Abstract
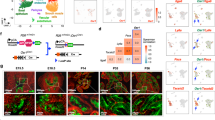
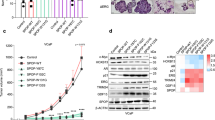
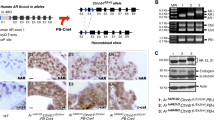
Studies of ETS-mediated prostate oncogenesis have been hampered by a lack of suitable experimental systems. Here we describe a new conditional mouse model that shows robust, homogenous ERG expression throughout the prostate. When combined with homozygous Pten loss, the mice developed accelerated, highly penetrant invasive prostate cancer. In mouse prostate tissue, ERG markedly increased androgen receptor (AR) binding. Robust ERG-mediated transcriptional changes, observed only in the setting of Pten loss, included the restoration of AR transcriptional output and upregulation of genes involved in cell death, migration, inflammation and angiogenesis. Similarly, ETS variant 1 (ETV1) positively regulated the AR cistrome and transcriptional output in ETV1-translocated, PTEN-deficient human prostate cancer cells. In two large clinical cohorts, expression of ERG and ETV1 correlated with higher AR transcriptional output in PTEN-deficient prostate cancer specimens. We propose that ETS factors cause prostate-specific transformation by altering the AR cistrome, priming the prostate epithelium to respond to aberrant upstream signals such as PTEN loss.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Tomlins, S.A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Tomlins, S.A. et al. TMPRSS2:ETV4 gene fusions define a third molecular subtype of prostate cancer. Cancer Res. 66, 3396–3400 (2006).
Helgeson, B.E. et al. Characterization of TMPRSS2:ETV5 and SLC45A3:ETV5 gene fusions in prostate cancer. Cancer Res. 68, 73–80 (2008).
Paulo, P. et al. FLI1 is a novel ETS transcription factor involved in gene fusions in prostate cancer. Genes Chromosom. Cancer 51, 240–249 (2012).
Mosquera, J.M. et al. Characterization of TMPRSS2-ERG fusion high-grade prostatic intraepithelial neoplasia and potential clinical implications. Clin. Cancer Res. 14, 3380–3385 (2008).
Börno, S.T. et al. Genome-wide DNA methylation events in TMPRSS2-ERG fusion-negative prostate cancers implicate an EZH2-dependent mechanism with miR-26a hypermethylation. Cancer Discov. 2, 1024–1035 (2012).
Grasso, C.S. et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature 487, 239–243 (2012).
King, J.C. et al. Cooperativity of TMPRSS2-ERG with PI3-kinase pathway activation in prostate oncogenesis. Nat. Genet. 41, 524–526 (2009).
Carver, B.S. et al. Reciprocal feedback regulation of PI3K and androgen receptor signaling in PTEN-deficient prostate cancer. Cancer Cell 19, 575–586 (2011).
Casey, O.M. et al. TMPRSS2-driven ERG expression in vivo increases self-renewal and maintains expression in a castration resistant subpopulation. PLoS ONE 7, e41668 (2012).
Tomlins, S.A. et al. Distinct classes of chromosomal rearrangements create oncogenic ETS gene fusions in prostate cancer. Nature 448, 595–599 (2007).
Shin, S. et al. Induction of prostatic intraepithelial neoplasia and modulation of androgen receptor by ETS variant 1/ETS-related protein 81. Cancer Res. 69, 8102–8110 (2009).
Clegg, N.J. et al. MYC cooperates with AKT in prostate tumorigenesis and alters sensitivity to mTOR inhibitors. PLoS ONE 6, e17449 (2011).
Wu, X. et al. Generation of a prostate epithelial cell–specific Cre transgenic mouse model for tissue-specific gene ablation. Mech. Dev. 101, 61–69 (2001).
Zong, Y. et al. ETS family transcription factors collaborate with alternative signaling pathways to induce carcinoma from adult murine prostate cells. Proc. Natl. Acad. Sci. USA 106, 12465–12470 (2009).
Tomlins, S.A. et al. Role of the TMPRSS2-ERG gene fusion in prostate cancer. Neoplasia 10, 177–188 (2008).
Klezovitch, O. et al. A causal role for ERG in neoplastic transformation of prostate epithelium. Proc. Natl. Acad. Sci. USA 105, 2105–2110 (2008).
Carver, B.S. et al. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat. Genet. 41, 619–624 (2009).
Wang, S. et al. Pten deletion leads to the expansion of a prostatic stem/progenitor cell subpopulation and tumor initiation. Proc. Natl. Acad. Sci. USA 103, 1480–1485 (2006).
Verhagen, A.P. et al. Colocalization of basal and luminal cell–type cytokeratins in human prostate cancer. Cancer Res. 52, 6182–6187 (1992).
Yu, J. et al. An integrated network of androgen receptor, polycomb, and TMPRSS2-ERG gene fusions in prostate cancer progression. Cancer Cell 17, 443–454 (2010).
Chng, K.R. et al. A transcriptional repressor co-regulatory network governing androgen response in prostate cancers. EMBO J. 31, 2810–2823 (2012).
Bailey, T.L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–208 (2009).
Bailey, T.L. DREME: motif discovery in transcription factor ChIP-seq data. Bioinformatics 27, 1653–1659 (2011).
Lupien, M. et al. FoxA1 translates epigenetic signatures into enhancer-driven lineage-specific transcription. Cell 132, 958–970 (2008).
Andreu-Vieyra, C. et al. Dynamic nucleosome-depleted regions at androgen receptor enhancers in the absence of ligand in prostate cancer cells. Mol. Cell Biol. 31, 4648–4662 (2011).
Heintzman, N.D. et al. Histone modifications at human enhancers reflect global cell-type–specific gene expression. Nature 459, 108–112 (2009).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Sahu, B. et al. Dual role of FoxA1 in androgen receptor binding to chromatin, androgen signalling and prostate cancer. EMBO J. 30, 3962–3976 (2011).
Wang, D. et al. Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 474, 390–394 (2011).
Lien, W.H. et al. Genome-wide maps of histone modifications unwind in vivo chromatin states of the hair follicle lineage. Cell Stem Cell 9, 219–232 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Taylor, B.S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
Rickman, D.S. et al. ERG cooperates with androgen receptor in regulating trefoil factor 3 in prostate cancer disease progression. Neoplasia 12, 1031–1040 (2010).
Huang da, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Chen, Z. et al. Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis. Nature 436, 725–730 (2005).
Mulholland, D.J. et al. Cell autonomous role of PTEN in regulating castration-resistant prostate cancer growth. Cancer Cell 19, 792–804 (2011).
Chi, P. et al. ETV1 is a lineage survival factor that cooperates with KIT in gastrointestinal stromal tumours. Nature 467, 849–853 (2010).
Cai, C. et al. ERG induces androgen receptor–mediated regulation of SOX9 in prostate cancer. J. Clin. Invest. 123, 1109–1122 (2013).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Ivanova, A. et al. In vivo genetic ablation by Cre-mediated expression of diphtheria toxin fragment A. Genesis 43, 129–135 (2005).
Murtaugh, L.C., Stanger, B.Z., Kwan, K.M. & Melton, D.A. Notch signaling controls multiple steps of pancreatic differentiation. Proc. Natl. Acad. Sci. USA 100, 14920–14925 (2003).
Scher, H.I. et al. Antitumour activity of MDV3100 in castration-resistant prostate cancer: a phase 1–2 study. Lancet 375, 1437–1446 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Acknowledgements
We thank the MSKCC Gene Targeting (C. Yang), Mouse Genetics Core (W. Mark and P. Romanienko), Genomics Core Laboratory (A. Viale), Molecular Cytogenetics (M. Leversha) and Molecular Cytology (K. Manova) core facilities and the Rockefeller University Genomics Core (S. Dewell). This work is supported in part by the Howard Hughes Medical Institute (C.L.S.), the US National Cancer Institute (K08CA140946, Y.C.; 1K08CA151660-01A1, P.C.; U01 CA141502, C.L.S.; and P50CA092629, C.L.S. and B.S.C.), the US Department of Defense (W81XWH-10-1-0197, Y.C.), the Prostate Cancer Foundation (Y.C. and B.S.C.), the Starr Cancer Consortium (Y.C., P.C., C.L.S. and D.Z.) and the US National Institute of Mental Health (R21MH099452, D.Z.). We thank J.P. Martinez-Barbera (University College London) for plasmids, C.D. Allis, F. Giancotti and J. Otero for conceptual input, N. Schultz for bioinformatics assistance and A. Gopalan and S. Couto for pathology input.
Author information
Authors and Affiliations
Contributions
Y.C. and C.L.S. conceived of the project. Y.C. performed the mouse experiments with technical support from J.W., T.S. and D.G. P.C. performed expression profiling and ChIP-seq with technical support from S.S. and I.S. S.R. and D.Z. performed bioinformatics analysis. P.J.I. performed the LNCaP ETV1 knockdown experiments. B.S.C. and H.I.S. provided major intellectual input for the initial project design and further troubleshooting. Y.C., P.C. and C.L.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–23 and Supplementary Tables 1–8 (PDF 4535 kb)
Rights and permissions
About this article
Cite this article
Chen, Y., Chi, P., Rockowitz, S. et al. ETS factors reprogram the androgen receptor cistrome and prime prostate tumorigenesis in response to PTEN loss. Nat Med 19, 1023–1029 (2013). https://doi.org/10.1038/nm.3216
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3216