Abstract
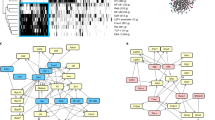
CD8+ T cells in chronic viral infections such as HIV develop functional defects including loss of interleukin-2 (IL-2) secretion and decreased proliferative potential that are collectively termed 'exhaustion'1. Exhausted T cells express increased amounts of multiple inhibitory receptors, such as programmed death-1 (PD-1)2,3, that contribute to impaired virus-specific T cell function. Although reversing PD-1 inhibition is therefore an attractive therapeutic strategy, the cellular mechanisms by which PD-1 ligation results in T cell inhibition are not fully understood. PD-1 is thought to limit T cell activation by attenuating T cell receptor (TCR) signaling4,5. It is not known whether PD-1 also acts by upregulating genes in exhausted T cells that impair their function. Here we analyzed gene expression profiles from HIV-specific CD8+ T cells in individuals with HIV and show that PD-1 coordinately upregulates a program of genes in exhausted CD8+ T cells from humans and mice. This program includes upregulation of basic leucine transcription factor, ATF-like (BATF), a transcription factor in the AP-1 family. Enforced expression of BATF was sufficient to impair T cell proliferation and cytokine secretion, whereas BATF knockdown reduced PD-1 inhibition. Silencing BATF in T cells from individuals with chronic viremia rescued HIV-specific T cell function. Thus, inhibitory receptors can cause T cell exhaustion by upregulating genes—such as BATF—that inhibit T cell function. Such genes may provide new therapeutic opportunities to improve T cell immunity to HIV.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Zajac, A.J. et al. Viral immune evasion due to persistence of activated T cells without effector function. J. Exp. Med. 188, 2205–2213 (1998).
Barber, D.L. et al. Restoring function in exhausted CD8 T cells during chronic viral infection. Nature 439, 682–687 (2006).
Blackburn, S.D. et al. Coregulation of CD8+ T cell exhaustion by multiple inhibitory receptors during chronic viral infection. Nat. Immunol. 10, 29–37 (2009).
Riley, J.L. PD-1 signaling in primary T cells. Immunol. Rev. 229, 114–125 (2009).
Chemnitz, J.M., Parry, R.V., Nichols, K.E., June, C.H. & Riley, J.L. SHP-1 and SHP-2 associate with immunoreceptor tyrosine-based switch motif of programmed death 1 upon primary human T cell stimulation, but only receptor ligation prevents T cell activation. J. Immunol. 173, 945–954 (2004).
Betts, M.R. et al. HIV nonprogressors preferentially maintain highly functional HIV-specific CD8+ T-cells. Blood 107, 4781–4789 (2006).
Migueles, S.A. et al. HIV-specific CD8+ T cell proliferation is coupled to perforin expression and is maintained in nonprogressors. Nat. Immunol. 3, 1061–1068 (2002).
Pereyra, F. et al. Genetic and immunologic heterogeneity among persons who control HIV infection in the absence of therapy. J. Infect. Dis. 197, 563–571 (2008).
Wherry, E.J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Nilsson, B., Håkansson, P., Johansson, M., Nelander, S. & Fioretos, T. Threshold-free high-power methods for the ontological analysis of genome-wide gene-expression studies. Genome Biol. 8, R74 (2007).
Haining, W.N. & Wherry, E.J. Integrating genomic signatures for immunologic discovery. Immunity 32, 152–161 (2010).
Wherry, E.J., Blattman, J.N., Murali-Krishna, K., van der Most, R. & Ahmed, R. Viral persistence alters CD8 T-cell immunodominance and tissue distribution and results in distinct stages of functional impairment. J. Virol. 77, 4911–4927 (2003).
Chemnitz, J.M. et al. RNA fingerprints provide direct evidence for the inhibitory role of TGFβ and PD-1 on CD4+ T cells in Hodgkin lymphoma. Blood 110, 3226–3233 (2007).
Parry, R.V. et al. CTLA-4 and PD-1 receptors inhibit T-cell activation by distinct mechanisms. Mol. Cell. Biol. 25, 9543–9553 (2005).
Day, C.L. et al. PD-1 expression on HIV-specific T cells is associated with T-cell exhaustion and disease progression. Nature 443, 350–354 (2006).
Trautmann, L. et al. Upregulation of PD-1 expression on HIV-specific CD8+ T cells leads to reversible immune dysfunction. Nat. Med. 12, 1198–1202 (2006).
Echlin, D.R., Tae, H.J., Mitin, N. & Taparowsky, E.J. B-ATF functions as a negative regulator of AP-1 mediated transcription and blocks cellular transformation by Ras and Fos. Oncogene 19, 1752–1763 (2000).
Williams, K.L. et al. Characterization of murine BATF: a negative regulator of activator protein-1 activity in the thymus. Eur. J. Immunol. 31, 1620–1627 (2001).
Haining, W.N. et al. Identification of an evolutionarily conserved transcriptional signature of CD8 memory differentiation that is shared by T and B cells. J. Immunol. 181, 1859–1868 (2008).
Petrovas, C. et al. PD-1 is a regulator of virus-specific CD8+ T cell survival in HIV infection. J. Exp. Med. 203, 2281–2292 (2006).
Lanier, L.L. Up on the tightrope: natural killer cell activation and inhibition. Nat. Immunol. 9, 495–502 (2008).
Clark, G.J., Ju, X., Tate, C. & Hart, D.N. The CD300 family of molecules are evolutionarily significant regulators of leukocyte functions. Trends Immunol. 30, 209–217 (2009).
Foletta, V.C., Segal, D.H. & Cohen, D.R. Transcriptional regulation in the immune system: all roads lead to AP-1. J. Leukoc. Biol. 63, 139–152 (1998).
Schraml, B.U. et al. The AP-1 transcription factor Batf controls TH17 differentiation. Nature 460, 405–409 (2009).
Betz, B.C. et al. Batf coordinates multiple aspects of B and T cell function required for normal antibody responses. J. Exp. Med. 207, 933–942 (2010).
Shin, H. et al. A role for the transcriptional repressor Blimp-1 in CD8+ T cell exhaustion during chronic viral infection. Immunity 31, 309–320 (2009).
Crotty, S., Johnston, R.J. & Schoenberger, S.P. Effectors and memories: Bcl-6 and Blimp-1 in T and B lymphocyte differentiation. Nat. Immunol. 11, 114–120 (2010).
Velu, V. et al. Enhancing SIV-specific immunity in vivo by PD-1 blockade. Nature 458, 206–210 (2009).
Haining, W.N. et al. High-throughput gene expression profiling of memory differentiation in primary human T cells. BMC Immunol. 9, 44 (2008).
Acknowledgements
We would like to thank the subjects for taking part in the study; E. Cutrell, B. Baker, K. Moss, A. Rathod and C. Brume for coordinating sample management; and A. Sharpe, T. Golub, G. Lauer, M. Altfeld and H. Joffe for valuable discussions. This work was supported by US National Institutes of Health grants AI082630, AI56299, HHSN26620050030C and HL092565, the International HIV Controllers Study and the Foundation for the National Institutes of Health through the Grand Challenges in Global Health Initiative.
Author information
Authors and Affiliations
Contributions
M.Q. designed and performed experiments, analyzed data and helped write the paper. F. Pereyra designed the clinical components of the study. B.N. and J.L.J. designed and performed computational experiments. F. Porichis, D.S.K., J.Z. and D.E.K. designed and performed siRNA experiments in samples from subjects with HIV. C.F., Q.E., B.J., K.B., S.I., K.R., I.T., A.P.-T., D.D. and L.F. all performed experiments. G.J.F. designed experiments and developed PD-L1–Ig. J.A., A.C., H.S. and E.J.W. designed and performed mouse experiments and analyzed data. W.N.H., B.E. and B.D.W. conceived of the study and designed the experiments. W.N.H. analyzed data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–6 and Supplementary Methods (PDF 1897 kb)
Rights and permissions
About this article
Cite this article
Quigley, M., Pereyra, F., Nilsson, B. et al. Transcriptional analysis of HIV-specific CD8+ T cells shows that PD-1 inhibits T cell function by upregulating BATF. Nat Med 16, 1147–1151 (2010). https://doi.org/10.1038/nm.2232
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2232
This article is cited by
-
Challenges and new technologies in adoptive cell therapy
Journal of Hematology & Oncology (2023)
-
Reprogramming T cell differentiation and exhaustion in CAR-T cell therapy
Journal of Hematology & Oncology (2023)
-
PD-1 signaling negatively regulates the common cytokine receptor γ chain via MARCH5-mediated ubiquitination and degradation to suppress anti-tumor immunity
Cell Research (2023)
-
Generation of functionally distinct T-cell populations by altering the viscoelasticity of their extracellular matrix
Nature Biomedical Engineering (2023)
-
Single cell transcriptomics shows that malaria promotes unique regulatory responses across multiple immune cell subsets
Nature Communications (2023)