Abstract
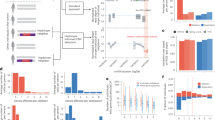
Structural variations of DNA greater than 1 kilobase in size account for most bases that vary among human genomes, but are still relatively under-ascertained. Here we use tiling oligonucleotide microarrays, comprising 42 million probes, to generate a comprehensive map of 11,700 copy number variations (CNVs) greater than 443 base pairs, of which most (8,599) have been validated independently. For 4,978 of these CNVs, we generated reference genotypes from 450 individuals of European, African or East Asian ancestry. The predominant mutational mechanisms differ among CNV size classes. Retrotransposition has duplicated and inserted some coding and non-coding DNA segments randomly around the genome. Furthermore, by correlation with known trait-associated single nucleotide polymorphisms (SNPs), we identified 30 loci with CNVs that are candidates for influencing disease susceptibility. Despite this, having assessed the completeness of our map and the patterns of linkage disequilibrium between CNVs and SNPs, we conclude that, for complex traits, the heritability void left by genome-wide association studies will not be accounted for by common CNVs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Data deposits
The CNV discovery and CNV genotyping data are available at ArrayExpress (http://www.ebi.ac.uk/microarray-as/ae/) under accession numbers E-MTAB-40 and E-MTAB-142, respectively. Normalized CNV discovery data are available at http://www.sanger.ac.uk/humgen/cnv/42mio. CNVs are displayed at the Database of Genomic Variants (http://projects.tcag.ca/variation). CNV locations and genotypes are reported in Supplementary Tables 1 and 2.
References
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004)
Levy, S. & Strausberg, R. L. Human genetics: Individual genomes diversify. Nature 456, 49–51 (2008)
Frazer, K. A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007)
Marchini, J. et al. A new multipoint method for genome-wide association studies by imputation of genotypes. Nature Genet. 39, 906–913 (2007)
Levy, S. et al. The diploid genome sequence of an individual human. PLoS Biol. 5, e254 (2007)
Wheeler, D. A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008)
Conrad, D. F. et al. A high-resolution survey of deletion polymorphism in the human genome. Nature Genet. 38, 75–81 (2006)
McCarroll, S. A. et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nature Genet. 40, 1166–1174 (2008)
Redon, R. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006)
Hurles, M. E., Dermitzakis, E. T. & Tyler-Smith, C. The functional impact of structural variation in humans. Trends Genet. 24, 238–245 (2008)
Stranger, B. E. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007)
Buchanan, J. A. & Scherer, S. W. Contemplating effects of genomic structural variation. Genet. Med. 10, 639–647 (2008)
McCarroll, S. A. et al. Deletion polymorphism upstream of IRiGM associated with altered IRGM expression and Crohn’s disease. Nature Genet. 40, 1107–1112 (2008)
Willer, C. J. et al. Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nature Genet. 41, 25–34 (2009)
de Cid, R. et al. Deletion of the late cornified envelope LCE3B and LCE3C genes as a susceptibility factor for psoriasis. Nature Genet. 41, 211–215 (2009)
Yang, T. L. et al. Genome-wide copy-number-variation study identified a susceptibility gene, UGT2B17, for osteoporosis. Am. J. Hum. Genet. 83, 663–674 (2008)
Lee, C., Iafrate, A. J. & Brothman, A. R. Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nature Genet. 39 (suppl). S48–S54 (2007)
Kidd, J. M. et al. Mapping and sequencing of structural variation from eight human genomes. Nature 453, 56–64 (2008)
Korbel, J. O. et al. Paired-end mapping reveals extensive structural variation in the human genome. Science 318, 420–426 (2007)
Gu, W., Zhang, F. & Lupski, J. R. Mechanisms for human genomic rearrangements. Pathogenetics 1, 4 (2008)
Barnes, C. et al. A robust statistical method for case-control association testing with copy number variation. Nature Genet. 40, 1245–1252 (2008)
Lohmueller, K. E. et al. Proportionally more deleterious genetic variation in European than in African populations. Nature 451, 994–997 (2008)
Ng, P. C. et al. Genetic variation in an individual human exome. PLoS Genet. 4, e1000160 (2008)
Kim, P. M., Korbel, J. O. & Gerstein, M. B. Positive selection at the protein network periphery: evaluation in terms of structural constraints and cellular context. Proc. Natl Acad. Sci. USA 104, 20274–20279 (2007)
Tuzun, E. et al. Fine-scale structural variation of the human genome. Nature Genet. 37, 727–732 (2005)
Jeffreys, A. J. et al. Human minisatellites, repeat DNA instability and meiotic recombination. Electrophoresis 20, 1665–1675 (1999)
Bacolla, A. & Wells, R. D. Non-B DNA conformations, genomic rearrangements, and human disease. J. Biol. Chem. 279, 47411–47414 (2004)
Myers, S. et al. A common sequence motif associated with recombination hot spots and genome instability in humans. Nature Genet. 40, 1124–1129 (2008)
Jeffreys, A. J. et al. Meiotic recombination hot spots and human DNA diversity. Phil. Trans. R. Soc. Lond. B 359, 141–152 (2004)
Huppert, J. L. & Balasubramanian, S. G-quadruplexes in promoters throughout the human genome. Nucleic Acids Res. 35, 406–413 (2007)
Down, T. A. & Hubbard, T. J. NestedMICA: sensitive inference of over-represented motifs in nucleic acid sequence. Nucleic Acids Res. 33, 1445–1453 (2005)
Sen, S. K. et al. Human genomic deletions mediated by recombination between Alu elements. Am. J. Hum. Genet. 79, 41–53 (2006)
Tian, D. et al. Single-nucleotide mutation rate increases close to insertions/deletions in eukaryotes. Nature 455, 105–108 (2008)
Campbell, P. J. et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nature Genet. 40, 722–729 (2008)
Pickeral, O. K., Makalowski, W., Boguski, M. S. & Boeke, J. D. Frequent human genomic DNA transduction driven by LINE-1 retrotransposition. Genome Res. 10, 411–415 (2000)
Gondo, Y. et al. High-frequency genetic reversion mediated by a DNA duplication: the mouse pink-eyed unstable mutation. Proc. Natl Acad. Sci. USA 90, 297–301 (1993)
Boyko, A. R. et al. Assessing the evolutionary impact of amino acid mutations in the human genome. PLoS Genet. 4, e1000083 (2008)
Emerson, J. J., Cardoso-Moreira, M., Borevitz, J. O. & Long, M. Natural selection shapes genome-wide patterns of copy-number polymorphism in Drosophila melanogaster . Science 320, 1629–1631 (2008)
Wang, L. L. et al. Intron-size constraint as a mutational mechanism in Rothmund-Thomson syndrome. Am. J. Hum. Genet. 71, 165–167 (2002)
Sabeti, P. C. et al. Genome-wide detection and characterization of positive selection in human populations. Nature 449, 913–918 (2007)
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006)
Smith, E. E. & Malik, H. S. The apolipoprotein L family of programmed cell death and immunity genes rapidly evolved in primates at discrete sites of host-pathogen interactions. Genome Res. 19, 850–858 (2009)
Pickrell, J. K. et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 19, 826–837 (2009)
Silva, A. M. et al. Ethnicity-related skeletal muscle differences across the lifespan. Am. J. Hum. Biol. 10.1002/ajhb.20956 (16 June 2009)
MacArthur, D. G. et al. Loss of ACTN3 gene function alters mouse muscle metabolism and shows evidence of positive selection in humans. Nature Genet. 39, 1261–1265 (2007)
Nielsen, R. et al. Darwinian and demographic forces affecting human protein coding genes. Genome Res. 19, 838–849 (2009)
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009)
Pique-Regi, R. et al. Sparse representation and Bayesian detection of genome copy number alterations from microarray data. Bioinformatics 24, 309–318 (2008)
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007)
Iafrate, A. J. et al. Detection of large-scale variation in the human genome. Nature Genet. 36, 949–951 (2004)
Acknowledgements
We would like to thank A. Boyko, J. J. Emerson, J. Pickrell, S. Kudaravalli, J. Pritchard, T. Down, S. McCarroll, J. Collins, C. Beazley, M. Dermitzakis, P. Eis, T. Richmond, M. Hogan, D. Bailey, S. Giles, G. Speight, N. Sparkes, D. Peiffer, C. Chen, K. Li, P. Oeth, D. Stetson and D. Church for advice, sharing data, sharing software and technical assistance. We are grateful for the efforts and support of our colleagues at NimbleGen, Agilent, Illumina, Applied Biosystems and Sequenom. We thank J. Barrett for comments on an earlier version of the manuscript. The Centre for Applied Genomics at the Hospital for Sick Children and Wellcome Trust Sanger Institute are acknowledged for database, technical assistance and bioinformatics support. This research was supported by the Wellcome Trust (grant no. 077006/Z/05/Z; to M.E.H., N.P.C., C.T.-S.), Canada Foundation of Innovation and Ontario Innovation Trust (to S.W.S.), Canadian Institutes of Health Research (CIHR) (to S.W.S.), Genome Canada/Ontario Genomics Institute (to S.W.S.), the McLaughlin Centre for Molecular Medicine (to S.W.S.), Ontario Ministry of Research and Innovation (to S.W.S.), the Hospital for Sick Children Foundation (to S.W.S.), the Department of Pathology at Brigham and Women’s Hospital (to C.L.) and the National Institutes of Health (NIH) (grants HG004221 and GM081533; to C.L.). K.K. is supported by the Academy of Finland. D.P. is supported by fellowships from the Royal Netherlands Academy of Arts and Sciences (TMF/DA/5801) and the Netherlands Organization for Scientific Research (Rubicon 825.06.031). S.W.S. holds the GlaxoSmithKline Pathfinder Chair in Genetics and Genomics at the University of Toronto and the Hospital for Sick Children.
Author Contributions C.T.-S., N.P.C., C.L., S.W.S. and M.E.H. are all joint senior authors, and planned and managed the project. D.F.C. and D.P. lead the data analysis. Data analyses were performed by D.F.C., D.P., R.R., L.F., O.G., Y.Z., J.A., T.D.A., C.B., P.C., T.F., M.H., C.H.I., K.K., D.G.M., J.R.M., I.O., A.W.C.P., S.R., K.S., A.V., K.W., J.W. and M.E.H. The WTCCC collaborated on array design. Validation experiments were performed by Y.Z. and M.H. D.F.C., D.P., S.W.S. and M.E.H. wrote the paper.
Author information
Authors and Affiliations
Consortia
Corresponding authors
Additional information
Lists of participants and affiliations appear in Supplementary Information.
Supplementary information
Supplementary Notes
This file contains Supplementary Notes, including Figures 1.1-1.12, Tables 1.1-1.7 and an Appendix of WTCCC authors and their affiliations. (PDF 3783 kb)
Supplementary Methods
This file contains Supplementary Methods, including Figures 2.1-2.30 and Tables 2.1-2.9, References and Appendices. (PDF 7430 kb)
Supplementary Table
This file contains Supplementary Table 1: CNV map. Genomic locations for all 11,700 candidate CNVs, including the number of CEU and YRI individuals in which the CNV was detected during the discovery experiment. (XLS 1860 kb)
Supplementary Table
This file contains Supplementary Table 2: CNV genotypes. Absolute integer copy number estimates for 5,238 CNVs in 450 individuals from 4 HapMap populations. (XLS 15811 kb)
Rights and permissions
About this article
Cite this article
Conrad, D., Pinto, D., Redon, R. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010). https://doi.org/10.1038/nature08516
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08516
This article is cited by
-
A cohort study of neurodevelopmental disorders and/or congenital anomalies using high resolution chromosomal microarrays in southern Brazil highlighting the significance of ASD
Scientific Reports (2024)
-
Ethnic and functional differentiation of copy number polymorphisms in Tunisian and HapMap population unveils insights on genome organizational plasticity
Scientific Reports (2024)
-
Genome integrity as a potential index of longevity in Ashkenazi Centenarian’s families
GeroScience (2024)
-
High-resolution structural variants catalogue in a large-scale whole genome sequenced bovine family cohort data
BMC Genomics (2023)
-
Exploring quantitative traits-associated copy number deletions through reanalysis of UK10K consortium whole genome sequencing cohorts
BMC Genomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.