Abstract
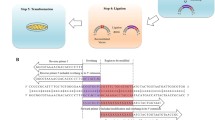
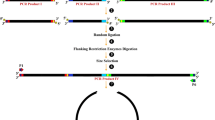
Inverse PCR is a powerful tool for the rapid introduction of desired mutations at desired positions in a circular double-stranded DNA sequence. Here, custom-designed mutant primers oriented in the inverse direction are used to amplify the entire circular template with incorporation of the required mutation(s). By careful primer design it can be used to perform such diverse modifications as the introduction of point mutations and multiple mutations, the insertion of new sequences, and even sequence deletions. Three primer formats are commonly used; nonoverlapping, partially overlapping and fully overlapping primers, and here we describe the use of nonoverlapping primers for introduction of a point mutation. Use of such a primer setup in the PCR reaction, with one of the primers containing the desired mismatch mutation, results in the amplification of a linear, double-stranded, mutated product. Methylated template DNA is removed from the nonmethylated PCR product by DpnI digestion and the PCR product is then phosphorylated by polynucleotide kinase treatment before being recircularized by ligation, and transformed to E. coli. This relatively simple site-directed mutagenesis procedure is of major importance in biology and biotechnology today where it is commonly employed for the study and engineering of DNA, RNA, and proteins.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bommarius AS, Paye MF (2013) Stabilizing biocatalysts. Chem Soc Rev 42:6534–6565. doi:10.1039/c3cs60137d
Collins T, De Vos D, Hoyoux A, Savvides SN, Gerday C, Van Beeumen J, Feller G (2005) Study of the active site residues of a glycoside hydrolase family 8 xylanase. J Mol Biol 354(2):425–435
Davids T, Schmidt M, Bottcher D, Bornscheuer UT (2013) Strategies for the discovery and engineering of enzymes for biocatalysis. Curr Opin Chem Biol 17(2):215–220. doi:10.1016/j.cbpa.2013.02.022
Sanjuan R (2010) Mutational fitness effects in RNA and single-stranded DNA viruses: common patterns revealed by site-directed mutagenesis studies. Philos Trans R Soc Lond B Biol Sci 365(1548):1975–1982. doi:10.1098/rstb.2010.0063
Steiner K, Malke H (1997) Primary structure requirements for in vivo activity and bidirectional function of the transcription terminator shared by the oppositely oriented skc/rel-orf1 genes of Streptococcus equisimilis H46A. Mol Gen Genet 255(6):611–618
Traboni C, Ciliberto G, Cortese R (1982) A novel method for site-directed mutagenesis: its application to an eukaryotic tRNAPro gene promoter. EMBO J 1(4):415–420
Hutchison CA, Phillips S, Edgell MH, Gillam S, Jahnke P, Smith M (1978) Mutagenesis at a specific position in a DNA sequence. J Biol Chem 253(18):6551–6560
Hutchison CA 3rd, Edgell MH (1971) Genetic assay for small fragments of bacteriophage phi X174 deoxyribonucleic acid. J Virol 8(2):181–189
Edgell MH, Hutchison CA 3rd, Sclair M (1972) Specific endonuclease R fragments of bacteriophage phiX174 deoxyribonucleic acid. J Virol 9(4):574–582
Reeves AR (ed) (2016) In vitro mutagenesis. Methods and protocols, series vol 1498, 1 edn. Humana, New York, NY. doi:10.1007/978-1-4939-6472-7
Ho SN, Hunt HD, Horton RM, Pullen JK, Pease LR (1989) Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 77(1):51–59
Stuckey S, Storici F (2013) Gene knockouts, in vivo site-directed mutagenesis and other modifications using the delitto perfetto system in Saccharomyces cerevisiae. Methods Enzymol 533:103–131. doi:10.1016/B978-0-12-420067-8.00008-8
Kren BT, Bandyopadhyay P, Steer CJ (1998) In vivo site-directed mutagenesis of the factor IX gene by chimeric RNA/DNA oligonucleotides. Nat Med 4(3):285–290
Higuchi R, Krummel B, Saiki RK (1988) A general method of in vitro preparation and specific mutagenesis of DNA fragments: study of protein and DNA interactions. Nucleic Acids Res 16(15):7351–7367
Triglia T, Peterson MG, Kemp DJ (1988) A procedure for in vitro amplification of DNA segments that lie outside the boundaries of known sequences. Nucleic Acids Res 16(16):8186
Ochman H, Gerber AS, Hartl DL (1988) Genetic applications of an inverse polymerase chain reaction. Genetics 120(3):621–623
Hemsley A, Arnheim N, Toney MD, Cortopassi G, Galas DJ (1989) A simple method for site-directed mutagenesis using the polymerase chain reaction. Nucleic Acids Res 17(16):6545–6551
Qi D, Scholthof KB (2008) A one-step PCR-based method for rapid and efficient site-directed fragment deletion, insertion, and substitution mutagenesis. J Virol Methods 149(1):85–90. doi:10.1016/j.jviromet.2008.01.002
Zheng L, Baumann U, Reymond JL (2004) An efficient one-step site-directed and site-saturation mutagenesis protocol. Nucleic Acids Res 32(14):e115. doi:10.1093/nar/gnh110
Liu H, Naismith JH (2008) An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol 8:91. doi:10.1186/1472-6750-8-91
Xia Y, Chu W, Qi Q, Xun L (2015) New insights into the QuikChange process guide the use of Phusion DNA polymerase for site-directed mutagenesis. Nucleic Acids Res 43(2):e12. doi:10.1093/nar/gku1189
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Silva, D., Santos, G., Barroca, M., Collins, T. (2017). Inverse PCR for Point Mutation Introduction. In: Domingues, L. (eds) PCR. Methods in Molecular Biology, vol 1620. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-7060-5_5
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7060-5_5
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-7059-9
Online ISBN: 978-1-4939-7060-5
eBook Packages: Springer Protocols