Biases in Illumina transcriptome sequencing caused by random hexamer priming
- PMID: 20395217
- PMCID: PMC2896536
- DOI: 10.1093/nar/gkq224
Biases in Illumina transcriptome sequencing caused by random hexamer priming
Abstract
Generation of cDNA using random hexamer priming induces biases in the nucleotide composition at the beginning of transcriptome sequencing reads from the Illumina Genome Analyzer. The bias is independent of organism and laboratory and impacts the uniformity of the reads along the transcriptome. We provide a read count reweighting scheme, based on the nucleotide frequencies of the reads, that mitigates the impact of the bias.
Figures


 . (b) As in (a), but the hexamers correspond to positions 25–30 of the aligned reads, with a correlation of
. (b) As in (a), but the hexamers correspond to positions 25–30 of the aligned reads, with a correlation of  .
.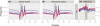
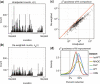
 goodness-of-fit statistics based on unadjusted and reweighted counts for 552 highly expressed regions of constant expression. (d) Smoothed histograms of the reduction in
goodness-of-fit statistics based on unadjusted and reweighted counts for 552 highly expressed regions of constant expression. (d) Smoothed histograms of the reduction in  goodness-of-fit statistics when using the re-weighting scheme, evaluated in five different experiments. Values greater than zero indicate that the re-weighting scheme improves the uniformity of the read distribution.
goodness-of-fit statistics when using the re-weighting scheme, evaluated in five different experiments. Values greater than zero indicate that the re-weighting scheme improves the uniformity of the read distribution.Similar articles
-
RNA sequencing and quantitation using the Helicos Genetic Analysis System.Methods Mol Biol. 2011;733:37-49. doi: 10.1007/978-1-61779-089-8_3. Methods Mol Biol. 2011. PMID: 21431761
-
Consistent errors in first strand cDNA due to random hexamer mispriming.PLoS One. 2013 Dec 30;8(12):e85583. doi: 10.1371/journal.pone.0085583. eCollection 2013. PLoS One. 2013. PMID: 24386481 Free PMC article.
-
Characterizing the mouse ES cell transcriptome with Illumina sequencing.Genomics. 2008 Oct;92(4):187-94. doi: 10.1016/j.ygeno.2008.05.011. Epub 2008 Aug 3. Genomics. 2008. PMID: 18602984
-
Substantial biases in ultra-short read data sets from high-throughput DNA sequencing.Nucleic Acids Res. 2008 Sep;36(16):e105. doi: 10.1093/nar/gkn425. Epub 2008 Jul 26. Nucleic Acids Res. 2008. PMID: 18660515 Free PMC article.
-
De novo assembly of short sequence reads.Brief Bioinform. 2010 Sep;11(5):457-72. doi: 10.1093/bib/bbq020. Epub 2010 Aug 19. Brief Bioinform. 2010. PMID: 20724458 Review.
Cited by
-
Enhancing RNA-seq analysis by addressing all co-existing biases using a self-benchmarking approach with 2D structural insights.Brief Bioinform. 2024 Sep 23;25(6):bbae532. doi: 10.1093/bib/bbae532. Brief Bioinform. 2024. PMID: 39428128 Free PMC article.
-
Enhancing RNA-seq bias mitigation with the Gaussian self-benchmarking framework: towards unbiased sequencing data.BMC Genomics. 2024 Sep 30;25(1):904. doi: 10.1186/s12864-024-10814-0. BMC Genomics. 2024. PMID: 39350040 Free PMC article.
-
Enhancing transcriptome expression quantification through accurate assignment of long RNA sequencing reads with TranSigner.bioRxiv [Preprint]. 2024 Aug 17:2024.04.13.589356. doi: 10.1101/2024.04.13.589356. bioRxiv. 2024. PMID: 39185147 Free PMC article. Preprint.
-
Sequencing accuracy and systematic errors of nanopore direct RNA sequencing.BMC Genomics. 2024 May 28;25(1):528. doi: 10.1186/s12864-024-10440-w. BMC Genomics. 2024. PMID: 38807060 Free PMC article.
-
BEERS2: RNA-Seq simulation through high fidelity in silico modeling.Brief Bioinform. 2024 Mar 27;25(3):bbae164. doi: 10.1093/bib/bbae164. Brief Bioinform. 2024. PMID: 38605641 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

