3, 513–525 (2019).
Davey Smith, G. What can Mendelian randomisation tell us about modifiable behavioural and environmental exposures? BMJ https://doi.org/10.1136/bmj.330.7499.1076 (2005).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Zheng, J.-S. et al. The association between circulating 25-hydroxyvitamin D metabolites and type 2 diabetes in European populations: a meta-analysis and Mendelian randomisation analysis. PLoS Med. 17, e1003394 (2020).
Nelson, C. P. et al. Association analyses based on false discovery rate implicate new loci for coronary artery disease. Nat. Genet. 49, 1385–1391 (2017).
Lind, J. M., Hennessy, A. & Chiu, C. L. Association between a woman’s age at first birth and high blood pressure. Medicine (Baltimore) 94, e697 (2015).
Patchen, L., Leoutsakos, J.-M. & Astone, N. M. Early parturition: is young maternal age at first birth associated with obesity? J. Pediatr. Adolesc. Gynecol. 30, 553–559 (2017).
Kim, J. H., Jung, Y., Kim, S. Y. & Bae, H. Y. Impact of age at first childbirth on glucose tolerance status in postmenopausal women: the 2008–2011 Korean national health and nutrition examination survey. Diabetes Care 37, 671–677 (2014).
Day, F. et al. Large-scale genome-wide meta-analysis of polycystic ovary syndrome suggests shared genetic architecture for different diagnosis criteria. PLoS Genet. 14, e1007813 (2018).
Eisenberg, D. T. A., Hayes, M. G. & Kuzawa, C. W. Delayed paternal age of reproduction in humans is associated with longer telomeres across two generations of descendants. Proc. Natl Acad. Sci. USA 109, 10251–10256 (2012).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014.e22 (2018).
Tabula Muris Consortium et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Yang, H., Robinson, P. N. & Wang, K. Phenolyzer: phenotype-based prioritization of candidate genes for human diseases. Nat. Methods 12, 841–843 (2015).
Vaez, A. et al. In silico post genome-wide association studies analysis of c-reactive protein loci suggests an important role for interferons. Circ. Cardiovasc. Genet. 8, 487–497 (2015).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
van der Wijst, M. G. P. et al. Single-cell RNA sequencing identifies celltype-specific cis-eQTLs and co-expression QTLs. Nat. Genet. 50, 493–497 (2018).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Ellsworth, B. S. et al. FOXL2 in the pituitary: molecular, genetic, and developmental analysis. Mol. Endocrinol 20, 2796–2805 (2006).
van Vliet, J. et al. Human KLF17 is a new member of the Sp/KLF family of transcription factors. Genomics 87, 474–482 (2006).
Governini, L. et al. FOXL2 in human endometrium: hyperexpressed in endometriosis. Reprod. Sci. 21, 1249–1255 (2014).
Rico, C. et al. HIF1 activity in granulosa cells is required for FSH-regulated Vegfa expression and follicle survival in mice. Biol. Reprod. 90, 135 (2014).
Dai, Z. et al. Caveolin-1 promotes trophoblast cell invasion through the focal adhesion kinase (FAK) signalling pathway during early human placental development. Reprod. Fertil. Dev. https://doi.org/10.1071/RD18296 (2019).
Artini, P. G. et al. Cumulus cells surrounding oocytes with high developmental competence exhibit down-regulation of phosphoinositol 1,3 kinase/protein kinase B (PI3K/AKT) signalling genes involved in proliferation and survival. Hum. Reprod. 32, 2474–2484 (2017).
Zheng, J. et al. Novel FSHβ mutation in a male patient with isolated FSH deficiency and infertility. Eur. J. Med. Genet. 60, 335–339 (2017).
Yan, W., Burns, K. H., Ma, L. & Matzuk, M. M. Identification of Zfp393, a germ cell-specific gene encoding a novel zinc finger protein. Mech. Dev. 118, 233–239 (2002).
Lin, Y.-N., Roy, A., Yan, W., Burns, K. H. & Matzuk, M. M. Loss of zona pellucida binding proteins in the acrosomal matrix disrupts acrosome biogenesis and sperm morphogenesis. Mol. Cell. Biol. 27, 6794–6805 (2007).
Wieser, F. et al. Expression and regulation of CCR1 in peritoneal macrophages from women with and without endometriosis. Fertil. Steril. 83, 1878–1881 (2005).
Mei, J. et al. CXCL16/CXCR6 interaction promotes endometrial decidualization via the PI3K/ AKT pathway. Reproduction https://doi.org/10.1530/REP-18-0417 (2019).
Gusev, F. E. et al. Epigenetic-genetic chromatin footprinting identifies novel and subject-specific genes active in prefrontal cortex neurons. FASEB J. 33, 8161–8173 (2019).
Quinn, J. P., Savage, A. L. & Bubb, V. J. Non-coding genetic variation shaping mental health. Curr. Opin. Psychol. 27, 18–24 (2019).
Barak, B. et al. Neuronal deletion of Gtf2i, associated with Williams syndrome, causes behavioral and myelin alterations rescuable by a remyelinating drug. Nat. Neurosci. 22, 700–708 (2019).
Li, Y. et al. Topoisomerase IIbeta is required for proper retinal development and survival of postmitotic cells. Biol. Open 3, 172–184 (2014).
Athanasiou, M. C. et al. The transcription factor E2F-1 in SV40 T antigen-induced cerebellar Purkinje cell degeneration. Mol. Cell. Neurosci. 12, 16–28 (1998).
Yang, X. et al. The association between NCAM1 levels and behavioral phenotypes in children with autism spectrum disorder. Behav. Brain Res. 359, 234–238 (2019).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Tu, S. et al. NitroSynapsin therapy for a mouse MEF2C haploinsufficiency model of human autism. Nat. Commun. 8, 1488 (2017).
Shamir, A. et al. The importance of the NRG-1/ErbB4 pathway for synaptic plasticity and behaviors associated with psychiatric disorders. J. Neurosci. 32, 2988–2997 (2012).
Yang, J.-M. et al. erbb4 deficits in chandelier cells of the medial prefrontal cortex confer cognitive dysfunctions: implications for schizophrenia. Cereb. Cortex 29, 4334–4346 (2019).
Day, F. R. et al. Causal mechanisms and balancing selection inferred from genetic associations with polycystic ovary syndrome. Nat. Commun. 6, 8464 (2015).
Baumgartner, H. K. et al. Characterization of choline transporters in the human placenta over gestation. Placenta 36, 1362–1369 (2015).
Peng, Z. et al. Liver X receptor β in the hippocampus: a potential novel target for the treatment of major depressive disorder? Neuropharmacology 135, 514–528 (2018).
Mahajan, A. et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat. Genet. 50, 1505–1513 (2018).
Tropf, F. C. et al. Human fertility, molecular genetics, and natural selection in modern societies. PLoS One 10, e0126821 (2015).
Waren, E. B. & et al. Heterogeneity in polygenic scores for common human traits. Preprint at bioRxiv https://doi.org/10.1101/106062 (2017).
Ripke, S. et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Mills, M. C. & Rahal, C. The GWAS diversity monitor tracks diversity by disease in real time. Nat. Genet. 52, 242–243 (2020).
Chen, X.-K. et al. Teenage pregnancy and adverse birth outcomes: a large population based retrospective cohort study. Int. J. Epidemiol. 36, 368–373 (2007).
Bongaarts, J., Mensch, B. S. & Blanc, A. K. Trends in the age at reproductive transitions in the developing world: the role of education. Popul. Stud. (N. Y.). 71, 139–154 (2017).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: Fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Finucane, H. K. et al. Partionining heritability by functional category using GWAS summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Altman, D. G. & Bland, J. M. Interaction revisited: the difference between two estimates. Br. Med. J. 326, 219 (2003).
Turley, P. et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat. Genet. 50, 229–237 (2018).
Harris, K. M. & et al. The National Longitudinal Study of Adolescent to Adult Health: research design. Carolina Population Center http://www.cpc.unc.edu/projects/addhealth/design (2009).
Buck, N. & McFall, S. Understanding society: design overview. Longit. Life Course Stud. 3, 5–17 (2012).
Euesden, J., Lewis, C. M. & O’Reilly, P. F. PRSice: polygenic risk score software. Bioinformatics 31, btu848–btu1468 (2014).
Vilhjálmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. https://doi.org/10.1038/s41588-018-0147-3 (2018).
Burgess, S., Butterworth, A. & Thompson, S. G. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet. Epidemiol. 37, 658–665 (2013).
Bowden, J., Smith, G. D. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. https://doi.org/10.1093/ije/dyv080 (2015).
Hemani, G., Bowden, J. & Davey Smith, G. Evaluating the potential role of pleiotropy in Mendelian randomization studies. Hum. Mol. Genet. 27, R195–R208 (2018).
Burgess, S. & Thompson, S. G. Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects. Am. J. Epidemiol. 181, 251–260 (2015).
Day, F. R., Loh, P.-R., Scott, R. A., Ong, K. K. & Perry, J. R. B. A robust example of collider bias in a genetic association study. Am. J. Hum. Genet. 98, 392–393 (2016).
Woods, L. M. Geographical variation in life expectancy at birth in England and Wales is largely explained by deprivation. J. Epidemiol. Community Health 59, 115–120 (2005).
Timshel, P. N., Thompson, J. J. & Pers, T. H. Genetic mapping of etiologic brain cell types for obesity. eLife 9, e55851 (2020).
Vosa, U. & Al., E. Unraveling the polygenic architecture of complex traits using blood eQTL metaanalysis. Preprint at bioRxiv https://doi.org/10.1101/447367 (2018).
Qi, T. et al. Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood. Nat. Commun. 9, 2282 (2018).
Bult, C. J. et al. Mouse genome database (MGD) 2019. Nucleic Acids Res. 47, D801–D806 (2019).
Acknowledgements
A detailed list of funding and other acknowledgements for each cohort can be found in Supplementary Sect. 14. This research was conducted using the UK Biobank resource under application 22276 and 9905. Funding was provided to M.C.M. by the ERC, SOCIOGENOME (615603), CHRONO (835079), ESRC/UKRI SOCGEN (ES/N011856/1), Wellcome Trust ISSF, Leverhulme Trust and Leverhulme Centre for Demographic Science, to N.B. by ERC GENPOP (865356), to F.C.T. by LabEx Ecode, French National Research Agency (ANR) Investissements d’Avenir (ANR-11-LABX-0047), to M.d.H. by Swedish Heart-Lung Foundation (20170872, 20200781, 20140543, 20170678, 20180706 and 20200602), Kjell and Märta Beijer Foundation and Swedish Research Council (2015-03657, 2019-01417). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. This study received ethical approval from the Department of Sociology, University of Oxford, and relevant ethical approval was obtained at the local level for the contributing datasets. The authors thank E. T. Akimova and S. Møllegaard for administrative work in the organization of the cohort information and author list.
Author information
Authors and Affiliations
Consortia
Contributions
M.C.M. and F.R.D. designed and led the study. M.C.M. wrote the paper and Supplementary Information with contributions by authors for respective analyses and comments by all authors. D.M.B. conducted phenotypic changes, phenotype preparation, LD score and genetic correlations, GenomicSEM and EFA and sex-specific effects. N.B. conducted GWAS meta-analysis, MTAG, PGS prediction, survival models and Cox models of longevity. F.C.T. and F.R.D. conducted the cohort QC. F.C.T. conducted GREML cohort heritability analysis and phenotype preparation in UKBB. F.R.D. ran MR and conducted GWAS analyses, J.R.B.P. conducted COJO and X chromosome analysis and K.K.O. provided comments and expertise throughout. N.v.Z. conducted DEPICT and Phenolyzer analyses. A.V. and H.S. conducted in silico sequencing and SMR analyses. T.H.P. conducted cell type enrichment analyses. M.d.H. integrated gene prioritization results and performed downstream analyses, for example, Human Protein Atlas; Entrez, GeneCards and Uniprot mining; and STRING protein–protein interaction analyses. Authors in the Human Reproductive Behaviour Consortium contributed valuable data, conducted cohort-specific GWAS and other analyses, and contributed through the administration, management and data collection for the participating cohorts. The eQTLGen and BIOS Consortia provided data for additional analyses. All authors reviewed and approved the final version of the paper, and code relies upon the standard packages described above.
Corresponding authors
Ethics declarations
Competing interests
The main authors declare no competing interests. The views expressed in this article are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health. M.I.M. has served on advisory panels for Pfizer, NovoNordisk and Zoe Global, has received honoraria from Merck, Pfizer, Novo Nordisk and Eli Lilly and research funding from Abbvie, Astra Zeneca, Boehringer Ingelheim, Eli Lilly, Janssen, Merck, NovoNordisk, Pfizer, Roche, Sanofi Aventis, Servier and Takeda. As of June 2019, M.I.M. is an employee of Genentech and a holder of Roche stock.
Additional information
Peer review information Nature Human Behaviour thanks Ahmed Elhakeem and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information
Supplementary Figs. 1–19, Supplementary Discussion, Supplementary Tables 3a (excerpt) and 14, Supplementary Authorship and Detailed Acknowledgements.
Supplementary tables
Supplementary Tables 1–19.
Rights and permissions
About this article
Cite this article
Mills, M.C., Tropf, F.C., Brazel, D.M. et al. Identification of 371 genetic variants for age at first sex and birth linked to externalising behaviour. Nat Hum Behav 5, 1717–1730 (2021). https://doi.org/10.1038/s41562-021-01135-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41562-021-01135-3
This article is cited by
-
The causal effect of reproductive factors on pelvic floor dysfunction: a Mendelian randomization study
BMC Women's Health (2024)
-
Identifying causal associations between women’s reproductive traits and risk of schizophrenia: a multivariate validated two-sample Mendelian randomization analysis
BMC Psychiatry (2024)
-
Mortality risk predicts global, local, and individual patterns of human reproduction
BMC Public Health (2024)
-
Causal effect of the age at first birth with depression: a mendelian randomization study
BMC Medical Genomics (2024)
-
Shared genetics of ADHD, cannabis use disorder and cannabis use and prediction of cannabis use disorder in ADHD
Nature Mental Health (2024)