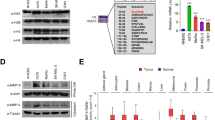
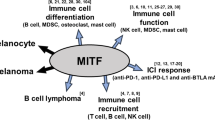
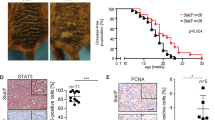
Abstract
Malignant melanoma is an aggressive cancer known for its notorious resistance to most current therapies. The basic helix-loop-helix microphthalmia transcription factor (MITF) is the master regulator determining the identity and properties of the melanocyte lineage, and is regarded as a lineage-specific ‘oncogene’ that has a critical role in the pathogenesis of melanoma. MITF promotes melanoma cell proliferation, whereas sustained supression of MITF expression leads to senescence. By combining chromatin immunoprecipitation coupled to high throughput sequencing (ChIP-seq) and RNA sequencing analyses, we show that MITF directly regulates a set of genes required for DNA replication, repair and mitosis. Our results reveal how loss of MITF regulates mitotic fidelity, and through defective replication and repair induces DNA damage, ultimately ending in cellular senescence. These findings reveal a lineage-specific control of DNA replication and mitosis by MITF, providing new avenues for therapeutic intervention in melanoma. The identification of MITF-binding sites and gene-regulatory networks establish a framework for understanding oncogenic basic helix-loop-helix factors such as N-myc or TFE3 in other cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Abraham J, Lemmers B, Hande MP, Moynahan ME, Chahwan C, Ciccia A et al. (2003). Eme1 is involved in DNA damage processing and maintenance of genomic stability in mammalian cells. EMBO J 22: 6137–6147.
Artandi SE, DePinho RA . (2000). Mice without telomerase: what can they teach us about human cancer? Nat Med 6: 852–855.
Artandi SE, DePinho RA . (2010). Telomeres and telomerase in cancer. Carcinogenesis 31: 9–18.
Bentley DJ, Harrison C, Ketchen AM, Redhead NJ, Samuel K, Waterfall M et al. (2002). DNA ligase I null mouse cells show normal DNA repair activity but altered DNA replication and reduced genome stability. J Cell Sci 115: 1551–1561.
Bentley NJ, Eisen T, Goding CR . (1994). Melanocyte-specific expression of the human tyrosinase promoter: activation by the microphthalmia gene product and role of the initiator. Mol Cell Biol 14: 7996–8006.
Bertolotto C, Busca R, Abbe P, Bille K, Aberdam E, Ortonne JP et al. (1998). Different cis-acting elements are involved in the regulation of TRP1 and TRP2 promoter activities by cyclic AMP: pivotal role of M boxes (GTCATGTGCT) and of microphthalmia. Mol Cell Biol 18: 694–702.
Beuret L, Flori E, Denoyelle C, Bille K, Busca R, Picardo M et al. (2007). Up-regulation of MET expression by alpha-melanocyte-stimulating hormone and MITF allows hepatocyte growth factor to protect melanocytes and melanoma cells from apoptosis. J Biol Chem 282: 14140–14147.
Busca R, Berra E, Gaggioli C, Khaled M, Bille K, Marchetti B et al. (2005). Hypoxia-inducible factor 1\{alpha\} is a new target of microphthalmia-associated transcription factor (MITF) in melanoma cells. J Cell Biol 170: 49–59.
Carreira S, Goodall J, Aksan I, La Rocca SA, Galibert MD, Denat L et al. (2005). Mitf cooperates with Rb1 and activates p21Cip1 expression to regulate cell cycle progression. Nature 433: 764–769.
Carreira S, Goodall J, Denat L, Rodriguez M, Nuciforo P, Hoek KS et al. (2006). Mitf regulation of Dia1 controls melanoma proliferation and invasiveness. Genes Dev 20: 3426–3439.
Carroll CW, Silva MC, Godek KM, Jansen LE, Straight AF . (2009). Centromere assembly requires the direct recognition of CENP-A nucleosomes by CENP-N. Nat Cell Biol 11: 896–902.
Cheeseman IM, Chappie JS, Wilson-Kubalek EM, Desai A . (2006). The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell 127: 983–997.
Chiaverini C, Beuret L, Flori E, Busca R, Abbe P, Bille K et al. (2008). Microphthalmia-associated transcription factor regulates RAB27A gene expression and controls melanosome transport. J Biol Chem 283: 12635–12642.
Delacroix L, Moutier E, Altobelli G, Legras S, Poch O, Choukrallah MA et al. (2010). Cell-specific interaction of retinoic acid receptors with target genes in mouse embryonic fibroblasts and embryonic stem cells. Mol Cell Biol 30: 231–244.
Du J, Widlund HR, Horstmann MA, Ramaswamy S, Ross K, Huber WE et al. (2004). Critical role of CDK2 for melanoma growth linked to its melanocyte-specific transcriptional regulation by MITF. Cancer Cell 6: 565–576.
Dynek JN, Chan SM, Liu J, Zha J, Fairbrother WJ, Vucic D . (2008). Microphthalmia-associated transcription factor is a critical transcriptional regulator of melanoma inhibitor of apoptosis in melanomas. Cancer Res 68: 3124–3132.
Fagiani E, Giardina G, Luzi L, Cesaroni M, Quarto M, Capra M et al. (2007). RaLP, a new member of the Src homology and collagen family, regulates cell migration and tumor growth of metastatic melanomas. Cancer Res 67: 3064–3073.
Foltz DR, Jansen LE, Black BE, Bailey AO, Yates III JR, Cleveland DW . (2006). The human CENP-A centromeric nucleosome-associated complex. Nat Cell Biol 8: 458–469.
Garraway LA, Weir BA, Zhao X, Widlund H, Beroukhim R, Berger A et al. (2005a). ‘Lineage addiction’ in human cancer: lessons from integrated genomics. Cold Spring Harb Symp Quant Biol 70: 25–34.
Garraway LA, Widlund HR, Rubin MA, Getz G, Berger AJ, Ramaswamy S et al. (2005b). Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature 436: 117–122.
Giuliano S, Cheli Y, Ohanna M, Bonet C, Beuret L, Bille K et al. (2010). Microphthalmia-associated transcription factor controls the DNA damage response and a lineage-specific senescence program in melanomas. Cancer Res 70: 3813–3822.
Goding CR . (2000a). Mitf from neural crest to melanoma: signal transduction and transcription in the melanocyte lineage. Genes Dev 14: 1712–1728.
Goding CR . (2000b). Melanocyte development and malignant melanoma. Forum (Genova) 10: 176–187.
Gomez-Baldo L, Schmidt S, Maxwell CA, Bonifaci N, Gabaldon T, Vidalain PO et al. (2010). TACC3-TSC2 maintains nuclear envelope structure and controls cell division. Cell Cycle 9: 1143–1155.
Goodall J, Carreira S, Denat L, Kobi D, Davidson I, Nuciforo P et al. (2008). Brn-2 represses microphthalmia-associated transcription factor expression and marks a distinct subpopulation of microphthalmia-associated transcription factor-negative melanoma cells. Cancer Res 68: 7788–7794.
Hoek KS, Eichhoff OM, Schlegel NC, Dobbeling U, Kobert N, Schaerer L et al. (2008a). in vivo switching of human melanoma cells between proliferative and invasive states. Cancer Res 68: 650–656.
Hoek KS, Schlegel NC, Brafford P, Sucker A, Ugurel S, Kumar R et al. (2006). Metastatic potential of melanomas defined by specific gene expression profiles with no BRAF signature. Pigment Cell Res 19: 290–302.
Hoek KS, Schlegel NC, Eichhoff OM, Widmer DS, Praetorius C, Einarsson SO et al. (2008b). Novel MITF targets identified using a two-step DNA microarray strategy. Pigment Cell Melanoma Res 21: 665–676.
Hou L, Pavan WJ . (2008). Transcriptional and signaling regulation in neural crest stem cell-derived melanocyte development: do all roads lead to Mitf? Cell Res 18: 1163–1176.
Kim DS, Park SH, Park KC . (2004). Transforming growth factor-beta1 decreases melanin synthesis via delayed extracellular signal-regulated kinase activation. Int J Biochem Cell Biol 36: 1482–1491.
Kim NW, Piatyszek MA, Prowse KR, Harley CB, West MD, Ho PL et al. (1994). Specific association of human telomerase activity with immortal cells and cancer. Science 266: 2011–2015.
Kobi D, Steunou AL, Dembele D, Legras S, Larue L, Nieto L et al. (2010). Genome-wide analysis of POU3F2/BRN2 promoter occupancy in human melanoma cells reveals Kitl as a novel regulated target gene. Pigment Cell Melanoma Res 23: 404–418.
Koyanagi K, O'Day SJ, Gonzalez R, Lewis K, Robinson WA, Amatruda TT et al. (2006). Microphthalmia transcription factor as a molecular marker for circulating tumor cell detection in blood of melanoma patients. Clin Cancer Res 12: 1137–1143.
Krebs A, Frontini M, Tora L . (2008). GPAT: retrieval of genomic annotation from large genomic position datasets. BMC Bioinformatics 9: 533.
Larribere L, Hilmi C, Khaled M, Gaggioli C, Bille K, Auberger P et al. (2005). The cleavage of microphthalmia-associated transcription factor, MITF, by caspases plays an essential role in melanocyte and melanoma cell apoptosis. Genes Dev 19: 1980–1985.
Levy C, Khaled M, Robinson KC, Veguilla RA, Chen PH, Yokoyama S et al. (2010). Lineage -specific transcriptional regulation of DICER by MITF in melanocytes. Cell 141: 994–1005.
McGill GG, Horstmann M, Widlund HR, Du J, Motyckova G, Nishimura EK et al. (2002). Bcl2 regulation by the melanocyte master regulator Mitf modulates lineage survival and melanoma cell viability. Cell 109: 707–718.
Moore R, Champeval D, Denat L, Tan SS, Faure F, Julien-Grille S et al. (2004). Involvement of cadherins 7 and 20 in mouse embryogenesis and melanocyte transformation. Oncogene 23: 6726–6735.
Neumann B, Walter T, Heriche JK, Bulkescher J, Erfle H, Conrad C et al. (2010). Phenotypic profiling of the human genome by time-lapse microscopy reveals cell division genes. Nature 464: 721–727.
Nishimura EK, Suzuki M, Igras V, Du J, Lonning S, Miyachi Y et al. (2010). Key roles for transforming growth factor beta in melanocyte stem cell maintenance. Cell Stem Cell 6: 130–140.
Ouhtit A, Gaur RL, Abd Elmageed ZY, Fernando A, Thouta R, Trappey AK et al. (2009). Towards understanding the mode of action of the multifaceted cell adhesion receptor CD146. Biochim Biophys Acta 1795: 130–136.
Pinner S, Jordan P, Sharrock K, Bazley L, Collinson L, Marais R et al. (2009). Intravital imaging reveals transient changes in pigment production and Brn2 expression during metastatic melanoma dissemination. Cancer Res 69: 7969–7977.
Rothhammer T, Poser I, Soncin F, Bataille F, Moser M, Bosserhoff AK . (2005). Bone morphogenic proteins are overexpressed in malignant melanoma and promote cell invasion and migration. Cancer Res 65: 448–456.
Ruchaud S, Carmena M, Earnshaw WC . (2007). Chromosomal passengers: conducting cell division. Nat Rev Mol Cell Biol 8: 798–812.
Rudolph KL, Chang S, Lee HW, Blasco M, Gottlieb GJ, Greider C et al. (1999). Longevity, stress response, and cancer in aging telomerase-deficient mice. Cell 96: 701–712.
Schmit TL, Zhong W, Setaluri V, Spiegelman VS, Ahmad N . (2009). Targeted depletion of Polo-like kinase (Plk) 1 through lentiviral shRNA or a small-molecule inhibitor causes mitotic catastrophe and induction of apoptosis in human melanoma cells. J Invest Dermatol 129: 2843–2853.
Soengas MS, Lowe SW . (2003). Apoptosis and melanoma chemoresistance. Oncogene 22: 3138–3151.
Steingrimsson E . (2008). All for one, one for all: alternative promoters and Mitf. Pigment Cell Melanoma Res 21: 412–414.
Tomkinson AE, Mackey ZB . (1998). Structure and function of mammalian DNA ligases. Mutat Res 407: 1–9.
Wang W . (2007). Emergence of a DNA-damage response network consisting of Fanconi anaemia and BRCA proteins. Nat Rev Genet 8: 735–748.
Wu G, Lin YT, Wei R, Chen Y, Shan Z, Lee WH . (2008). Hice1, a novel microtubule-associated protein required for maintenance of spindle integrity and chromosomal stability in human cells. Mol Cell Biol 28: 3652–3662.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE et al. (2008). Model-based analysis of ChIP-Seq (MACS). Genome Biol 9: R137.
Acknowledgements
We thank B Jost for Illumina sequencing, A Hamiche and E Soutoglou for antibodies. This work was supported by grants from the CNRS, the INSERM, the Association pour la Recherche contre le Cancer grant 4985 and 7823, the Ligue Nationale contre le Cancer, the Institut national du Cancer grant R08009AP and the ANR Regulome project grant. We thank the Conseil Regional de la Région Provence-Alpes-Cote d’Azur, the Nice University Hospital and the Conseil Général des Alpes Maritimes for funding of the C3M MiCa Cell Imaging Facility. ID and CB are ‘équipes labellisées’ of the Ligue Nationale contre le Cancer.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Strub, T., Giuliano, S., Ye, T. et al. Essential role of microphthalmia transcription factor for DNA replication, mitosis and genomic stability in melanoma. Oncogene 30, 2319–2332 (2011). https://doi.org/10.1038/onc.2010.612
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.612
Keywords
This article is cited by
-
ZEB1 controls a lineage-specific transcriptional program essential for melanoma cell state transitions
Oncogene (2024)
-
Novel cellular systems unveil mucosal melanoma initiating cells and a role for PI3K/Akt/mTOR pathway in mucosal melanoma fitness
Journal of Translational Medicine (2024)
-
Fanca deficiency is associated with alterations in osteoclastogenesis that are rescued by TNFα
Cell & Bioscience (2023)
-
GREB1 isoform 4 is specifically transcribed by MITF and required for melanoma proliferation
Oncogene (2023)
-
Transglutaminase Type 2-MITF axis regulates phenotype switching in skin cutaneous melanoma
Cell Death & Disease (2023)