Abstract
Post-acute sequelae of COVID-19 (PASC) is a persistent complication of severe acute respiratory syndrome coronavirus 2 infection that includes symptoms, such as fatigue, cognitive impairment, and respiratory distress. These symptoms severely affect the quality of life of patients after their recovery from COVID-19. In this study, a group of machine learning algorithms analyzed the whole blood RNA-seq data from patients with different PASC levels. The purpose of this analysis was to identify the gene markers associated with PASC and the special expression patterns for different PASC levels. By comparing the quality of life of patients after the acute phase of COVID-19 and before the disease, samples in the dataset were divided into three groups, namely, “Better,” “The Same,” and “Worse.” Each patient was represented by the expression levels of 58,929 genes. The machine learning-based workflow included six feature-ranking algorithms, incremental feature selection (IFS), and four classification algorithms. The feature ranking algorithms were in charge of assessing feature importance, whereas IFS with classification algorithms were used to extract essential genes and to construct efficient classifiers and classification rules. The expression of top genes in the results was associated with the immune response to viral infection, which is supported by the published literature. For example, patients with low CCDC18 expression and high CPED1 expression had good quality of life, whereas those with low CDC16 expression had poor quality of life.
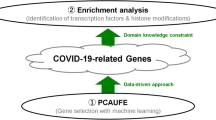
Graphical Abstract







Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.References
World Health Organization. Geneva (Switzerland): World Health Organization; 2020. WHO Director-General's opening remarks at the media briefing on COVID-19 - 11 March 2020 [Internet] [cited 2023 Jan. 26]. Available from: https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020
Nalbandian A et al (2021) Post-acute COVID-19 syndrome. Nat Med 27(4):601–615
Ladds E et al (2020) Persistent symptoms after COVID-19: qualitative study of 114 “long COVID” patients and draft quality principles for services. BMC Health Serv Res 20(1):1144
Greenhalgh T et al (2020) Management of post-acute COVID-19 in primary care. bmj 370:m3026
Huang C et al (2021) 6-month consequences of COVID-19 in patients discharged from hospital: a cohort study. Lancet 397(10270):220–232
Al-Jahdhami I, Al-Naamani K, Al-Mawali A (2021) The post-acute COVID-19 syndrome (long COVID). Oman Med J 36(1):e220
Carfì A, Bernabei R, Landi F (2020) Persistent symptoms in patients after acute COVID-19. JAMA 324(6):603–605
Arnold DT et al (2021) Patient outcomes after hospitalisation with COVID-19 and implications for follow-up: results from a prospective UK cohort. Thorax 76(4):399–401
Knight DR et al (2022) Perception, prevalence, and prediction of severe infection and post-acute sequelae of COVID-19. Am J Med Sci 363(4):295–304
Baj J et al (2020) COVID-19: specific and non-specific clinical manifestations and symptoms: the current state of knowledge. J Clin Med 9(6):1753
Jin X et al (2020) Epidemiological, clinical and virological characteristics of 74 cases of coronavirus-infected disease 2019 (COVID-19) with gastrointestinal symptoms. Gut 69(6):1002–1009
Wong SH, Lui RN, Sung JJ (2020) COVID-19 and the digestive system. J Gastroenterol Hepatol 35(5):744–748
Zhou Z et al (2020) Effect of gastrointestinal symptoms in patients with COVID-19. Gastroenterology 158(8):2294–2297
Guotao L et al (2020) SARS-CoV-2 infection presenting with hematochezia. Med Mal Infect 50(3):293
Munipalli B et al (2022) Post-acute sequelae of COVID-19 (PASC): a meta-narrative review of pathophysiology, prevalence, and management. SN Compr Clin Med 4(1):90
Lieberman NA et al (2020) In vivo antiviral host transcriptional response to SARS-CoV-2 by viral load, sex, and age. PLoS Biol 18(9):e3000849
Townsend L et al (2020) Persistent fatigue following SARS-CoV-2 infection is common and independent of severity of initial infection. PLoS One 15(11):e0240784
Sudre CH et al (2021) Attributes and predictors of long COVID. Nat Med 27(4):626–631
Petersen MS et al (2021) Long COVID in the Faroe Islands: a longitudinal study among nonhospitalized patients. Clin Infect Dis 73(11):e4058–e4063
Patel JA et al (2020) Poverty, inequality and COVID-19: the forgotten vulnerable. Public Health 183:110
McClure ES et al (2020) Racial capitalism within public health—how occupational settings drive COVID-19 disparities. Am J Epidemiol 189(11):1244–1253
Xu R et al. Co‐reactivation of human herpesvirus alpha subfamily (HSV I and VZV) in critically ill patient with COVID‐19. Br J Dermatol 183(6):1145–1147
Hirschtick JL et al (2021) Population-based estimates of post-acute sequelae of SARS-CoV-2 infection (PASC) prevalence and characteristics. Clin Infect Dis 73(11):2055–2064
Chen L et al (2021) Identifying COVID-19-specific transcriptomic biomarkers with machine learning methods. Biomed Res Int 2021:9939134
Huang F et al (2022) Identifying COVID-19 severity-related SARS-CoV-2 mutation using a machine learning method. Life 12(6):806
Chen L et al (2022) Recognition of immune cell markers of COVID-19 severity with machine learning methods. Biomed Res Int 2022:6089242
Lu J et al (2022) Identification of COVID-19 severity biomarkers based on feature selection on single-cell RNA-Seq data of CD8(+) T cells. Front Genet 13:1053772
Chen L et al (2022) Identification of DNA methylation signature and rules for SARS-CoV-2 associated with age. Front Biosci (Landmark Ed) 27(7):204
Liu H, Setiono R (1998) Incremental feature selection. Appl Intell 9(3):217–230
Thompson RC et al (2023) Molecular states during acute COVID-19 reveal distinct etiologies of long-term sequelae. Nat Med 29(1):236–246
Tibshirani R (1996) Regression shrinkage and selection via the LASSO. J Roy Stat Soc: Ser B (Methodol) 58(1):267–288
Ke G et al (2017) LightGBM: a highly efficient gradient boosting decision tree. Adv Neural Inf Process Syst 30:3146–3154
Draminski M et al (2008) Monte Carlo feature selection for supervised classification. Bioinformatics 24(1):110–117
Breiman L (2001) Random forests. Mach Learn 45(1):5–32
Dorogush AV, Ershov V, A Gulin (2018) CatBoost: gradient boosting with categorical features support. arXiv preprint arXiv:1810.11363
Chen T, Guestrin C (2016) XGBoost: a scalable tree boosting system. in The 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. Assoc Comput Mach 785–794
Li H et al (2022) Identifying functions of proteins in mice with functional embedding features. Front Genet 13:909040
Li H et al (2022) Identification of COVID-19-specific immune markers using a machine learning method. Front Mol Biosci 9:952626
Li Z et al (2022) Identifying key microRNA signatures for neurodegenerative diseases with machine learning methods. Front Genet 13:880997
Huang F et al (2023) Analysis and prediction of protein stability based on interaction network, gene ontology, and KEGG pathway enrichment scores. BBA - Proteins Proteomics 1871(3):140889
Huang F et al (2023) Identification of smoking associated transcriptome aberration in blood with machine learning methods. Biomed Res Int 2023:5333361
Ren J et al (2023) Identification of genes associated with the impairment of olfactory and gustatory functions in COVID-19 via machine-learning methods. Life 13(3):798
Zhao X, Chen L, Lu J (2018) A similarity-based method for prediction of drug side effects with heterogeneous information. Math Biosci 306:136–144
Chawla NV et al (2002) SMOTE: synthetic minority over-sampling technique. J Artif Intell Res 16:321–357
Safavian SR, Landgrebe D (1991) A survey of decision tree classifier methodology. IEEE Trans Syst Man Cybern 21(3):660–674
Cover T, Hart P (1967) Nearest neighbor pattern classification. IEEE Trans Inf Theory 13(1):21–27
Cortes C, Vapnik V (1995) Support-vector networks. Mach Learn 20(3):273–297
Powers D (2011) Evaluation: from precision, recall and F-measure to ROC, informedness, markedness & correlation. J Mach Learn Technol 2(1):37–63
Chen L et al (2022) Predicting RNA 5-methylcytosine sites by using essential sequence features and distributions. Biomed Res Int 2022:4035462
Chen L, Chen K, Zhou B (2023) Inferring drug-disease associations by a deep analysis on drug and disease networks. Math Biosci Eng 20(8):14136–14157
Wu C, Chen L (2023) A model with deep analysis on a large drug network for drug classification. Math Biosci Eng 20(1):383–401
Yang Y, Chen L (2022) Identification of drug–disease associations by using multiple drug and disease networks. Curr Bioinform 17(1):48–59
Kohavi R (1995) A study of cross-validation and bootstrap for accuracy estimation and model selection. in International joint Conference on artificial intelligence. Lawrence Erlbaum Associates Ltd
Wang H, Chen L (2023) PMPTCE-HNEA: predicting metabolic pathway types of chemicals and enzymes with a heterogeneous network embedding algorithm. Curr Bioinform 18(9):748–759
Tang S, Chen L (2022) iATC-NFMLP: identifying classes of anatomical therapeutic chemicals based on drug networks, fingerprints and multilayer perceptron. Curr Bioinform 17(9):814–824
Matthews B (1975) Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochimica et Biophysica Acta (BBA)-Protein Struct 405(2):442–451
Magin C, Löwer R, Löwer J (1999) cORF and RcRE, the Rev/Rex and RRE/RxRE homologues of the human endogenous retrovirus family HTDV/HERV-K. J Virol 73(11):9496–9507
Gray LR et al (2019) HIV-1 Rev interacts with HERV-K RcREs present in the human genome and promotes export of unspliced HERV-K proviral RNA. Retrovirology 16:1–17
Zhang L, et al. (2020) SARS-CoV-2 RNA reverse-transcribed and integrated into the human genome. BioRxiv 2020.12. 12.422516
Crooke PS et al (2021) Cutting edge: reduced adenosine-to-inosine editing of endogenous Alu RNAs in severe COVID-19 disease. J Immunol 206(8):1691–1696
Pang X, et al. (2021) Emerging SARS-CoV-2 mutation hotspots associated with clinical outcomes. bioRxiv 2021: 2021.03. 31.437666.
Picardi E, Mansi L, Pesole G (2021) Detection of A-to-I RNA editing in SARS-COV-2. Genes 13(1):41
Russo RC et al (2014) The CXCL8/IL-8 chemokine family and its receptors in inflammatory diseases. Expert Rev Clin Immunol 10(5):593–619
Park JH, Lee HK (2020) Re-analysis of single cell transcriptome reveals that the NR3C1-CXCL8-neutrophil axis determines the severity of COVID-19. Front Immunol 11:2145
Pius-Sadowska E et al (2022) CXCL8, CCL2, and CMV seropositivity as new prognostic factors for a severe COVID-19 course. Int J Mol Sci 23(19):11338
Huang Y et al (2020) The associations between fasting plasma glucose levels and mortality of COVID-19 in patients without diabetes. Diabetes Res Clin Pract 169:108448
Nouailles G et al (2021) Temporal omics analysis in Syrian hamsters unravel cellular effector responses to moderate COVID-19. Nat Commun 12(1):4869
Zhang J-Y et al (2020) Single-cell landscape of immunological responses in patients with COVID-19. Nat Immunol 21(9):1107–1118
Wang Y, et al. Single-cell transcriptomic atlas of individuals receiving inactivated COVID-19 vaccines reveals distinct immunological responses between vaccine and natural SARS-CoV-2 infection. medRxiv, 2021: 2021.08. 30.21262863
Vastrad BM, Vastrad CM (2021) Bioinformatics analysis of expression profiling by high throughput sequencing for identification of potential key genes among SARS-CoV-2/COVID 19. Researchsquare
Sarohan AR, et al. Retinol depletion in severe COVID-19. medRxiv 2021: 2021.01. 30.21250844
Guardela BMJ et al (2021) 50-gene risk profiles in peripheral blood predict COVID-19 outcomes: a retrospective, multicenter cohort study. EBioMedicine 69:103439
Hsu Y-L et al (2017) Identification of novel gene expression signature in lung adenocarcinoma by using next-generation sequencing data and bioinformatics analysis. Oncotarget 8(62):104831
Charitou T et al (2022) Drug genetic associations with COVID-19 manifestations: a data mining and network biology approach. Pharmacogenomics J 22(5–6):294–302
Gorodin V et al (2021) Role of polymorphisms of genes involved in hemostasis in COVID-19 pathogenesis. Infektsionnye Bolezni 19(2):16–26
Fu L et al (2022) Using bioinformatics and systems biology to discover common pathogenetic processes between sarcoidosis and COVID-19. Gene Rep 27:101597
Nikitopoulou I et al (2021) Increased autotaxin levels in severe COVID-19, correlating with IL-6 levels, endothelial dysfunction biomarkers, and impaired functions of dendritic cells. Int J Mol Sci 22(18):10006
Duhalde Vega M et al (2022) PD-1/PD-L1 blockade abrogates a dysfunctional innate-adaptive immune axis in critical β-coronavirus disease. Sci Adv 8(38):eabn6545
Funding
This work was supported by the National Key R&D Program of China (2022YFF1203202), Strategic Priority Research Program of Chinese Academy of Sciences (XDA26040304, XDB38050200), the Fund of the Key Laboratory of Tissue Microenvironment and Tumor of Chinese Academy of Sciences (202002), and Shandong Provincial Natural Science Foundation (ZR2022MC072).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Conceptualization: Tao Huang, Yu-Dong Cai; methodology: JingXin Ren, Qian Gao, Lei Chen, KaiYan Feng; formal analysis and investigation: JingXin Ren, XianChao Zhou, Wei Guo; writing — original draft preparation: JingXin Ren, Qian Gao, XianChao Zhou; writing — review and editing: Tao Huang; funding acquisition: Tao Huang, Yu-Dong Cai; supervision: Yu-Dong Cai.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ren, J., Gao, Q., Zhou, X. et al. Identification of key gene expression associated with quality of life after recovery from COVID-19. Med Biol Eng Comput 62, 1031–1048 (2024). https://doi.org/10.1007/s11517-023-02988-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-023-02988-8