Abstract
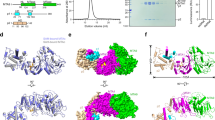
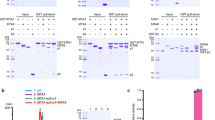
SET domain protein methyltransferases catalyze the transfer of methyl groups from the cofactor S-adenosylmethionine (AdoMet) to specific lysine residues of protein substrates, such as the N-terminal tails of histones H3 and H4 and the large subunit of the Rubisco holoenzyme complex. The crystal structures of pea Rubisco large subunit methyltransferase (LSMT) in ternary complexes with either lysine or ε-N-methyllysine (MeLys) and the product S-adenosylhomocysteine (AdoHcy) were determined to resolutions of 2.65 and 2.55 Å, respectively. The ζ-methyl group of MeLys is bound to the enzyme via carbon–oxygen hydrogen bonds that play a key role in catalysis. The methyl donor and acceptor are aligned in a linear geometry for SN2 nucleophilic transfer of the methyl group during catalysis. Differences in hydrogen bonding between the MeLys ε-amino group and Rubisco LSMT and SET7/9 explain why Rubisco LSMT generates multiply methylated Lys, wheras SET7/9 generates only MeLys.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Kouzarides, T. Histone methylation in transcriptional control. Curr. Opin. Genet. Dev. 12, 198–209 (2002).
Lachner, M. & Jenuwein, T. The many faces of histone lysine methylation. Curr. Opin. Cell Biol. 14, 286–298 (2002).
Rice, J.C. & Allis, C.D. Histone methylation versus histone acetylation: new insights into epigenetic regulation. Curr. Opin. Cell Biol. 13, 263–273 (2001).
Strahl, B.D., Ohba, R., Cook, R.G. & Allis, C.D. Methylation of histone H3 at lysine 4 is highly conserved and correlates with transcriptionally active nuclei in Tetrahymena. Proc. Natl. Acad. Sci. USA 96, 14967–14972 (1999).
Noma, K., Allis, C.D. & Grewal, S.I. Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science 293, 1150–1155 (2001).
Houtz, R.L., Stults, J.T., Mulligan, R.M. & Tolbert, N.E. Post-translational modifications in the large subunit of ribulose bisphosphate carboxylase/oxygenase. Proc. Natl. Acad. Sci. USA 86, 1855–1859 (1989).
Klein, R.R. & Houtz, R.L. Cloning and developmental expression of pea ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit N-methyltransferase. Plant Mol. Biol. 27, 249–261 (1995).
Schultz, J., Milpetz, F., Bork, P. & Ponting, C.P. SMART, a simple modular architecture research tool: identification of signaling domains. Proc. Natl. Acad. Sci. USA 95, 5857–5864 (1998).
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000).
Lachner, M., O'Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001).
Bannister, A.J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001).
Nakayama, J., Rice, J.C., Strahl, B.D., Allis, C.D. & Grewal, S.I. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292, 110–113 (2001).
Bernstein, B.E. et al. Methylation of histone H3 Lys 4 in coding regions of active genes. Proc. Natl. Acad. Sci. USA 99, 8695–8700 (2002).
Santos-Rosa, H. et al. Active genes are tri-methylated at K4 of histone H3. Nature 419, 407–411 (2002).
Zhang, X. et al. Structure of the Neurospora SET domain protein DIM-5, a histone H3 lysine methyltransferase. Cell 111, 117–127 (2002).
Jacobs, S.A. et al. The active site of the SET domain is constructed on a knot. Nat. Struct. Biol. 9, 833–838 (2002).
Kwon, T. et al. Mechanism of histone lysine methyl transfer revealed by the structure of SET7/9-AdoMet. EMBO J. 22, 292–303 (2003).
Wilson, J.R. et al. Crystal structure and functional analysis of the histone methyltransferase SET7/9. Cell 111, 105–115 (2002).
Xiao, B. et al. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature 421, 652–656 (2003).
Min, J., Zhang, X., Cheng, X., Grewal, S.I. & Xu, R.M. Structure of the SET domain histone lysine methyltransferase Clr4. Nat. Struct. Biol. 9, 828–832 (2002).
Manzur, K.L. et al. A dimeric viral SET domain methyltransferase specific to Lys27 of histone H3. Nat. Struct. Biol. 10, 187–196 (2003).
Trievel, R.C., Beach, B.M., Dirk, L.M., Houtz, R.L. & Hurley, J.H. Structure and catalytic mechanism of a SET domain protein methyltransferase. Cell 111, 91–103 (2002).
Marmorstein, R. Structure of SET domain proteins: a new twist on histone methylation. Trends Biochem. Sci. 28, 59–62 (2003).
Yeates, T.O. Structures of SET domain proteins: protein lysine methyltransferases make their mark. Cell 111, 5–7 (2002).
Dutnall, R.N. & Denu, J.M. Methyl magic and HAT tricks. Nat. Struct. Biol. 9, 888–891 (2002).
Houtz, R.L., Poneleit, L., Jones, S.B., Royer, M. & Stults, J.T. Posttranslational modifications in the amino-terminal region of the large subunit of ribulose-1,5-bisphosphate carboxylase/oxygenase from several plant species. Plant Physiol. 98, 1170–1174 (1992).
Houtz, R.L., Royer, M. & Salvucci, M.E. Partial purification and characterization of ribulose 1,5-bisphosphate carboxylase/oxygenase large subunit N-methyltransferase. Plant Physiol. 97, 913–920 (1991).
Segel, I.H. Enzyme Kinetics (John Wiley & Sons, New York; 1975).
Derewenda, Z.S., Lee, L. & Derewenda, U. The occurrence of C-H...O hydrogen bonds in proteins. J. Mol. Biol. 252, 248–262 (1995).
Bella, J. & Berman, H.M. Crystallographic evidence for Cα-H...O=C hydrogen bonds in a collagen triple helix. J. Mol. Biol. 264, 734–742 (1996).
Fabiola, G.F., Krishnaswamy, S., Nagarajan, V. & Pattabhi, V. C-H...O hydrogen bonds in β-sheets. Acta Crystallogr. D 53, 316–320 (1997).
Wahl, M.C. & Sundaralingam, M. C-H...O hydrogen bonding in biology. Trends Biochem. Sci. 22, 97–101 (1997).
Senes, A., Ubarretxena-Belandia, I. & Engelman, D.M. The Cα-H...O hydrogen bond: a determinant of stability and specificity in transmembrane helix interactions. Pro. Natl. Acad. Sci. USA 98, 9056–9061 (2001).
Weiss, M.S., Brandl, M., Suhnel, J., Pal, D. & Hilgenfeld, R. More hydrogen bonds for the (structural) biologist. Trends Biochem. Sci. 26, 521–523 (2001).
Scheiner, S., Kar, T. & Gu, Y. Strength of the Cα-H...O hydrogen bond of amino acid residues. J. Biol. Chem. 276, 9832–9837 (2001).
Derewenda, Z.S., Derewenda, U. & Kobos, P.M. (His)Cε-H...O=C < hydrogen bond in the active sites of serine hydrolases. J. Mol. Biol. 241, 83–93 (1994).
Bach, R.D., Thorpe, C., & Dmitrenko, O. C-H...carboxylate oxygen hydrogen bonding in substrate activation by acyl-CoA dehydrogenases: synergy between the H-bonds. J. Phys. Chem. 106, 4325–4335 (2002).
Coward, J.K. Chemical mechanisms of methyl transfer reactions: comparison of methylases with nonenyzmatic 'model reactions' in The Biochemistry of Adenosylmethionine (eds. Salvatore, F., Borek, E., Zappia, V., William-Ashman, H.G. & Schlenk, F.) 127–144 (Columbia University Press, New York; 1977).
Duff, A.P., Andrews, T.J. & Curmi, P.M. The transition between the open and closed states of rubisco is triggered by the inter-phosphate distance of the bound bisphosphate. J. Mol. Biol. 298, 903–916 (2000).
Rebouche, C.J. & Broquist, H.P. Carnitine biosynthesis in Neurospora crassa: enzymatic conversion of lysine to ε-N-trimethyllysine. J. Bacteriol. 126, 1207–1214 (1976).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. Model. 15, 132–134 (1997).
Esnouf, R.M. Further additions to MolScript version 1.4, including reading and contouring of electron-density maps. Acta Crystallogr. D 55, 938–940 (1999).
Merritt, E.A. & Bacon, D.J. Raster3D version 2.0: a program for photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Guex, N. & Peitsch, M.C. SWISS-MODEL and the Swiss-Pdb Viewer: an environment for comparative protein modeling. Electrophoresis 18, 2714–2723 (1997).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 24, 946–950 (1993).
Acknowledgements
We would like to thank S. Gamblin for SET7/9–AdoHcy–histone H3 ternary complex coordinates, and D. Engelman and G. Felsenfeld for insightful discussions. Research by R.L.H. and E.M.F. was supported by the DOE, Division of Energy Biosciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Trievel, R., Flynn, E., Houtz, R. et al. Mechanism of multiple lysine methylation by the SET domain enzyme Rubisco LSMT. Nat Struct Mol Biol 10, 545–552 (2003). https://doi.org/10.1038/nsb946
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb946