Abstract
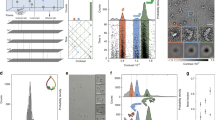
Native mass spectrometry is an emerging technology that allows the topological investigation of intact protein complexes with high sensitivity and a theoretically unrestricted mass range. This unique tool provides complementary information to established technologies in structural biology, and also provides a link to high-throughput interactomics studies, which do not generate information on exact protein complex-composition, structure or dynamics. Here I review the current state of native mass spectrometry technology and discuss several important biological applications. I also describe current experimental challenges in native mass spectrometry, encouraging readers to contribute to solutions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Alberts, B. The cell as a collection of protein machines: preparing the next generation of molecular biologists. Cell 92, 291–294 (1998).
Stelzl, U. et al. A human protein-protein interaction network: a resource for annotating the proteome. Cell 122, 957–968 (2005).
Kocher, T. & Superti-Furga, G. Mass spectrometry-based functional proteomics: from molecular machines to protein networks. Nat. Methods 4, 807–815 (2007).
Gingras, A.C., Gstaiger, M., Raught, B. & Aebersold, R. Analysis of protein complexes using mass spectrometry. Nat. Rev. Mol. Cell Biol. 8, 645–654 (2007).
Krogan, N.J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006).
Gavin, A.C. et al. Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636 (2006).
Robinson, C.V., Sali, A. & Baumeister, W. The molecular sociology of the cell. Nature 450, 973–982 (2007).
Sprangers, R., Velyvis, A. & Kay, L.E. Solution NMR of supramolecular complexes: providing new insights into function. Nat. Methods 4, 697–703 (2007).
Loo, J.A. Studying noncovalent protein complexes by electrospray ionization mass spectrometry. Mass Spectrom. Rev. 16, 1–23 (1997).
van den Heuvel, R.H. & Heck, A.J. Native protein mass spectrometry: from intact oligomers to functional machineries. Curr. Opin. Chem. Biol. 8, 519–526 (2004).
Kaddis, C.S. & Loo, J.A. Native protein MS and ion mobility large flying proteins with ESI. Anal. Chem. 79, 1778–1784 (2007).
Sharon, M. & Robinson, C.V. The role of mass spectrometry in structure elucidation of dynamic protein complexes. Annu. Rev. Biochem. 76, 167–193 (2007).
Uetrecht, C. et al. High-resolution mass spectrometry of viral assemblies: molecular composition and stability of dimorphic hepatitis B virus capsids. Proc. Natl. Acad. Sci. USA 105, 9216–9220 (2008).
Uetrecht, C. et al. Stability and shape of hepatitis B virus capsids in vacuo. Angew. Chem. Int. Ed. 47, 6247–6251 (2008).
Heck, A.J. & Van Den Heuvel, R.H. Investigation of intact protein complexes by mass spectrometry. Mass Spectrom. Rev. 23, 368–389 (2004).
Benesch, J.L., Ruotolo, B.T., Simmons, D.A. & Robinson, C.V. Protein complexes in the gas phase: technology for structural genomics and proteomics. Chem. Rev. 107, 3544–3567 (2007).
Hernandez, H. & Robinson, C.V. Determining the stoichiometry and interactions of macromolecular assemblies from mass spectrometry. Nat. Protoc. 2, 715–726 (2007).
van Duijn, E. et al. Tandem mass spectrometry of intact GroEL-substrate complexes reveals substrate-specific conformational changes in the trans ring. J. Am. Chem. Soc. 128, 4694–4702 (2006).
van Duijn, E. et al. Monitoring macromolecular complexes involved in the chaperonin-assisted protein folding cycle by mass spectrometry. Nat. Methods 2, 371–376 (2005).
Vaughan, C.K. et al. Structure of an Hsp90-Cdc37-Cdk4 complex. Mol. Cell 23, 697–707 (2006).
Sharon, M., Taverner, T., Ambroggio, X.I., Deshaies, R.J. & Robinson, C.V. Structural organization of the 19S proteasome lid: insights from MS of intact complexes. PLoS Biol. 4, e267 (2006).
Sharon, M. et al. 20S proteasomes have the potential to keep substrates in store for continual degradation. J. Biol. Chem. 281, 9569–9575 (2006).
Sharon, M., Witt, S., Glasmacher, E., Baumeister, W. & Robinson, C.V. Mass spectrometry reveals the missing links in the assembly pathway of the bacterial 20 S proteasome. J. Biol. Chem. 282, 18448–18457 (2007).
Synowsky, S.A., van den Heuvel, R.H., Mohammed, S., Pijnappel, P.W. & Heck, A.J. Probing genuine strong interactions and post-translational modifications in the heterogeneous yeast exosome protein complex. Mol. Cell. Proteomics 5, 1581–1592 (2006).
Hernandez, H., Dziembowski, A., Taverner, T., Seraphin, B. & Robinson, C.V. Subunit architecture of multimeric complexes isolated directly from cells. EMBO Rep. 7, 605–610 (2006).
Taverner, T. et al. Subunit architecture of intact protein complexes from mass spectrometry and homology modeling. Acc. Chem. Res. 41, 617–627 (2008).
Lorenzen, K., Vannini, A., Cramer, P. & Heck, A.J. Structural biology of RNA polymerase III: mass spectrometry elucidates subcomplex architecture. Structure 15, 1237–1245 (2007).
Sprangers, R. & Kay, L.E. Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature 445, 618–622 (2007).
Medalia, O. et al. Macromolecular architecture in eukaryotic cells visualized by cryoelectron tomography. Science 298, 1209–1213 (2002).
Loo, J.A. et al. Electrospray ionization mass spectrometry and ion mobility analysis of the 20S proteasome complex. J. Am. Soc. Mass Spectrom. 16, 998–1008 (2005).
Cramer, P., Bushnell, D.A. & Kornberg, R.D. Structural basis of transcription: RNA polymerase II at 2.8 angstrom resolution. Science 292, 1863–1876 (2001).
Fernandez-Tornero, C. et al. Insights into transcription initiation and termination from the electron microscopy structure of yeast RNA polymerase III. Mol. Cell 25, 813–823 (2007).
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 17, 1030–1032 (1999).
Synowsky, S.A. & Heck, A.J. The yeast Ski complex is a hetero-tetramer. Protein Sci. 17, 119–125 (2007).
Benesch, J.L., Aquilina, J.A., Ruotolo, B.T., Sobott, F. & Robinson, C.V. Tandem mass spectrometry reveals the quaternary organization of macromolecular assemblies. Chem. Biol. 13, 597–605 (2006).
Lorenzen, K., Olia, A.S., Uetrecht, C., Cingolani, G. & Heck, A.J. Determination of stoichiometry and conformational changes in the first step of the P22 tail assembly. J. Mol. Biol. 379, 385–396 (2008).
Demmers, J.A., Haverkamp, J., Heck, A.J., Koeppe, R.E. II & Killian, J.A. Electrospray ionization mass spectrometry as a tool to analyze hydrogen/deuterium exchange kinetics of transmembrane peptides in lipid bilayers. Proc. Natl. Acad. Sci. USA 97, 3189–3194 (2000).
Doyle, D.A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998).
Sharon, M., Ilag, L.L. & Robinson, C.V. Evidence for micellar structure in the gas phase. J. Am. Chem. Soc. 129, 8740–8746 (2007).
Barrera, N.P., Di Bartolo, N., Booth, P.J. & Robinson, C.V. Micelles protect membrane complexes from solution to vacuum. Science 321, 243–246 (2008).
Rinner, O. et al. Identification of cross-linked peptides from large sequence databases. Nat. Methods 5, 315–318 (2008).
Sinz, A. Chemical cross-linking and mass spectrometry to map three-dimensional protein structures and protein-protein interactions. Mass Spectrom. Rev. 25, 663–682 (2006).
Aloy, P. et al. Structure-based assembly of protein complexes in yeast. Science 303, 2026–2029 (2004).
Eckers, C., Laures, A.M., Giles, K., Major, H. & Pringle, S. Evaluating the utility of ion mobility separation in combination with high-pressure liquid chromatography/mass spectrometry to facilitate detection of trace impurities in formulated drug products. Rapid Commun. Mass Spectrom. 21, 1255–1263 (2007).
Ruotolo, B.T. et al. Evidence for macromolecular protein rings in the absence of bulk water. Science 310, 1658–1661 (2005).
Pinkse, M.W., Maier, C.S., Kim, J.I., Oh, B.H. & Heck, A.J. Macromolecular assembly of Helicobacter pylori urease investigated by mass spectrometry. J. Mass Spectrom. 38, 315–320 (2003).
Sobott, F., Hernandez, H., McCammon, M.G., Tito, M.A. & Robinson, C.V. A tandem mass spectrometer for improved transmission and analysis of large macromolecular assemblies. Anal. Chem. 74, 1402–1407 (2002).
van den Heuvel, R.H. et al. Improving the performance of a quadrupole time-of-flight instrument for macromolecular mass spectrometry. Anal. Chem. 78, 7473–7483 (2006).
Shvartsburg, A.A., Mashkevich, S.V., Baker, E.S. & Smith, R.D. Optimization of algorithms for ion mobility calculations. J. Phys. Chem. 111, 2002–2010 (2007).
Ruotolo, B.T. et al. Ion mobility-mass spectrometry reveals long-lived, unfolded intermediates in the dissociation of protein complexes. Angew. Chem. Int. Ed. 46, 8001–8004 (2007).
Ruotolo, B.T., Benesch, J.L., Sandercock, A.M., Hyung, S.J. & Robinson, C.V. Ion mobility-mass spectrometry analysis of large protein complexes. Nat. Protoc. 3, 1139–1152 (2008).
Acknowledgements
I thank all colleagues in native MS who have contributed in making this area of research come to maturity. I acknowledge all members of my group, most notably the native MS (former) members whose material and input I have used here; K. Lorenzen, H. Mazon, M. Pinkse, S. Synowsky, C. Uetrecht, R. van den Heuvel, E. van Duijn and C. Versluis. K. Lorenzen and C. Uetrecht helped construct the figures. Additionally, I thank all collaborators whose samples, support and expertise have been instrumental in our work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Heck, A. Native mass spectrometry: a bridge between interactomics and structural biology. Nat Methods 5, 927–933 (2008). https://doi.org/10.1038/nmeth.1265
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1265
This article is cited by
-
Direct determination of oligomeric organization of integral membrane proteins and lipids from intact customizable bilayer
Nature Methods (2023)
-
Determining the gas-phase structures of α-helical peptides from shape, microsolvation, and intramolecular distance data
Nature Communications (2023)
-
Structure and dynamics of endogenous cardiac troponin complex in human heart tissue captured by native nanoproteomics
Nature Communications (2023)
-
Rehydration Post-orientation: Investigating Field-Induced Structural Changes via Computational Rehydration
The Protein Journal (2023)
-
Native mass spectrometry-based metabolomics identifies metal-binding compounds
Nature Chemistry (2022)